Development: Computing away the magic?
Multicellular organisms employ a variety of mechanisms to ensure that genes are expressed at the right time and place throughout their life cycles. The transcription of DNA into RNA is augmented by activators and diminished by repressors. Both classes of regulatory proteins bind to specific sequences contained within enhancers, which are the key agents of gene regulation in higher organisms. Elucidating how enhancers work is critical for understanding gene regulation in development and disease.
It is over 30 years since Banerji and Schaffner discovered that enhancers can be physically separate from the genes they regulate (Banerji et al., 1981). Enhancers can map quite far—1 million base pairs or more—from their target genes (Amano et al., 2009). This action at a distance is a defining property of complex organisms, and contrasts with what happens in simple bacteria, where most activator and repressor binding sites are found quite close to their target genes (see, e.g., Levine and Tjian, 2003).
One of the most widely studied enhancers is the eve stripe 2 enhancer in the fruit fly Drosophila melanogaster (Small et al., 1992). The body of the Drosophila embryo is made up of 14 segments, and a gene called eve (even-skipped) is expressed in the even-numbered segments, giving rise to a distinctive pattern of seven stripes (Figure 1A). It was initially thought that the long-range diffusion of morphogens (Turing, 1952)—signaling molecules that influence tissue development through their formation of concentration gradients—coordinated the expression of all seven eve stripes (Meinhardt, 1986). The discovery that eve stripe 2 had its own dedicated enhancer led one researcher to complain of the ‘inelegance’ of such a mechanism (Akam, 1989). However, we have now come full circle: I cannot help but complain that the new models for the regulation of eve expression described by Nicholas Luscombe and co-workers in eLife seem to strip the mystique from the eve stripe 2 enhancer (Ilsley et al., 2013).
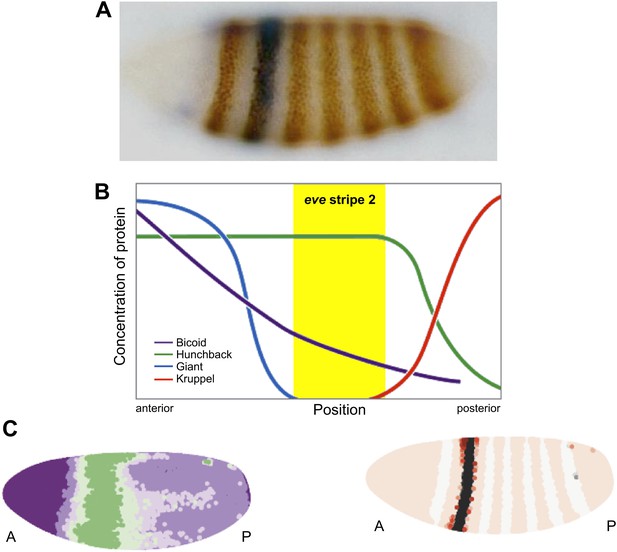
Regulation of eve stripe 2. The gene eve is expressed in the even-numbered body segments of Drosophila embryos, giving rise to a distinctive pattern of stripes. A, Transgenic embryo expressing an eve.2>lacZ fusion gene. The endogenous eve stripes are stained brown, while stripe 2 is stained blue (Small et al., 1992). B, The transcription factors Krüppel and Giant (repressors) and Bicoid and Hunchback (activators) are expressed in distinct patterns along the Drosophila embryo, and their combined effects dictate the position of eve stripe 2 (Watson et al., 2014). C, Computer simulations can be used to model the expression gradient of Bicoid (left) and the resulting effect on the position of eve stripe 2 (right). A, anterior; P, posterior.
The stripe 2 enhancer is regulated by four different transcription factors in the early Drosophila embryo—two activators, Bicoid and Hunchback; and two repressors, Giant and Krüppel (Small et al., 1992). There are 12 binding sites for these transcription factors distributed over the length of the enhancer, and the combined effects of these four proteins dictate the location of the second eve stripe (Figure 1B). In principle, Bicoid and Hunchback can activate the eve stripe 2 enhancer in the entire anterior half of the embryo (from the head to the anterior thorax); however, localized repressors—Giant and Krüppel—delineate eve expression within the stripe 2 domain.
Luscombe and co-workers—including Garth Ilsley as first author—investigated how these four transcription factors produce the stripe 2 expression pattern (Figure 1B), by combining quantitative imaging with computer simulations of different mathematical models. They used this same approach to model the enhancer that regulates stripes 3 and 7, but for simplicity I will restrict my discussion to stripe 2. The resulting models provide new insights into the mechanisms of stripe formation during development. First, IIsley et al. argue that the order of the Bicoid, Hunchback, Giant and Krüppel binding sites is unlikely to be important for stripe 2 expression. They base this on the observation that models in which the effects of activators can simply be added to those of repressors are sufficient to produce the stripe 2 pattern, and there is no need to assume that activators bound to adjacent sites cooperate with each other to augment their activities. Moreover, there is no indication of nonlinear effects such as ‘repression dominance’, whereby repressors downregulate transcription more than activators upregulate it (Arnosti et al., 1996). Rather, the models call for a simple balance between the effects of activators and those of repressors.
The most interesting implication of this work is that Bicoid might not function solely as an activator (Driever et al., 1989; Struhl et al., 1989). Luscombe and co-workers were able to achieve more faithful simulations of the stripe 2 expression pattern by assuming that Bicoid, which is most abundant in the anterior region of the embryo and gradually declines in concentration towards the posterior end, acts as both an activator and a repressor. Ilsley et al. propose that high levels of Bicoid repress expression of stripe 2 in anterior regions, while lower levels in the more central regions activate its expression (Figure 1C).
The idea that a transcription factor can mediate both activation and repression is not new. However, this is the first time that such a dual mechanism has been suggested for Bicoid, the lynchpin of anterior–posterior patterning. This dual function of Bicoid can explain why eve, and many other segmentation genes, are silent at the anterior pole of the Drosophila embryo (Andrioli et al., 2002).
In summary, the eve stripe 2 enhancer produces an exquisite on/off pattern of expression in response to crude gradients of transcription factors, and its ability to do so has previously been explained by nonlinear interactions between proteins. By arguing against such nonlinearity, Ilsley et al. seemingly strip the magic from the stripe 2 enhancer. But is the magic really gone? How the enhancer determines whether Bicoid functions as an activator or a repressor is uncertain. Hence, I believe that the concept of the enhancer as a template for weak protein interactions is alive and well, and yes, still a mystery.
References
-
Anterior repression of a Drosophila stripe enhancer requires three position-specific mechanismsDevelopment 129:4931–4940.
-
The gap protein knirps mediates both quenching and direct repression in the Drosophila embryoEMBO J 15:3659–3666.
-
The Chemical Basis of MorphogenesisPhil Trans Royal Soc London Series B 237:37–72.https://doi.org/10.1098/rstb.1952.0012
Article and author information
Author details
Publication history
- Version of Record published: August 6, 2013 (version 1)
Copyright
© 2013, Levine
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 1,228
- views
-
- 81
- downloads
-
- 4
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Developmental Biology
Inhibitory G alpha (GNAI or Gαi) proteins are critical for the polarized morphogenesis of sensory hair cells and for hearing. The extent and nature of their actual contributions remains unclear, however, as previous studies did not investigate all GNAI proteins and included non-physiological approaches. Pertussis toxin can downregulate functionally redundant GNAI1, GNAI2, GNAI3, and GNAO proteins, but may also induce unrelated defects. Here, we directly and systematically determine the role(s) of each individual GNAI protein in mouse auditory hair cells. GNAI2 and GNAI3 are similarly polarized at the hair cell apex with their binding partner G protein signaling modulator 2 (GPSM2), whereas GNAI1 and GNAO are not detected. In Gnai3 mutants, GNAI2 progressively fails to fully occupy the sub-cellular compartments where GNAI3 is missing. In contrast, GNAI3 can fully compensate for the loss of GNAI2 and is essential for hair bundle morphogenesis and auditory function. Simultaneous inactivation of Gnai2 and Gnai3 recapitulates for the first time two distinct types of defects only observed so far with pertussis toxin: (1) a delay or failure of the basal body to migrate off-center in prospective hair cells, and (2) a reversal in the orientation of some hair cell types. We conclude that GNAI proteins are critical for hair cells to break planar symmetry and to orient properly before GNAI2/3 regulate hair bundle morphogenesis with GPSM2.
-
- Computational and Systems Biology
- Developmental Biology
Organisms utilize gene regulatory networks (GRN) to make fate decisions, but the regulatory mechanisms of transcription factors (TF) in GRNs are exceedingly intricate. A longstanding question in this field is how these tangled interactions synergistically contribute to decision-making procedures. To comprehensively understand the role of regulatory logic in cell fate decisions, we constructed a logic-incorporated GRN model and examined its behavior under two distinct driving forces (noise-driven and signal-driven). Under the noise-driven mode, we distilled the relationship among fate bias, regulatory logic, and noise profile. Under the signal-driven mode, we bridged regulatory logic and progression-accuracy trade-off, and uncovered distinctive trajectories of reprogramming influenced by logic motifs. In differentiation, we characterized a special logic-dependent priming stage by the solution landscape. Finally, we applied our findings to decipher three biological instances: hematopoiesis, embryogenesis, and trans-differentiation. Orthogonal to the classical analysis of expression profile, we harnessed noise patterns to construct the GRN corresponding to fate transition. Our work presents a generalizable framework for top-down fate-decision studies and a practical approach to the taxonomy of cell fate decisions.





