Meiosis I chromosome segregation is established through regulation of microtubule–kinetochore interactions
Figures
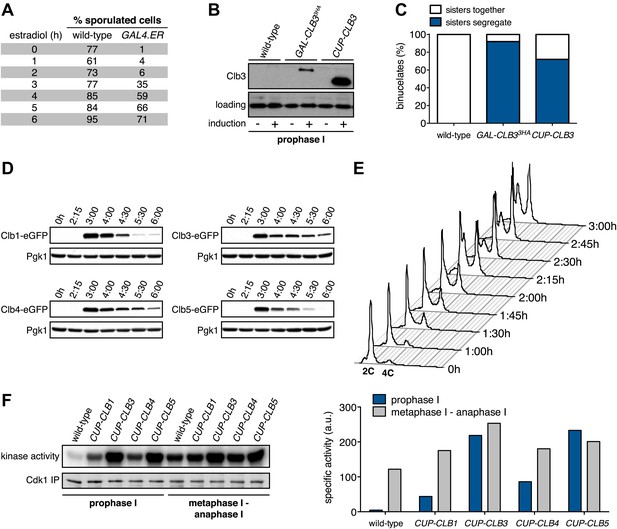
Characterization of premature cyclin expression and corresponding total CDK activity.
(A) Wild-type (A4962) and GAL4-ER (A19151) cells were induced to sporulate. At the indicated time points, an aliquot was removed and treated with estradiol (1 μM). The percentage of cells that had sporulated after 24 hr was calculated as the sum of dyads, triads and tetrads divided by the total number of cells (n > 100 cells counted for each condition). (B) Wild-type (A18686), GAL-CLB3-3HA (A23084) and CUP-CLB3 (A23086) cells also carrying the GAL4-ER fusion were induced to sporulate. After 3 hr, CLB3 was induced. Each culture was treated with estradiol (1 μM) and CuSO4 (50 μM). Cells were harvested after 1 hr of estradiol and CuSO4 treatment for protein extraction. Levels of Clb3 were examined by Western blot analysis. A cross-reacting band was used as a loading reference. (C) Segregation of sister chromatids (equational division) using heterozygous GFP dots integrated at LEU2 (∼20 kb from CENIII) was quantified in binucleate cells from wild-type (A18686), GAL-CLB3-3HA (A23084) and CUP-CLB3 (A23086). Note that the samples were collected from the same experiment described in (B) at a time point when a fraction of the cells had completed meiosis I (6 hr 30 min and 7 hr after induction of sporulation) (n > 100 for each sample). Using a chi-square test (df 1), the fraction of binucleates that display a reductional or equational division was compared between wild-type and GAL-CLB3-3HA χ2 = 166.4, p<0.0001 and between wild-type and CUP-CLB3 χ2 = 108.7, p<0.0001. (D) Wild-type or CUP-CLB-eGFP cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate. After 2 hr 15 min, cyclins were induced by addition of CuSO4 (50 μM). Cells were released from the NDT80 block at 4 hr 30 min post transfer to sporulation medium. Cyclin levels monitored by Western blot at the indicated time points in CUP-CLB1-eGFP (A28531), CUP-CLB3-eGFP (A28533), CUP-CLB4-eGFP (A28535) and CUP-CLB5-eGFP (A33199) cells. Pgk1 was used as a loading control. (E) Wild-type (A22678) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Samples were taken at indicated time points to determine DNA content by flow cytometry. By 2 hr 15 min 43% of cells had a 4C DNA content. (F) Left: Wild-type (A28663), CUP-CLB1 (A28665), CUP-CLB3 (A28667), CUP-CLB4 (A28669) and CUP-CLB5 (A28671) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. In vitro kinase assays were performed with Cdc28-3V5 (Cdk1) immunoprecipitated from prophase I samples (collected 4 hr 30 min after sporulation induction, at the time of NDT80 block-release) and metaphase I–anaphase I samples (collected 1 hr 30 min after release from the NDT80 block). Right: specific activity was calculated by normalizing the amount of phosphorylated Histone H1 to the amount of immunoprecipitated Cdc28-3V5 using ImageQuant software (Molecular Dynamics, Sunnyvale, CA).
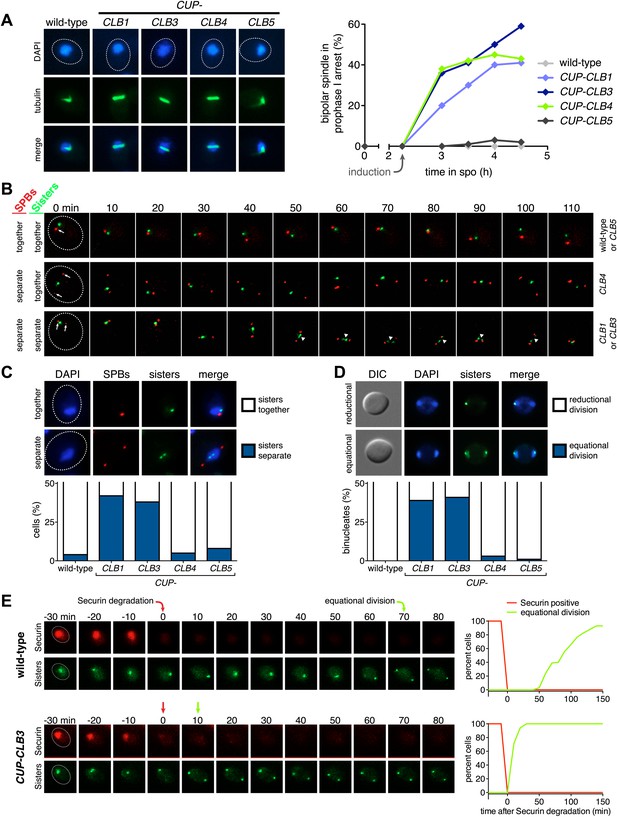
Premature expression of CLB1 or CLB3 causes sister kinetochore biorientation during prophase I and sister chromatid segregation in meiosis I.
Wild-type or CUP-CLB cells were induced to sporulate. After 2 hr 15 min, cyclins were induced by addition of CuSO4 (50 μM). Cells were either arrested during prophase I or released from an NDT80 block 4 hr 30 min after induction of sporulation. (A) Bipolar spindle formation determined in wild-type (A22678), CUP-CLB1 (A27421), CUP-CLB3 (A22702), CUP-CLB4 (A27423) and CUP-CLB5 (A27425) during prophase I (n = 100 per time point). Images on left show spindle formation in CUP-CLB cells 4 hr after induction of sporulation; in this and all subsequent Figures microtubules are shown in green and DNA in blue. The dotted line depicts the cell membrane. (B) Microtubule–kinetochore engagement monitored during prophase I, starting at 1 hr after CuSO4 addition in wild-type (A30700), CUP-CLB1 (A30702), CUP-CLB3 (A30704), CUP-CLB4 (A30707) and CUP-CLB5 (A30708) by live cell microscopy. SPBs (marked by arrow) and heterozygous CENV-GFP dots are shown (arrowheads mark separated CENV dots). In this and all subsequent figures SPBs are in red, GFP dots are in green. (C) Top panel: representative images of wild-type (A30700) and CUP-CLB3 (A30704). Bottom panel: separation of heterozygous CENV-GFP dots in prophase I-arrested cells quantified in wild-type (A22678), CUP-CLB1 (A27421), CUP-CLB3 (A22702), CUP-CLB4 (A27423) and CUP-CLB5 (A27425) by live cell microscopy (over the duration of 8 hr, n > 100) as described in the ‘Materials and methods’. The fraction of nuclei that display sister kinetochores as separate or together for each CUP-CLB strain was compared to wild-type using a chi-square test (df 1): CUP-CLB1, χ2 = 40.77, p<0.0001; CUP-CLB3, χ2 = 34.84, p<0.0001; CUP-CLB4, χ2 = 0.1163, p=0.7330; CUP-CLB5, χ2 = 1.418, p=0.2337. (D) Segregation of sister chromatids (equational division) using heterozygous CENV-GFP dots quantified in binucleates from wild-type (A22678), CUP-CLB1 (A27421), CUP-CLB3 (A22702), CUP-CLB4 (A27423) and CUP-CLB5 (A27425) (n = 100). The fraction of binucleates that display a reductional or equational division for each CUP-CLB strain was compared to wild-type using a chi-square test (df 1): CUP-CLB1, χ2 = 45.13, p<0.0001; CUP-CLB3, χ2 = 48.22, p<0.0001; CUP-CLB4, χ2 = 1.020, p=0.3124; CUP-CLB5, χ2 = 0, p=1. (E) Wild-type (A31019) and CUP-CLB3 (A31021) cells monitored for segregation of heterozygous CENV-GFP dots with respect to Pds1 (Securin, red) degradation by live cell microscopy (n > 17). Time of Pds1 degradation set to t = 0, percent cells were plotted as a Kaplan–Meier curve. Note that for A31021, the analysis of cells that segregate sister chromatids in the first nuclear division is shown. Pds1 accumulation during prophase II is not observed using the Pds1-tdTomato construct, likely due to delayed maturation of the fluorophore (Katis et al., 2010).
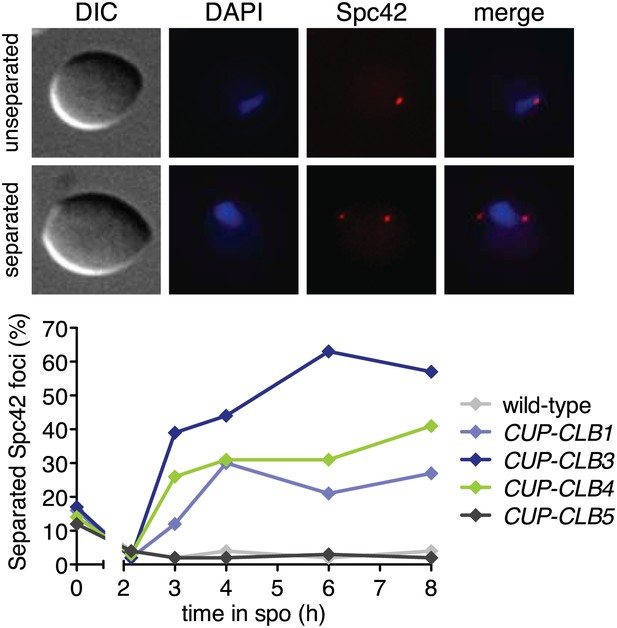
Spindle pole body separation in CUP-CLB cells.
Wild-type or CUP-CLB cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate. After 2 hr 15 min, cyclins were induced by addition of CuSO4 (50 μM). Cells were arrested during prophase I and the percentage of cells with separated Spc42 foci (red dots) was determined at indicated time points in wild-type (A29581), CUP-CLB1 (A29582), CUP-CLB3 (A29583), CUP-CLB4 (A29584) and CUP-CLB5 (A29585) (n > 100 for each time point).
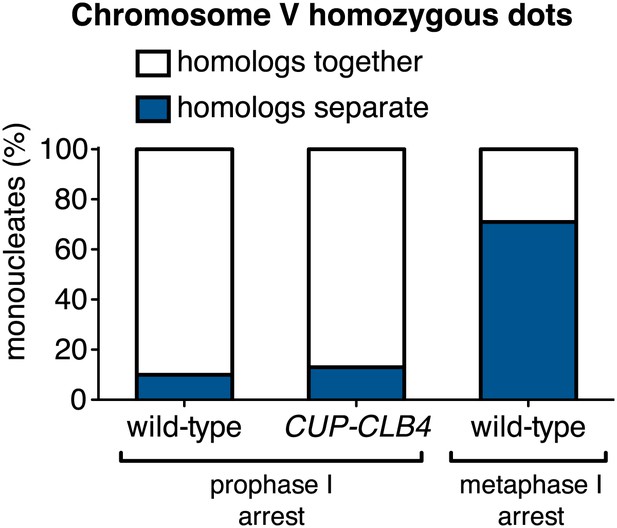
Homolog separation in CUP-CLB4 cells.
Wild-type (A22688), CUP-CLB4 (A32470) also carrying the GAL4-ER and GAL-NDT80 fusions or cdc20-mn (A15163) cells all carrying homozygous CENV GFP dots were induced to sporulate. After 2 hr 15 min, cyclins were induced by addition of CuSO4 (50 μM). Cells were arrested either during prophase I (A22688, A324470) or metaphase I (A15163). Separated GFP foci (homologs separate) were analyzed 6 hr (prophase I-arrest) or 8 hr 30 min (metaphase I-arrest) after induction of sporulation (n > 100 for each time point). Using a chi-square test (df 1), the fraction of mononucleates that display homologs as together or separate during a prophase I arrest was compared between wild-type and CUP-CLB4 χ2 = 0.4422, p=0.5061.
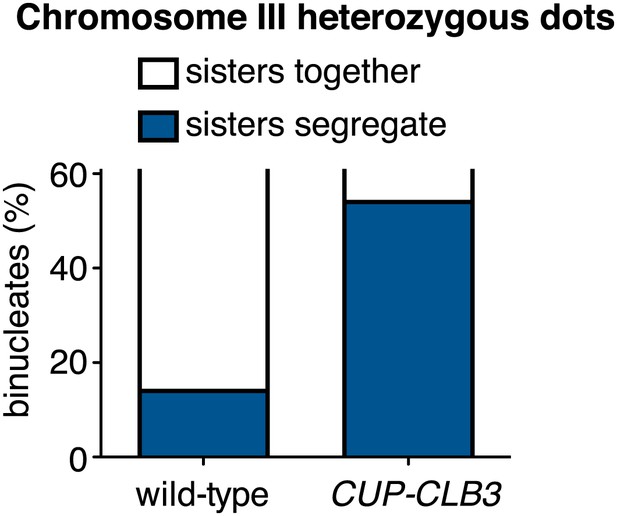
Chromosome III sister chromatid segregation in CUP-CLB3 cells.
Wild-type (A18185) and CUP-CLB3 (A22682) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate. After 3 hr, CLB3 was induced by addition of CuSO4 (50 μM). At 6 hr, cells were released from the NDT80 block. Subsequently, segregation of sister chromatids (equational division) using heterozygous GFP dots integrated at LEU2 (∼20 kb from CENIII) was quantified in binucleate cells. The appearance of segregated sister chromatids in wild-type is likely due to recombination between LEU2 and CEN3. Using a chi-square test (df 1), the fraction of binucleates that display a reductional or equational division was compared between wild-type and CUP-CLB3 χ2 = 35.65, p<0.0001.
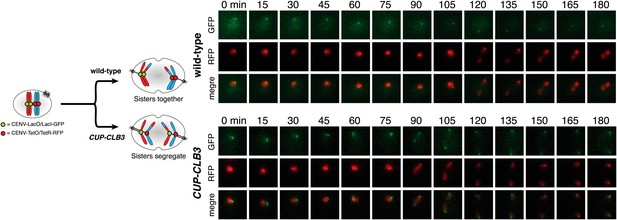
Sister chromatid segregation in CUP-CLB3 cells using dual-color marked chromosomes.
Wild-type (A27476) and CUP-CLB3 (A27480) cells carrying the GAL4-ER and GAL-NDT80 fusions and CENV-LacO/LacI-GFP on one homolog of chromosome V (green) and CENV-tetO/tetR-RFP on the other homolog of chromosome V (red) were induced to sporulate and CuSO4 (50 μM) was added at 2 hr 15 min. At 4 hr 30 min, cells were released from NDT80 block and monitored by live cell microscopy starting 30 min after estradiol addition, and monitored every 15 min for 8 hr.
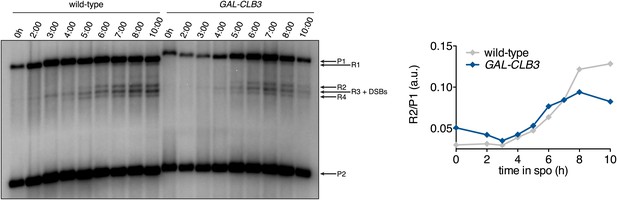
Recombination in CUP-CLB3 cells.
Left panel: wild-type (A21104) and GAL-CLB3 (A21105) cells were induced to sporulate and estradiol (1 μM) was added 3 hr after transfer into sporulation medium. Genomic DNA was prepared and digested with XhoI and MluI and hybridized with Probe A. See Storlazzi et al. (1995) for details. Right panel: recombination products were quantified as R2/P1. Note: A21104 and A21105 contain auxotrophies and have reduced meiotic kinetics relative to prototrophic strains.
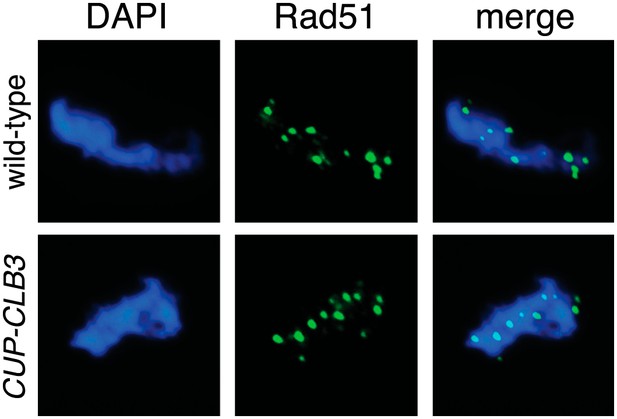
Localization of Rad51 in CUP-CLB3 cells.
Wild-type (A22864) and CUP-CLB3 (A22866) cells were induced to sporulate and CuSO4 (50 μM) was added 3 hr after transfer into sporulation medium. Localization of the double-strand break repair protein Rad51 (green) was determined by nuclear spreads 4 hr after transfer to sporulation medium. DNA is shown in blue.
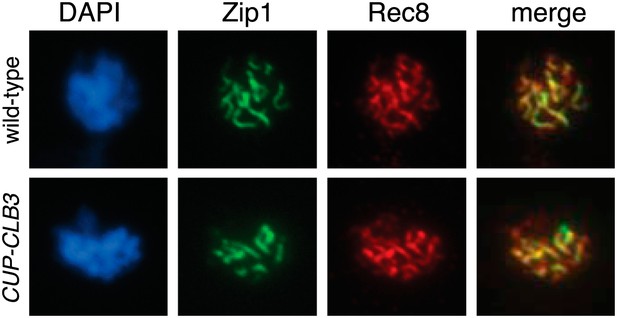
Localization of Zip1 in CUP-CLB3 cells.
Wild-type (A22836) and CUP-CLB3 (A22838) cells were induced to sporulate and CuSO4 (50 μM) was added 3 hr after transfer into sporulation medium. Localization of the synaptonemal complex component Zip1 (green) and the cohesin subunit Rec8-13myc (red) was determined by nuclear spreads 5 hr after transfer to sporulation medium. DNA is shown in blue.
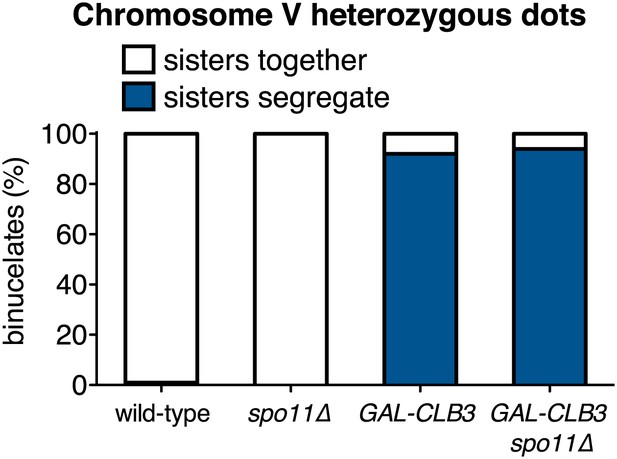
Preventing homologous recombination does not affect the phenotypes caused by premature CLB3 expression.
Wild-type (A19396), GAL-CLB3 (A19400), spo11∆ (A21193) and spo11∆ GAL-CLB3 (A21194) cells were induced to sporulate and estradiol (1 μM) was added 3 hr after transfer into sporulation medium. Subsequently, segregation of sister chromatids (equational division) was quantified using heterozygous CENV GFP dots in binucleate cells (n = 100). Note that CLB3-induced meiosis I sister chromatid segregation is higher in GAL-CLB3 cells than in CUP-CLB3 cells. This is presumably due to the more homogenous expression of CLB3 in cells where expression is driven from the GAL1-10 promoter. Using a chi-square test (df 1), the fraction of binucleates that display a reductional or equational division was compared between GAL-CLB3 and GAL-CLB3 spo11∆ χ2 = 0.3072, p=0.5794.
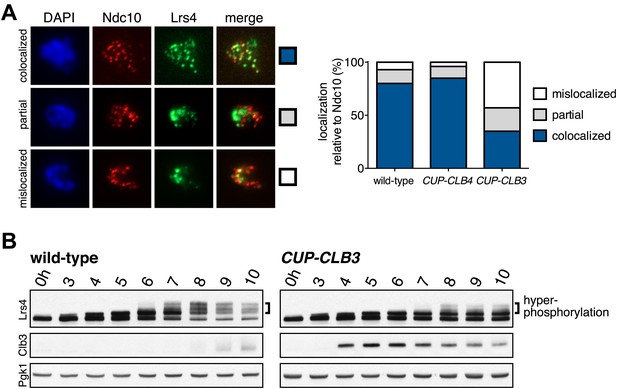
Premature CLB3 expression disrupts monopolin function.
(A) Lrs4-13myc (green) localization relative to Ndc10-6HA (red) was determined in spread nuclei from wild-type (A9217), CUP-CLB3 (A26278) and CUP-CLB4 (A29643) harboring a Cdc20 depletion allele (cdc20-mn) were induced to undergo sporulation and arrested in metaphase I due to depletion of Cdc20. CuSO4 was added at 3 hr after induction of sporulation (n > 40). The fraction of spread nuclei that display colocalized, partial or mislocalized Lrs4 with respect to Ndc10 was compared to wild-type using a chi-square test (df 2): CUP-CLB4, χ2 = 1.136, p=0.5666; CUP-CLB3, χ2 = 45.84, p<0.0001. (B) Western blots for Lrs4-13myc, Clb3 and Pgk1 from wild-type (A9217) and CUP-CLB3 (A26278) cells. Cells were sporulated as described in (A).
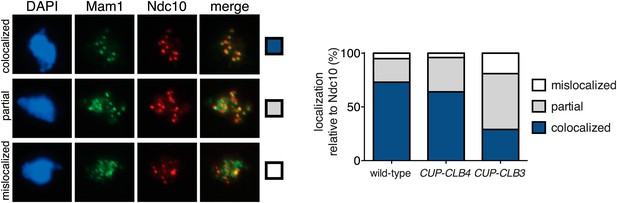
Monopolin association with kinetochores is disrupted in CUP-CLB3 but not in CUP-CLB4 cells.
Wild-type (A7450), CUP-CLB3 (A28673) and CUP-CLB4 (A28674) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Mam1-9myc (green) localization relative to Ndc10-6HA (red) was determined in spread nuclei from metaphase I-arrested cells (n > 40). DNA is shown in blue. Using a chi-square test (df 2) the fraction of spread nuclei that display colocalized, partial or mislocalized Mam1 with respect to Ndc10 was compared to wild-type: CUP-CLB4, χ2 = 2.554, p=0.2788; CUP-CLB3, χ2 = 39.31, p<0.0001.
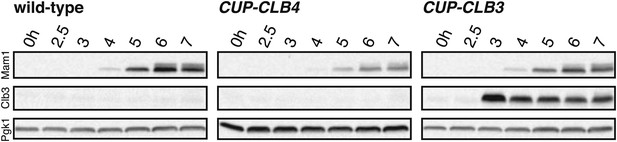
Premature Clb3 expression does not interfere with Mam1 expression.
Wild-type (A7450), CUP-CLB3 (A28673) and CUP-CLB4 (A28674) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Mam1 protein levels were analyzed to determine whether premature Clb3 expression interferes with Mam1 expression. Pgk1 was used as a loading control.
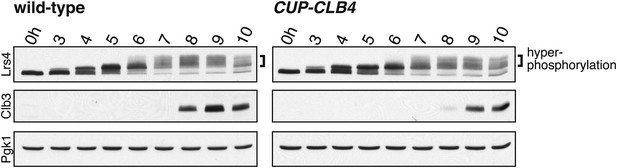
Lrs4 phosphorylation is not disrupted in CUP-CLB4 cells.
Wild-type (A26277) and CUP-CLB4 (A29643) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Levels of Lrs4, Clb3 and Pgk1 from cells arrested in metaphase I were examined by Western blot analysis.
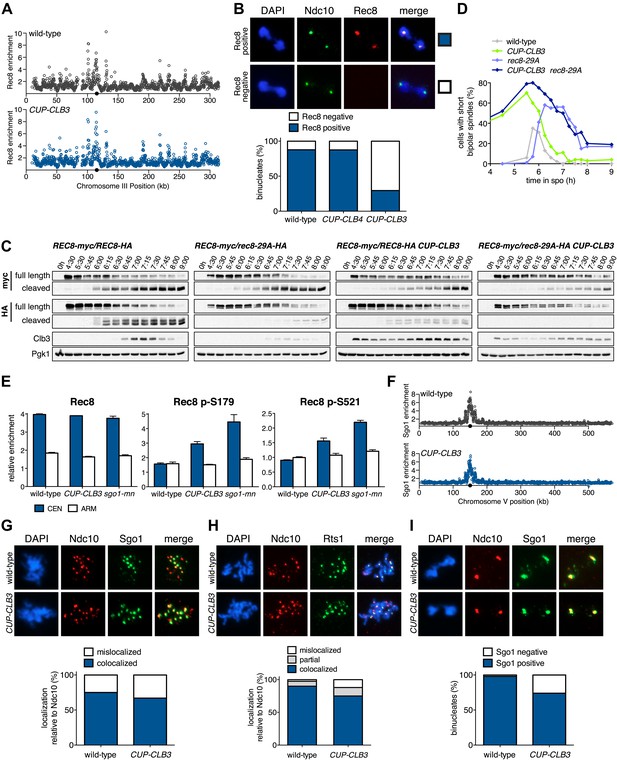
CLB3 misexpression disrupts protection of centromeric cohesin.
Cyclin expression was induced after 2 hr 15 min (C) and (D), 2 hr 30 min (A), (B), (E), (F) and (H) or 3 hr (G) and (I) of sporulation. (A) Chromosomal association of Rec8-13myc was monitored by ChIP-chip in wild-type (A28716) and CUP-CLB3 (A28718) during prophase I arrest. Centromere position is identified by a black circle. (B) Centromeric Rec8 localization was monitored in spread nuclei from wild-type (A28684), CUP-CLB3 (A28685) and CUP-CLB4 (A28686) cells carrying REC8-3HA (red) and NDC10-13myc (green) (n > 40). The fraction of spread nuclei that were Rec8 positive or negative was compared to wild-type using a chi-square test (df 1): CUP-CLB4, χ2 = 0.001323, p=0.9710; CUP-CLB3, χ2 = 32.79, p<0.0001. (C) Rec8 cleavage monitored by Western blot after release from an NDT80 block (4 hr 30 min) in wild-type and CUP-CLB3 carrying both a myc-tagged REC8 allele as well as either HA-tagged REC8 or rec8-29A allele (left to right: A29957, A29959, A29961, A29963). (D) Percentage of cells with short bipolar spindles was determined at indicated times in wild-type (A22804), CUP-CLB3 (A29965), rec8-29A (A22803) and CUP-CLB3 rec8-29A (A29967) after release from an NDT80 block (4 hr 30 min) (n = 100 per time point). (E) ChIP analysis for total Rec8, p-S179 Rec8 or p-S521 Rec8 from metaphase I-arrested (cdc20-mn) wild-type (A28681), CUP-CLB3 (A28682) and Sgo1-depleted (sgo1-mn; A29994) cells. Relative occupancy at a chromosome arm site (c194) or at a centromeric site (CENV) was determined relative to a low binding region (c281). Error bars represent range (n = 2). (F) Chromosomal association of Sgo1-3V5 was monitored by ChIP-chip in wild-type (A29795) and CUP-CLB3 (A29799) cells during prophase I-arrest. Centromere position is identified by a black circle. (G), (H) Localization of Sgo1-9myc (G, green) or Rts1-13myc (H, green) relative to Ndc10-6HA (red) determined by nuclear spreads in (G) wild-type (A22868) and CUP-CLB3 (A22870) or (H) wild-type (A28329) and CUP-CLB3 (A28330) during prophase I (n > 40). For (G), the fraction of spread nuclei that display colocalized or mislocalized Sgo1 relative to Ndc10 was compared between wild-type and CUP-CLB3 using a chi-square test (df 1) χ2 = 1.554, p=0.2125. For (H), the fraction of spread nuclei that display colocalized, partial or mislocalized Rts1 relative to Ndc10 was compared between wild-type and CUP-CLB3 using a chi-square test (df 2) χ2 = 3.712, p=0.1563. (I) Localization of Sgo1-9myc (green) in binucleates relative to Ndc10-6HA (red) determined by nuclear spreads from wild-type (A22868) and CUP-CLB3 (A22870) (n > 40). The fraction of spread nuclei that were Sgo1 positive or negative was compared between wild-type and CUP-CLB3 using a chi-square test (df 1) χ2 = 23.92, p<0.0001.
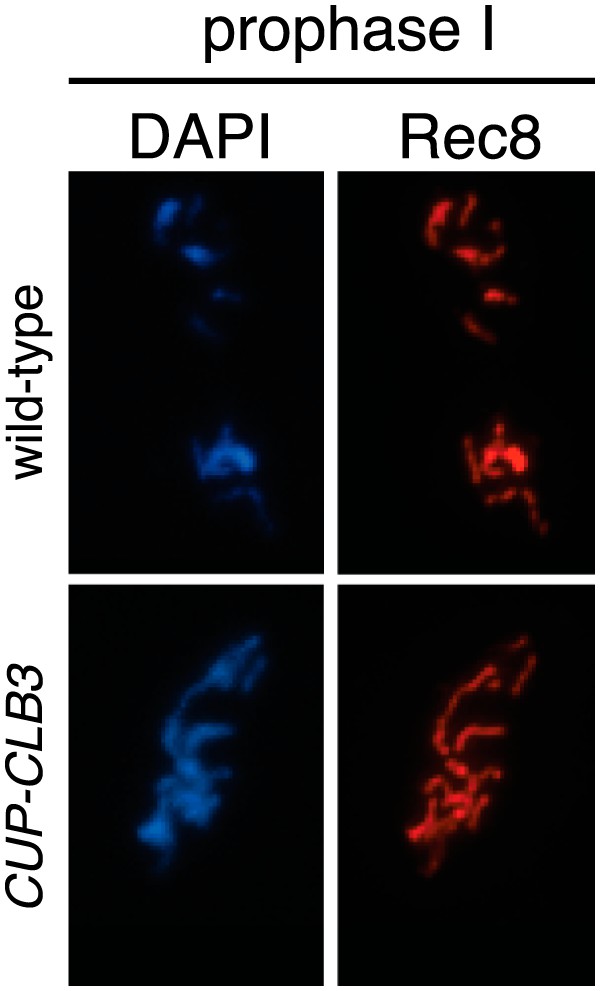
Chromosomal association of Rec8 in CUP-CLB3 cells.
Wild-type (A26547) and CUP-CLB3 (A26548) cells were induced to sporulate and CuSO4 (50 μM) was added 3 hr after transfer into sporulation medium. Rec8-3HA localization (red) was determined in spread nuclei from prophase I cells. DNA is shown in blue.
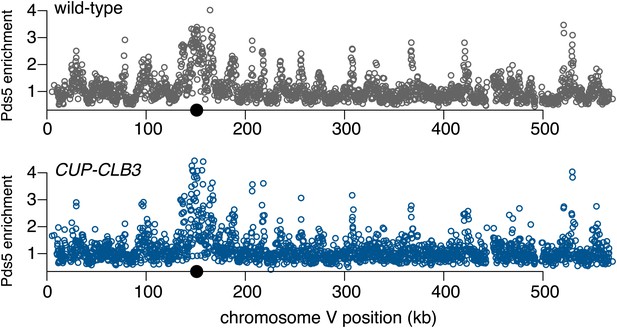
Chromosomal association of Pds5 in CUP-CLB3 cells.
Wild-type (A28681) and CUP-CLB3 (A28682) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Pds5 localization was determined by ChIP-chip from metaphase I-arrested cells. Black balls depict centromere positions.
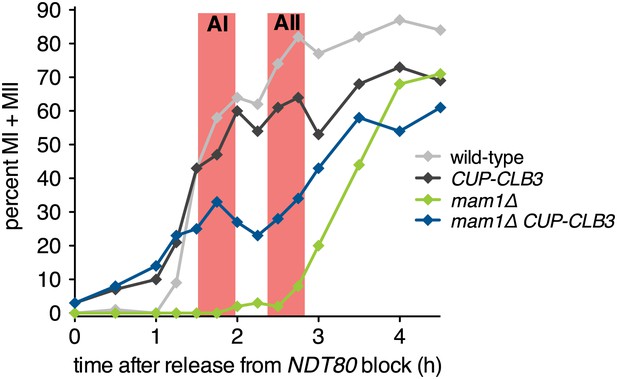
CUP-CLB3 cells partially bypass the nuclear division delay of mam1∆ cells.
Wild-type (A22678), CUP-CLB3 (A22702), mam1∆ (A31340) and mam1∆ CUP-CLB3 (A31342) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Cells were released from the NDT80 block 4 hr 30 min after transfer into sporulation medium. The percentage of cells that had undergone one or two meiotic divisions was determined at the indicated time points (n = 100 per time point).
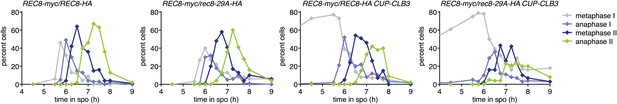
Meiotic progression of the cells analyzed for Rec8 cleavage in Figure 4C.
REC8-myc/REC8-HA (A29957), REC8-myc/rec8-29A-HA (A29961), REC8-myc/REC8-HA CUP-CLB3 (A29959) and REC8-myc/rec8-29A-HA CUP-CLB3 (A29963) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Cells were released from the NDT80 block 4 hr 30 min after transfer into sporulation medium. The percentage of cells in metaphase I (grey symbols), anaphase I (violet symbols), metaphase II (dark blue symbols) and anaphase II (green symbols) was determined at the indicated times (n = 100 per time point).
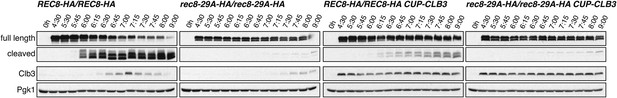
Analysis of Rec8 cleavage in cells used for Figure 4D.
Wild-type (A22804), CUP-CLB3 (A29965), rec8-29A (A22803) and rec8-29A CUP-CLB3 (A29967) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Cells were released from the NDT80 block 4 hr 30 min after transfer into sporulation medium. Levels of full-length Rec8, cleaved Rec8, Clb3 and Pgk1 were monitored by Western blot.
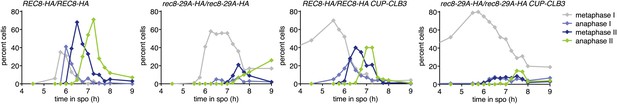
Meiotic progression of the cells analyzed for Rec8 cleavage in Figure 4D.
Wild-type (A22804), CUP-CLB3 (A29965), rec8-29A (A22803) and rec8-29A CUP-CLB3 (A29967) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Cells were released from the NDT80 block 4 hr 30 min after transfer into sporulation medium. The percentage of cells in metaphase I (grey symbols), anaphase I (violet symbols), metaphase II (dark blue symbols) and anaphase II (green symbols) was determined at the indicated times (n = 100 per time point).
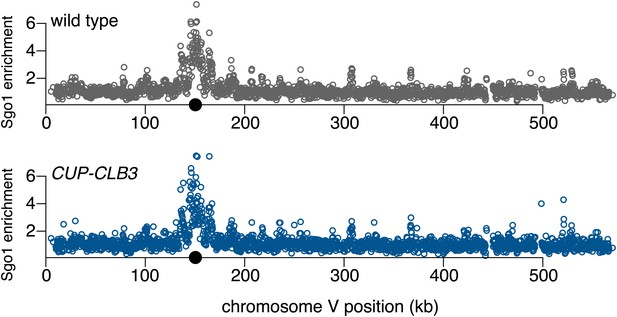
Chromosomal association of Sgo1 in CUP-CLB3 cells.
Wild-type (A28712) and CUP-CLB3 (A28713) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Sgo1-3V5 localization was determined by ChIP-chip, 7 hr after transfer into sporulation medium when cells were arrested in metaphase I. Arm peaks for Sgo1 correspond to cohesin-associated regions. The basis for Sgo1 enrichment at these sites is currently unclear. Black balls depict centromere positions.
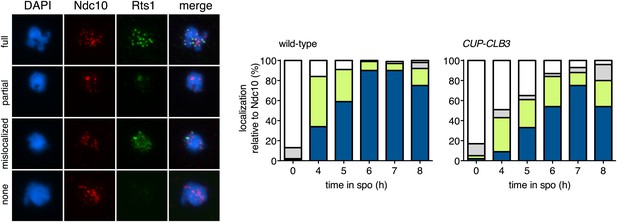
Localization of Rts1 in CUP-CLB3 cells.
Wild-type (A28331) and CUP-CLB3 (A28332) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Rts1-13myc (green) localization relative to Ndc10-6HA (red) was determined in spread nuclei from metaphase I-arrested cells (n > 40).
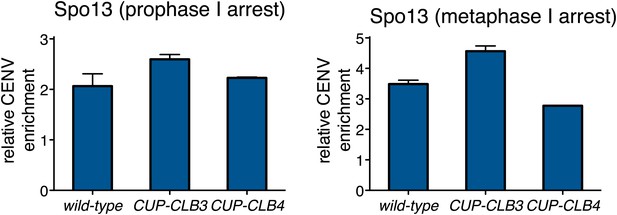
Chromosomal association of Spo13 in CUP-CLB3 cells.
(Left panel) wild-type (A30856), CUP-CLB3 (A30858) and CUP-CLB4 (A30860) cells carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate and CuSO4 (50 μM) was added 2 hr 15 min after transfer into sporulation medium. Spo13-3V5 localization was determined by ChIP from prophase I-arrested cells. Relative occupancy at a centromeric site (CEN5) relative to a low binding region (HMR) was determined. Error bars represent the range (n = 2). (Right panel) wild-type (A30743), CUP-CLB3 (A30745) and CUP-CLB4 (A30747) cells carrying the cdc20-mn allele were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Spo13-3V5 localization was determined by ChIP 7 hr after transfer into sporulation medium when cells were arrested in metaphase I. Relative occupancy at a centromeric site (CENV) relative to a low binding region (HMR) was determined. Error bars represent the range (n = 2).
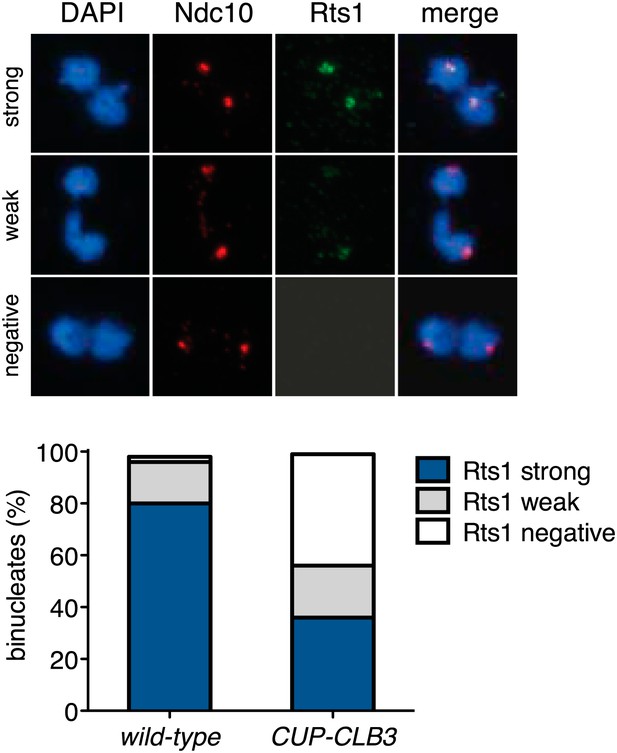
Rts1 localization in binucleate CUP-CLB3 cells.
Wild-type (A28329) and CUP-CLB3 (A28330) cells were induced to sporulate and CuSO4 (50 μM) was added 2 hr 30 min after transfer into sporulation medium. Rts1-13myc (green) localization relative to Ndc10-6HA (red) was determined in spread nuclei from binucleates (n > 40). Using a chi-square test (df 2) the fraction of spread nuclei that display strong, weak or negative Rts1 with respect to Ndc10 was compared between wild-type and CUP-CLB3 χ2 = 54.49, p<0.0001.
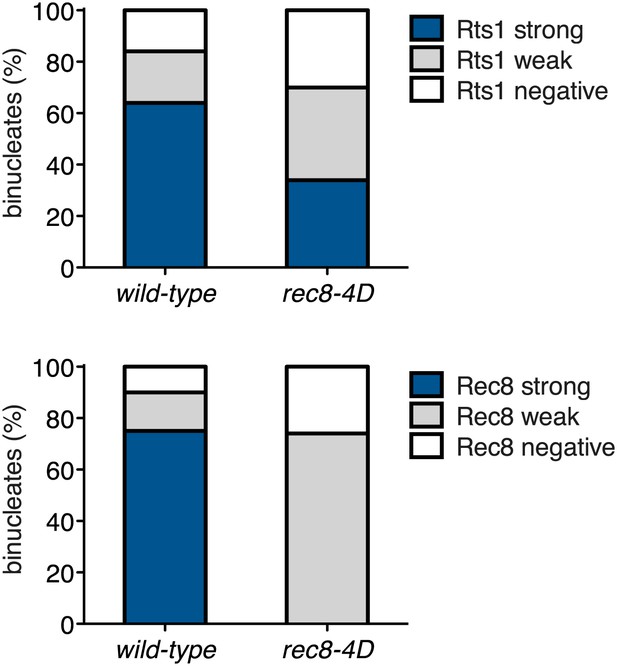
Analysis of Rts1 localization in Rec8 phosphomimetic mutants.
Wild-type (A29645) and rec8-S136D S179D S197D T209D (A29647) cells were induced to sporulate and Rec8-3HA/rec8-4D-3HA or Rts1-3V5 localization relative to Ndc10-13myc was determined in spread nuclei from binucleates (n > 40). Characterization of rec8-S136D S179D S197D T209D has been described in Katis et al. (2010). Note that strains carrying this allele fail to maintain centromeric cohesin beyond metaphase I (bottom panel). These binucleates also have weak Rts1 staining (top panel), suggesting that Rts1 maintenance at centromeric regions in anaphase I depends on cohesin. For top panel, using a chi-square test (df 2) the fraction of spread nuclei that display strong, weak or negative Rts1 with respect to Ndc10 was compared between wild-type and CUP-CLB3 χ2 = 18.02, p=0.0001. For bottom panel, using a chi-square test (df 2) the fraction of spread nuclei that display strong, weak or negative Rec8 with respect to Ndc10 was compared between wild-type and CUP-CLB3 χ2 = 121.2, p<0.0001.
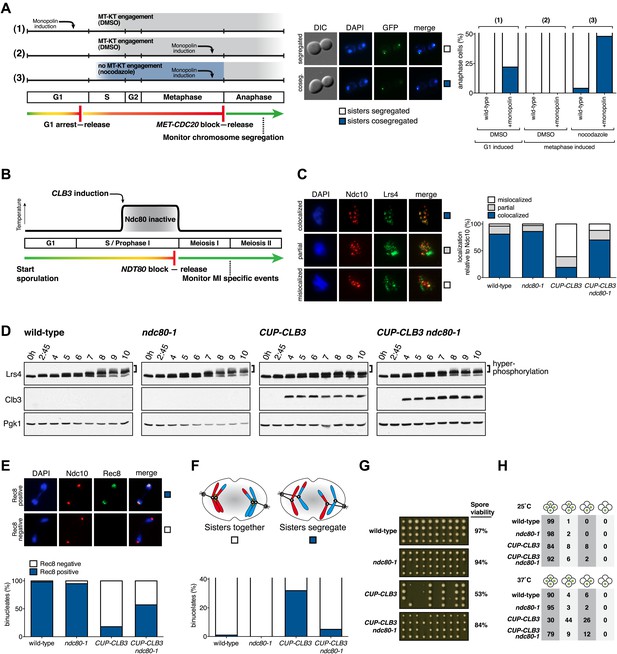
Transient disruption of microtubule–kinetochore interactions suppresses the chromosome segregation defects in CUP-CLB3 cells.
(A) Wild-type (A10684) and GAL-CDC5 GAL-MAM1 (A26546) cells, carrying a MET-CDC20 allele and CENIV-GFP dots, were monitored for chromosome segregation in anaphase (see ‘Materials and methods’ for details). MT = microtubule, KT = kinetochore, (n = 100). The fraction of anaphase cells that segregate or cosegregate sister chromatids was compared between GAL-CDC5 GAL-MAM1 condition (2) and GAL-CDC5 GAL-MAM1 condition (3) using a chi-square test (df 1) χ2 = 59.71, p<0.0001. (B) Schematic description of the experimental regime used for (C) through (H) see ‘Materials and methods’ for details. (C) Localization of Lrs4-13myc (green) in mononucleates relative to Ndc10-6HA (red) determined by nuclear spreads (n > 40) and (D) phosphorylation of Lrs4-13myc determined by gel mobility shift in wild-type (A29612), ndc80-1 (A29614), CUP-CLB3 (A29616) and CUP-CLB3 ndc80-1 (A29618). For (C), using a chi-square test (df 2) the fraction of spread nuclei that display colocalized, partial or mislocalized Lrs4 with respect to Ndc10 was compared between wild-type and ndc80-1 χ2 = 0.9668, p=0.6167 and between CUP-CLB3 and CUP-CLB3 ndc80-1 χ2 = 56.34, p<0.0001. (E) Localization of Rec8-13myc (green) in binucleates relative to Ndc10-6HA (red) determined by nuclear spreads in wild-type (A28716), ndc80-1 (A28720), CUP-CLB3 (A28718) and CUP-CLB3 ndc80-1 (A28722) (n > 40). Using a chi-square test (df 1) the fraction of spread nuclei that were Rec8 positive or negative was compared between wild-type and ndc80-1 χ2 = 1.185, p=0.2764 and between CUP-CLB3 and CUP-CLB3 ndc80-1 χ2 = 23.96, p<0.0001. (F) Segregation of sister chromatids using heterozygous CENV-GFP dots quantified in binucleates (n = 100) and (G) spore viability from wild-type (A22678), ndc80-1 (A28621), CUP-CLB3 (A22702) and CUP-CLB3 ndc80-1 (A28623) (n = 40 tetrads for wild-type and ndc80-1, n > 60 tetrads for CUP-CLB3 and CUP-CLB3 ndc80-1) (nonpermissive temperature >36°C). Using a chi-square test (df 1) the fraction of binucleates with a reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 ndc80-1 χ2 = 24.18, p<0.0001. (G) Segregation of chromosome V using homozygous CENV-GFP dots quantified in tetranucleates from wild-type (A22688), ndc80-1 (A28625), CUP-CLB3 (A22708) and CUP-CLB3 ndc80-1 (A28627). Top panel: cells kept at 25°C for the duration of the experiment. Bottom panel: Cells treated as in (B) but monitored after meiosis II (n = 100).
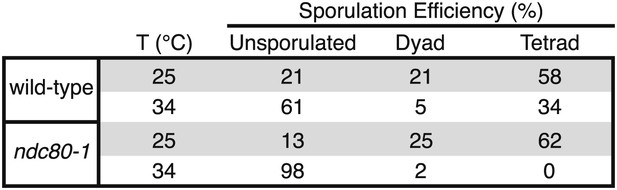
Sporulation efficiency of ndc80-1 mutants.
Wild-type (A22678) and ndc80-1 (A28221) cells were induced to sporulate at 25°C. 2 hr 30 min after transfer into sporulation medium, cells were shifted to the indicated temperature and sporulation efficiency was determined after 24 hr.
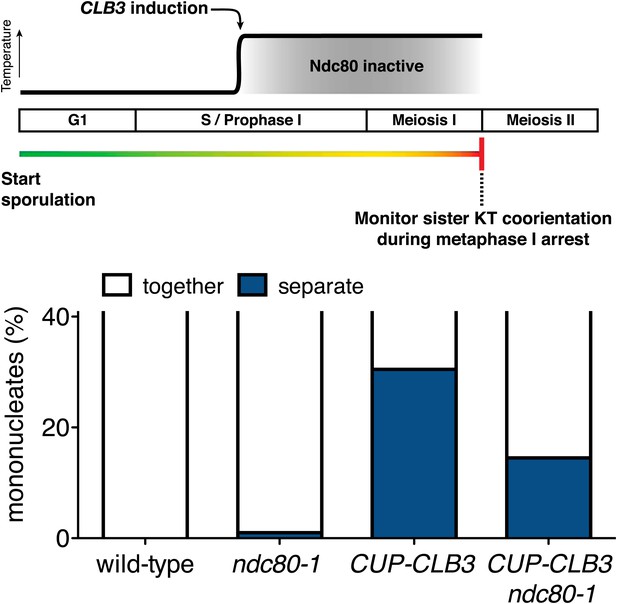
Sister kinetochore coorientation in ndc80-1 cells under a continuous inactivation regime at 34°C during a metaphase I arrest.
Wild-type (A7118), CUP-CLB3 (A23074), ndc80-1 (A29690) and ndc80-1 CUP-CLB3 (A29692) cells also carrying the cdc20-mn allele, were induced to sporulate at 25°C. 2 hr 45 min after transfer into sporulation medium, CuSO4 (50 μM) was added and concurrently, cultures were shifted to 34°C. The percentage of mononucleate cells with separated CENV-GFP dots was determined 7 hr 30 min after transfer into sporulation medium when cells were arrested in metaphase I (n = 100). The fraction of nuclei that display sister kinetochores as separate or together was compared between CUP-CLB3 and CUP-CLB3 ndc80-1 using a chi-square test (df 1) χ2 = 7.228, p=0.0072.
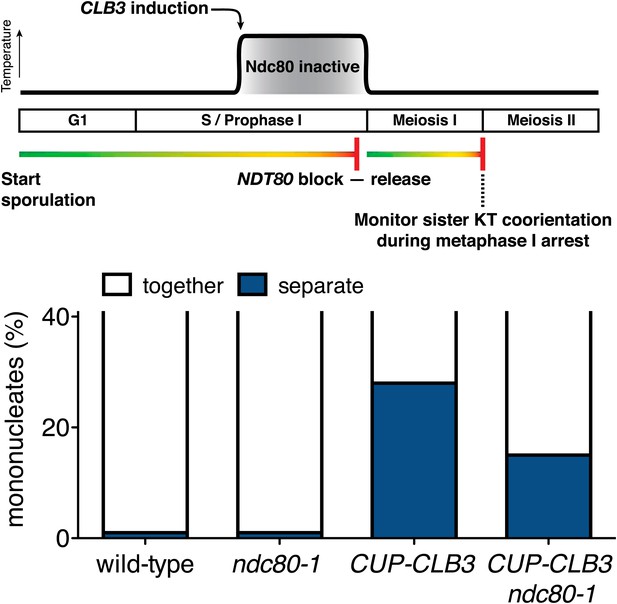
Sister kinetochore coorientation in ndc80-1 cells after transient inactivation regime at 34°C during a metaphase I arrest.
Wild-type (A20958), CUP-CLB3 (A23076), ndc80-1 (A29718) and ndc80-1 CUP-CLB3 (A29720) cells also carrying the GAL4-ER and GAL-NDT80 fusions and the cdc20-mn allele were induced to sporulate at 25°C. 2 hr 45 min after transfer into sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 34°C. After 5 hr, when cells had arrested in the NDT80 arrest, cells were released from the NDT80 block and transferred to 25°C. The percentage of mononucleate cells with separated CENV-GFP dots was determined 7 hr 30 min after transfer into sporulation medium when cells were arrested in metaphase I (n = 100). The fraction of nuclei that display sister kinetochores as separate or together was compared between CUP-CLB3 and CUP-CLB3 ndc80-1 using a chi-square test (df 1) χ2 = 5.007, p=0.0252.
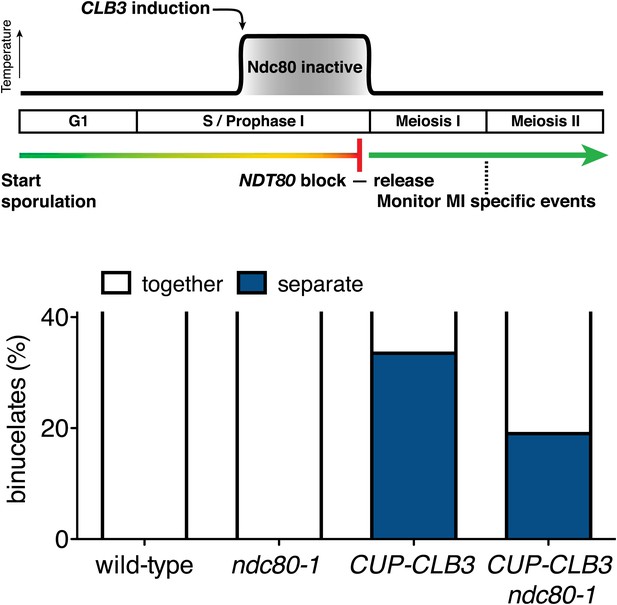
Meiosis I chromosome segregation in ndc80-1 cells after a transient inactivation regime at 34°C.
Wild-type (A22678), ndc80-1 (A28621), CUP-CLB3 (A22702) and CUP-CLB3 ndc80-1 (A28623) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer into sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 34°C. After 5 hr, when cells had arrested in the NDT80 block, cells were released and transferred to 25°C. The percentage of binucleate cells with segregated heterozygous CENV-GFP dots was determined 7 hr 30 min after transfer into sporulation medium (n = 100). Note that a greater suppression of meiosis I sister chromatid segregation was observed in ndc80-1 CUP-CLB3 cells when cells were incubated at temperatures higher than 34°C (Figure 5F and data not shown). The fraction of binucleates that underwent reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 ndc80-1 using a chi-square test (df 1) χ2 = 5.776, p=0.0162.
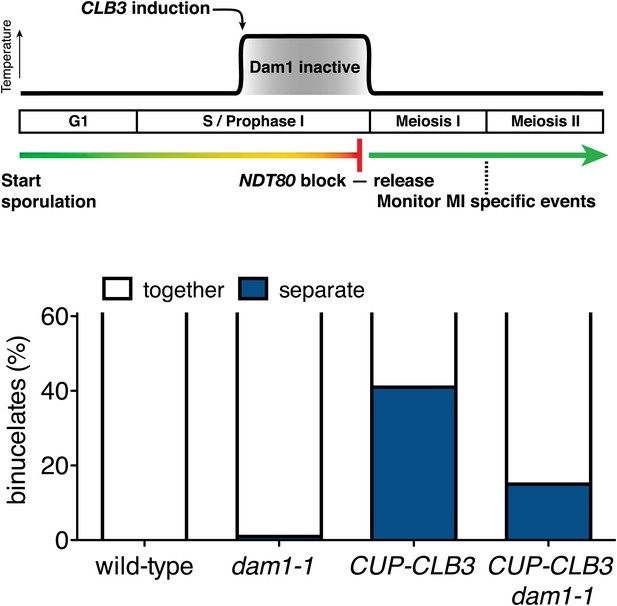
Transient disruption of microtubule–kinetochore interactions using dam1-1 allele restores meiosis I chromosome segregation in CUP-CLB3 cells.
Wild-type (A22678), dam1-1 (A28311), CUP-CLB3 (A22702) and CUP-CLB3 dam1-1 (A28341) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer into sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 34°C. After 5 hr, when cells had arrested in the NDT80 block, cells were released and transferred to 25°C. The percentage of binucleate cells with segregated heterozygous CENV-GFP dots was determined 7 hr 30 min after transfer into sporulation medium (n = 100). The fraction of binucleates that underwent reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 dam1-1 using a chi-square test (df 1) χ2 = 16.77, p<0.0001.
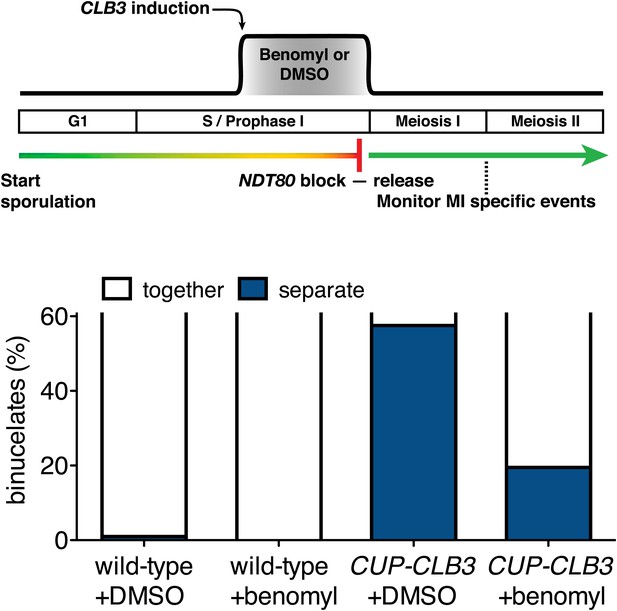
Transient disruption of microtubule–kinetochore interactions by benomyl treatment restores meiosis I chromosome segregation in CUP-CLB3 cells.
Wild-type (A22678) and CUP-CLB3 (A22702) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 30°C. 2 hr 15 min after transfer into sporulation medium CuSO4 (50 μM) was added and concurrently, cells were treated with DMSO or benomyl (120 μg/ml). Cells were subsequently released from NDT80 block 4 hr 30 min after transfer into sporulation medium and benomyl was washed out concomitant with NDT80-block release. The percentage of binucleate cells with segregated heterozygous CENV-GFP dots was determined 6 hr after transfer into sporulation medium (n = 100). See ‘Materials and methods’ for further details. The fraction of binucleates that underwent reductional or equational division was compared between CUP-CLB3 + DMSO and CUP-CLB3 + benomyl using a chi-square test (df 1) χ2 = 32.12, p<0.0001.
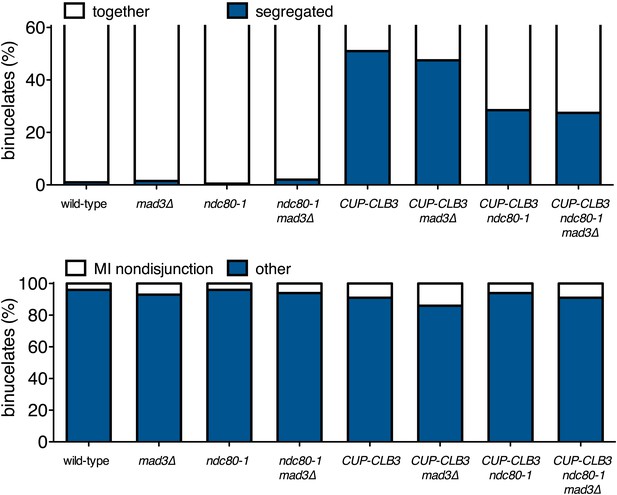
Transient disruption of microtubule–kinetochore interactions during S phase/prophase I suppresses CUP-CLB3-induced meiosis I sister chromatid segregation in a spindle assembly checkpoint independent manner.
(Top panel) wild-type (A22678), mad3∆ (A30386), ndc80-1 (A28621), ndc80-1 mad3∆ (A30390), CUP-CLB3 (A22702), CUP-CLB3 mad3∆ (A30388), CUP-CLB3 ndc80-1 (A28623) and CUP-CLB3 ndc80-1 mad3∆ (A30392) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer into sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 36°C. Cells were subsequently released from NDT80 block at 5 hr and transferred to 25°C. Percent binucleates with segregated heterozygous CENV-GFP dots was determined (n = 100). (Bottom panel) wild-type (A22688), mad3∆ (A30638), ndc80-1 (A28625), ndc80-1 mad3∆ (A30642), CUP-CLB3 (A22708), CUP-CLB3 mad3∆ (A30640), CUP-CLB3 ndc80-1 (A28627) and CUP-CLB3 ndc80-1 mad3∆ (A30644) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer to sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 36°C. After 5 hr, when cells had arrested in the NDT80 block, cells were released and transferred to 25°C. The percentage of binucleate cells with segregated homozygous CENV-GFP dots was determined 7 hr 30 min after transfer into sporulation medium. Binucleate cells with GFP signal in only one of the two nuclei were categorized as having experienced a meiosis I non-disjunction event (n = 100). For top panel, using a chi-square test (df 1) the fraction of binucleates that underwent reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 mad3∆ χ2 = 0.1800, p=0.6714 and between CUP-CLB3 ndc80-1 and CUP-CLB3 ndc80-1 mad3∆ χ2 = 0.02454, p=0.8755. For bottom panel, using a chi-square test (df 1) the fraction of binucleates that displayed MI nondisjunction or other was compared between CUP-CLB3 and CUP-CLB3 mad3∆ χ2 = 1.228, p=0.2678 and between CUP-CLB3 ndc80-1 and CUP-CLB3 ndc80-1 mad3∆ χ2 = 0.6486, p=0.4206.
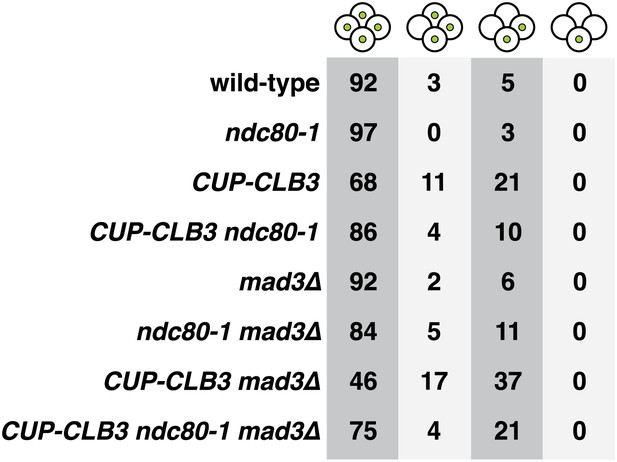
Transient disruption of microtubule–kinetochore interactions during S phase/prophase I restores meiotic chromosome segregation in CUP-CLB3 cells in a spindle assembly checkpoint independent manner.
Wild-type (A22688), ndc80-1 (A28625), CUP-CLB3 (A22708), CUP-CLB3 ndc80-1 (A28627), mad3∆ (A30638), ndc80-1 mad3∆ (A30642), CUP-CLB3 mad3∆ (A30640) and CUP-CLB3 ndc80-1 mad3∆ (A30644) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer to sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 36°C. After 5 hr, when cells had arrested in the NDT80 block, cells were released and transferred to 25°C. Segregation of homozygous CENV-GFP dots was determined in tetranucleates 12 hr after transfer into sporulation medium (n = 100).
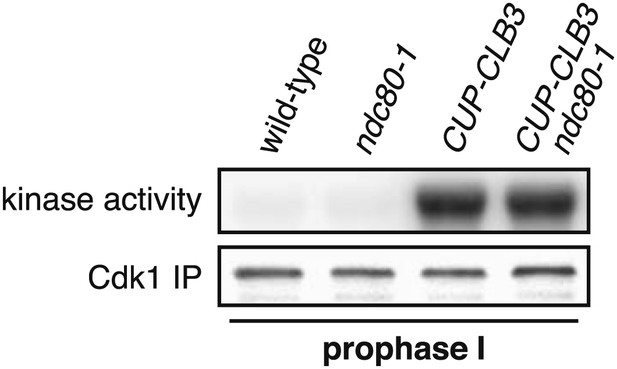
Transient ndc80-1 inactivation does not alter in vitro Cdk1 activity.
Wild-type (A25508), ndc80-1 (A33203), CUP-CLB3 (A33201) and CUP-CLB3 ndc80-1 (A33205) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate at 25°C. 2 hr 45 min after transfer to sporulation medium CuSO4 (50 μM) was added and concurrently, cultures were shifted to 35°C. Samples were harvested 5 hr post transfer to sporulation medium, when cells were arrested in the NDT80 block. In vitro kinase assays were performed with Cdc28-3V5 (Cdk1) immunoprecipitated from prophase I-arrested samples. Amounts of phosphorylated Histone H1 and immunoprecipitated Cdc28-3V5 are shown.
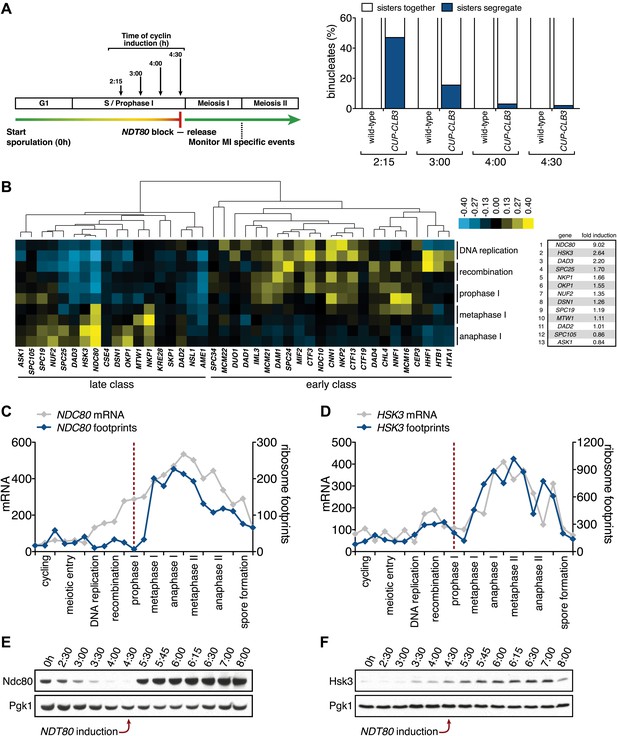
Meiosis I sister chromatid segregation correlates with presence of outer kinetochore components.
(A) Schematic description of the experimental regime and segregation of sister chromatids using heterozygous CENV-GFP dots quantified in binucleates from wild-type (A22678) and CUP-CLB3 (A29406) after cyclin induction at 2 hr 15 min, 3 hr, 4 hr and 4 hr 30 min post transfer to sporulation medium. Cells released from NDT80-block at 4 hr 30 min (n = 100). Using a chi-square test (df 1), the fraction of binucleates that display a reductional or equational division was compared between wild-type and CUP-CLB3 for each induction time point: (2:15), χ2 = 58.00, p<0.0001; (3:00), χ2 = 14.46, p=0.0001; (4:00), χ2 = 1.020, p=0.3124; (4:30), χ2 = 0.3384, p=0.5607. (B) Cluster analysis of kinetochore components from the indicated time points. Further details are in the ‘Materials and methods’ and in Brar et al. (2012). Inner kinetochore = Cse4 nucleosomes, Cbf3, Ctf19 complexes and Mif2. Outer kinetochore = Spc105, Mis12, Ndc80 and DASH complexes. Fold induction is calculated by dividing the average expression from metaphase I—anaphase I by the average expression from DNA replication-prophase I. (C) Ordered plot for mRNA-seq and ribosome footprinting data for NDC80 and (D) HSK3 at the indicated stages. Dotted line indicates time of release from NDT80 block. Further details are in the ‘Materials and methods’ and in Brar et al. (2012). (E) Western blot for Ndc80-3V5 and Pgk1 from A30340 cells and (F) Hsk3-3V5 and Pgk1 from A31861 cells. Cells induced to sporulate and released from NDT80 block at 4 hr 30 min.
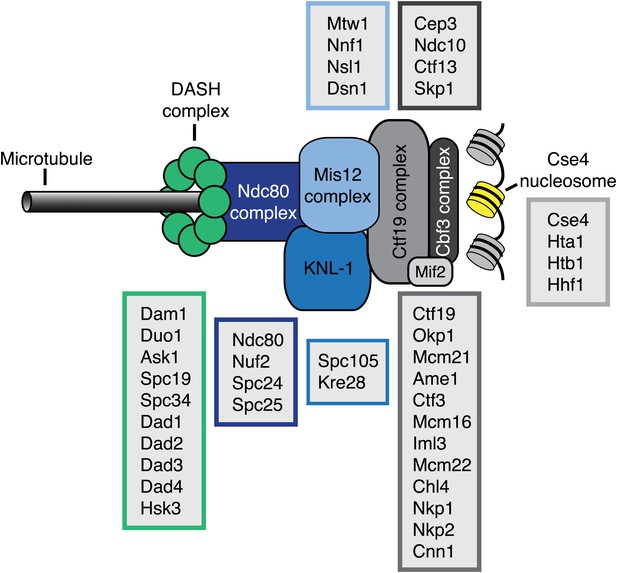
Schematic representation of the kinetochore–microtubule interface.
Components of the kinetochore subcomplexes are grouped in color coded boxes. Note that the schematic representation is not drawn to scale.
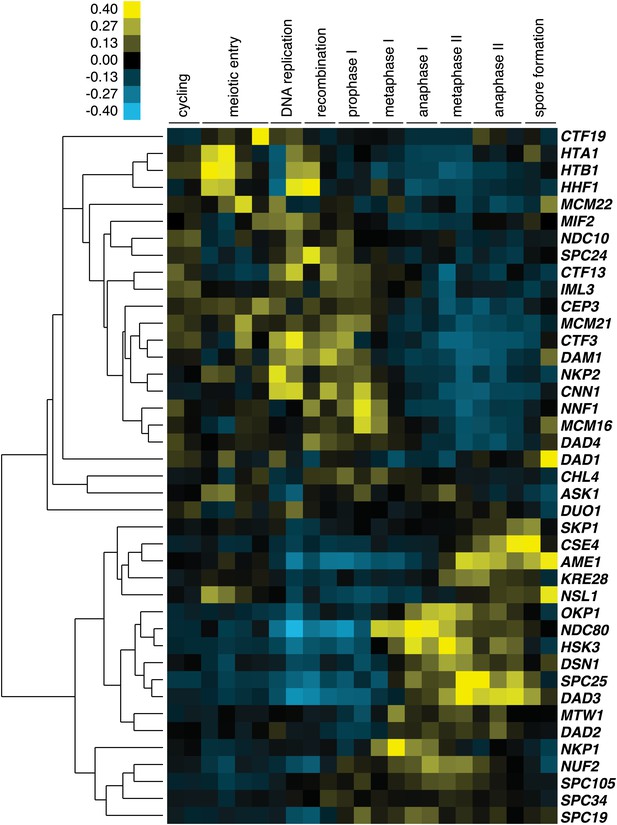
Meiotic cluster analysis of kinetochore components.
Cluster analysis of kinetochore components from the indicated time points. Further details are in ‘Materials and methods’ and in Brar et al. (2012).
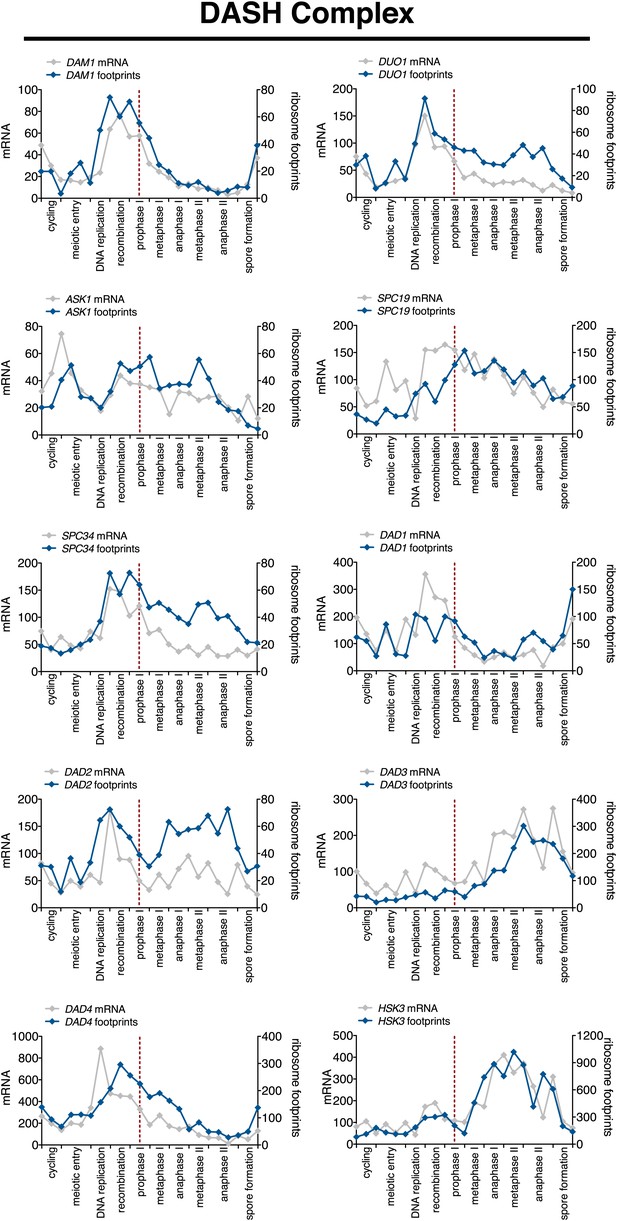
Meiotic expression of DASH complex subunits.
Ordered plot of mRNA-seq and ribosome footprinting data for the DASH complex components at indicated stages of sporulation. Dotted line indicates time of release from NDT80 block. See Brar et al. (2012) for details.
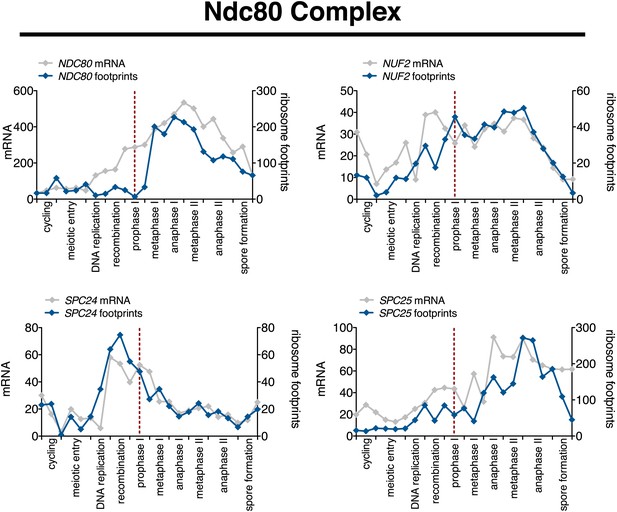
Meiotic expression of Ndc80 complex subunits.
Ordered plot of mRNA-seq and ribosome footprinting data for the Ndc80 complex at indicated stages. Dotted line indicates time of release from NDT80 block. See Brar et al. (2012) for details.
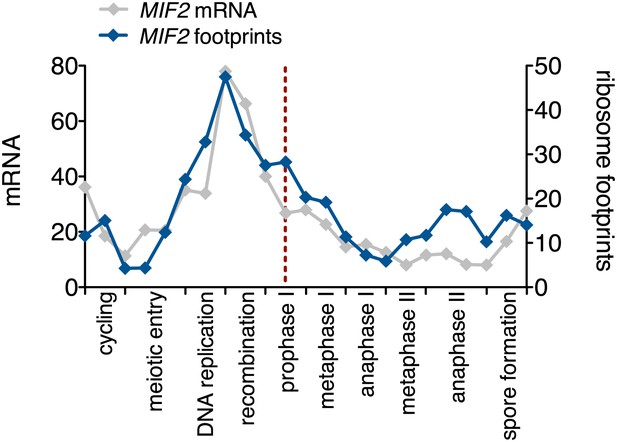
Meiotic expression of Mif2.
Ordered plot of the mRNA-seq and ribosome footprinting data for Mif2 at the indicated stages. Dotted line indicates time of release from the NDT80 block. See Brar et al. (2012) for details.
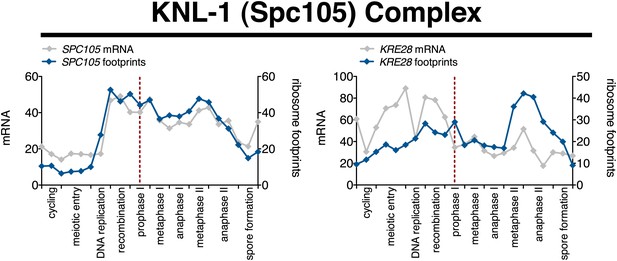
Meiotic expression of KNL-1 complex subunits.
Ordered plot of the mRNA-seq and ribosome footprinting data for KNL-1 complex subunits (Spc105 complex) at the indicated stages. Dotted line indicates time of release from the NDT80 block. See Brar et al. (2012) for details.
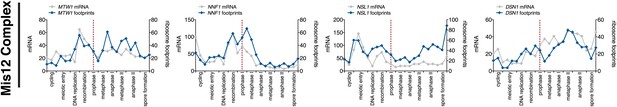

Meiotic expression of Mis12 complex subunits.
Ordered plot of the mRNA-seq and ribosome footprinting data for Mis12 complex subunits at the indicated stages. Dotted line indicates time of release from NDT80 block. See Brar et al. (2012) for details.
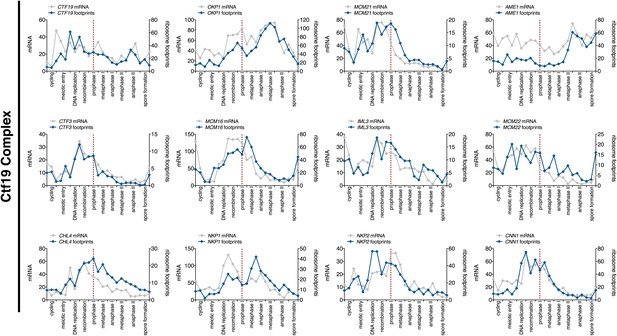
Meiotic expression of Ctf19 complex subunits.
Ordered plot for mRNA-seq and ribosome footprinting data for Ctf19 complex at indicated stages. Dotted line indicates time of release from the NDT80 block. See Brar et al. (2012) for details.
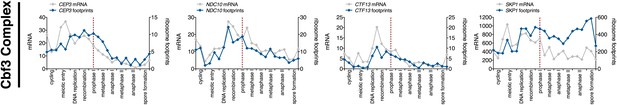
Meiotic expression of Cbf3 complex subunits.
Ordered plot of the mRNA-seq and ribosome footprinting data for Cbf3 complex subunits at the indicated stages. Dotted line indicates time of release from the NDT80 block. See Brar et al. (2012) for details.
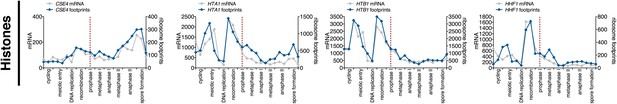

Meiotic expression of Histone subunits.
Ordered plot for mRNA-seq and ribosome footprinting data for the histones at indicated stages. Dotted line indicates time of release from NDT80 block See (Brar et al., 2012) for details.
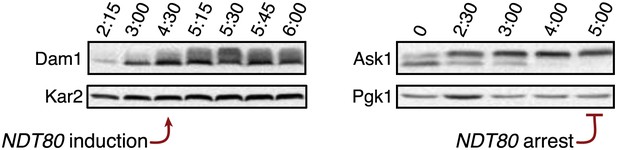
Meiotic expression of Dam1 and Ask1.
DAM1-3V5 (A28898) cells carrying the GAL4-ER and GAL-NDT80 fusion and ASK1-13myc (A29161) carrying ndt80∆ were induced to sporulate and were either released from (left panel) or arrested in (right panel) the NDT80 block. Levels of Dam1-3V5, Ask1-13myc, Kar2 and Pgk1 were monitored by Western blot analysis. Pgk1 and Kar2 served as loading controls.
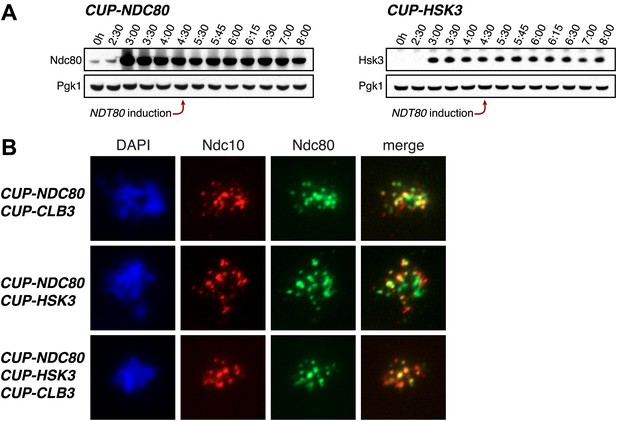
Characterization of NDC80 and HSK3 overexpression.
(A) CUP-NDC80-3V5 (A30342) and CUP-HSK3-3HA (A32060) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate. After 2 hr 30 min CuSO4 (50 μM) was added and cells were subsequently released from NDT80 block 4 hr 30 min after transfer into sporulation medium. The levels of Ndc80-3V5, Hsk3-3HA and Pgk1 were monitored by Western blot. (B) CUP-NDC80-3V5 CUP-CLB3 (A31949), CUP-NDC80-3V5 CUP-HSK3 (A31951) and CUP-NDC80-3V5 CUP-HSK3 CUP-CLB3 (A31953) cells were induced to sporulate. 4 hr after transfer into sporulation medium CuSO4 (50 μM) was added, and localization of Ndc80-3V5 (green) relative to Ndc10-6HA (red) was determined by nuclear spreads 5 hr after transfer into sporulation medium.
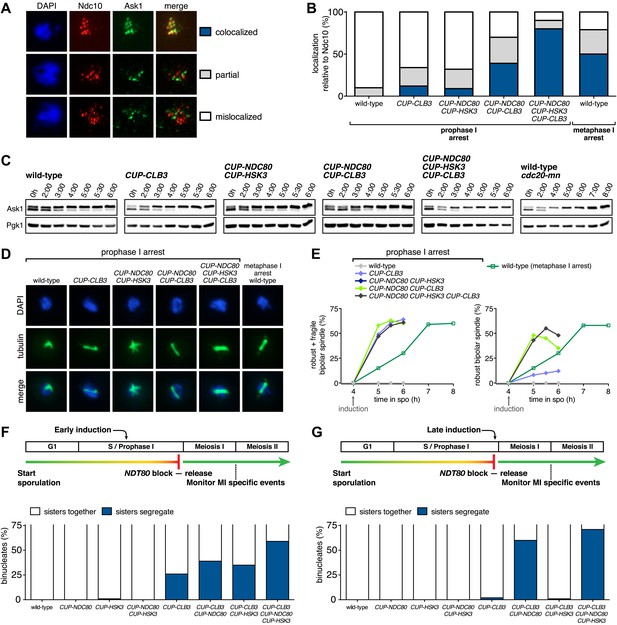
Expression of NDC80 and HSK3 in prophase I enhances Clb3-CDK-induced meiosis I sister chromatid segregation.
For (A)–(E), wild-type (A31945), CUP-CLB3 (A31947), CUP-NDC80 CUP-HSK3 (A31951), CUP-NDC80 CUP-CLB3 (A31949), CUP-NDC80 CUP-HSK3 CUP-CLB3 (A31953) and cdc20-mn (A31955) cells were induced to sporulate and CuSO4 (50 μM) was added at 4 hr after sporulation induction. (A) Representative images and (B) quantification of Ask1-13myc (green) in mononucleates relative to Ndc10-6HA (red) determined by nuclear spreads prepared after 1 hr of CuSO4 induction (n > 40 except for A31955 [n = 28]). For (B), using a chi-square test (df 2) the fraction of spread nuclei that display colocalized, partial or mislocalized Ask1 with respect to Ndc10 was compared between CUP-CLB3 and CUP-CLB3 CUP-NDC80 CUP-HSK3 χ2 = 51.49, p<0.0001. (C) Western blots of Ask1-13myc and Pgk1. (D) Bipolar spindle morphology and (E) left panel, total (robust + fragile) bipolar spindle formation, and right panel, robust bipolar spindle formation determined at the indicated time points (see ‘Materials and methods’ for further description) (n = 100 per time point). Note: CUP-NDC80 CUP-HSK3 (dark blue) data points occluded by wild-type (grey) data points. (F), (G) Segregation of sister chromatids using heterozygous CENV-GFP dots quantified in binucleates from wild-type (A30340), CUP-NDC80 (A30342), CUP-HSK3 (A31849), CUP-NDC80 CUP-HSK3 (A31855), CUP-CLB3 (A31847), CUP-CLB3 CUP-NDC80 (A31853), CUP-CLB3 CUP-HSK3 (A31851) and CUP-CLB3 CUP-NDC80 CUP-HSK3 (A31857) (early induction = 2:15 hr, late induction= 4:30 hr after induction of sporulation; release from NDT80 block at 4:30 hr) (n = 100). For (F), using a chi-square test (df 1) the fraction of binucleates with a reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 CUP-NDC80 CUP-HSK3 χ2 = 22.28, p<0.0001. For (G), using a chi-square test (df 1) the fraction of binucleates with a reductional or equational division was compared between CUP-CLB3 and CUP-CLB3 CUP-NDC80 CUP-HSK3 χ2 = 102.7, p<0.0001.
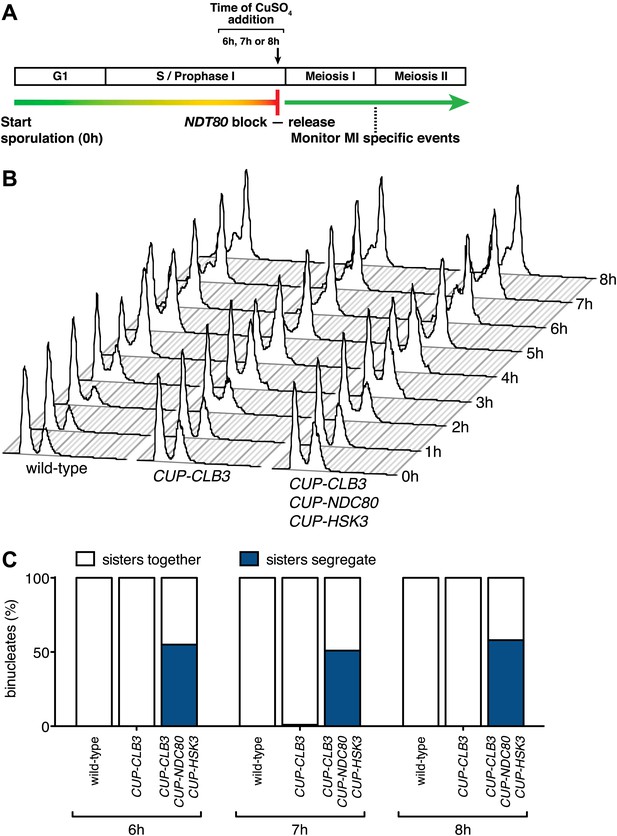
CUP-CLB3 CUP-NDC80 CUP-HSK3-induced meiosis I sister chromatid segregation is independent of the length of the prophase I arrest.
(A) Schematic description of the experimental regime used in (B) and (C). (B), (C) Wild-type (A22678), CUP-CLB3 (A22702) and CUP-CLB3 CUP-NDC80 CUP-HSK3 (A31857) cells also carrying the GAL4-ER and GAL-NDT80 fusions were induced to sporulate. Cells were released from the NDT80 block and concurrently pCUP1-dependent expression was induced at either 6 hr, 7 hr or 8 hr post transfer to sporulation medium (by addition of 1 μM estradiol and 50 μM CuSO4 respectively). Samples were taken at the indicated time points to determine DNA content (B) and the percentage of binucleate cells with segregated sister chromatids (C). For (C), using a chi-square test (df 1), the fraction of binucleates that display a reductional or equational division in CUP-CLB3 CUP-NDC80 CUP-HSK3 cells was compared between 6 hr and 7 hr induction χ2 = 0.3212, p=0.5709 and between 6 hr and 8 hr induction χ2 = 0.1831, p=0.6687.
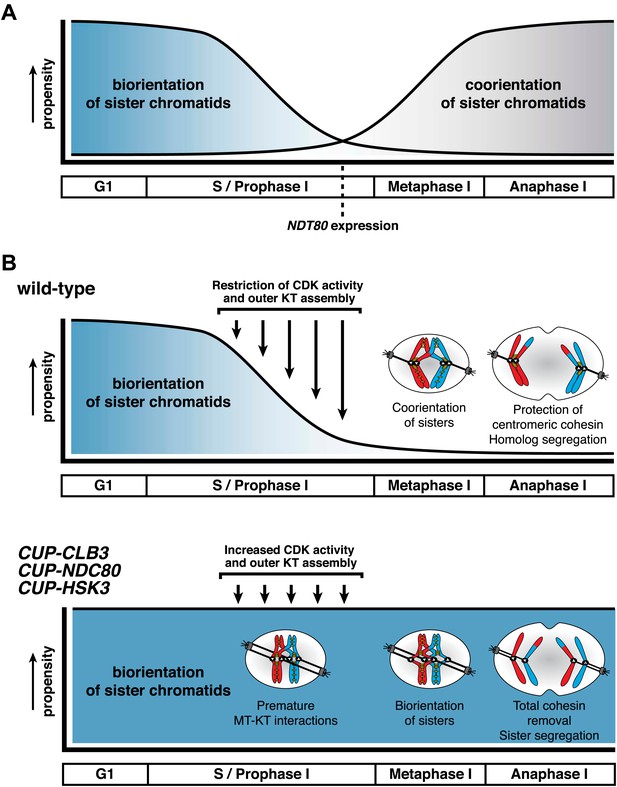
Model for temporal regulation of microtubule–kinetochore interactions during meiosis.
(A) As prophase I progresses, the propensity of sister chromatids to biorient decreases and the ability to coorient sister chromatids increases. (B) Top panel: inhibiting Clb-CDK activity and outer kinetochore (KT) assembly during prophase I establishes a meiosis I-specific chromosome segregation pattern by allowing sister kinetochore coorientation and protection of centromeric cohesin. Bottom panel: disrupting the regulation of microtubule–kinetochore (MT–KT) interactions causes sister chromatid segregation in meiosis I.
Additional files
-
Supplementary file 1
Strains used in this study.
- https://doi.org/10.7554/eLife.00117.055





