Origins and functional consequences of somatic mitochondrial DNA mutations in human cancer
Figures
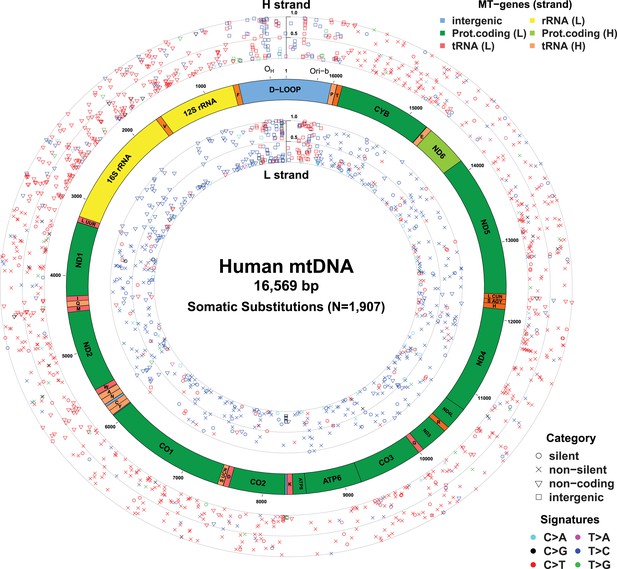
Mitochondrial somatic substitutions identified from 1675 Tumor–Normal pairs.
mtDNA genes and intergenic regions are shown. The strand of genes is shown based on mtDNA strand containing equivalent sequences of transcribed RNA. Substitution categories (silent, non-silent (missense and nonsense), non-coding (tRNA and rRNA), and intergenic) are shown by the shapes of each substitution. Six classes of substitutions are presented color-coded. The substitutions on the H, and L strand (when six substitutional classes were considered) are shown outside and inside of mtDNA genes, respectively. Vertical axes for H and L strand substitutions represent the VAF of each variant.
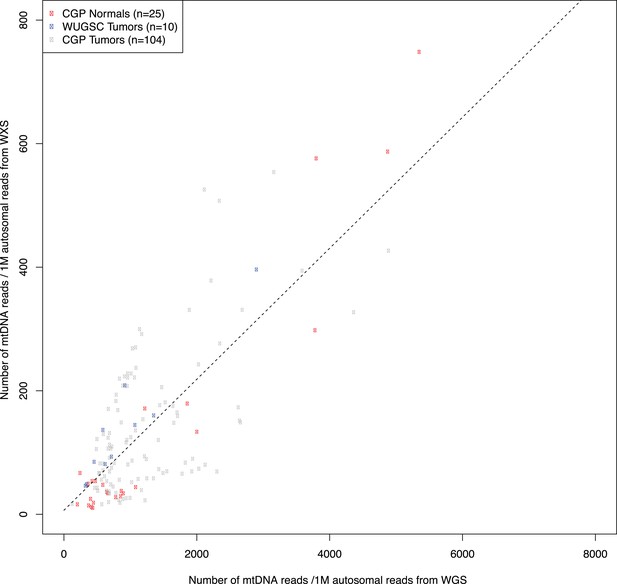
Correlation in amount of mtDNA reads between whole-genome and whole-exome sequencing.
139 DNA samples, either from tumors or bloods, sequenced by whole-genome sequencing were additionally sequenced by whole-exome sequencing. We compared the amount of mtDNA reads between whole-genome and whole-exome sequencing. As shown in this figure, we found strong positive correlation. * CGP; Cancer Genome Project, Wellcome Trust Sanger Institute, WUGSC; Washington University Genome Sequencing Center.
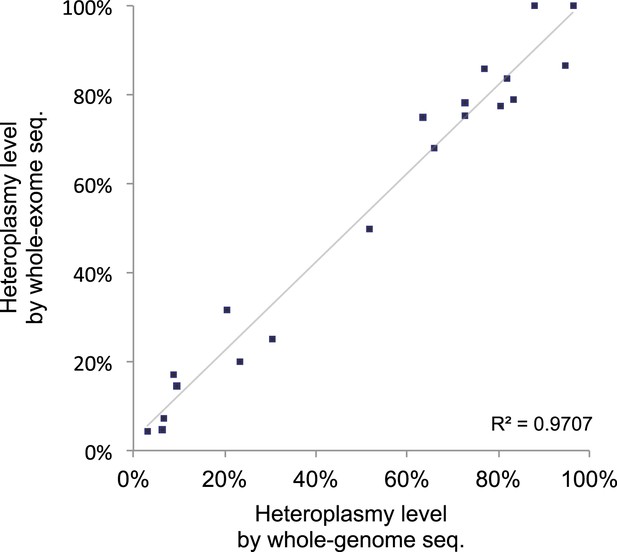
Correlation of heteroplasmy levels between whole-genome and whole-exome sequencing.
To validate the sensitivity and specificity of variant calling in this study, 19 tumor and normal pairs (which were originally whole-genome sequenced) were whole-exome sequenced and mtDNA variants were assessed independently. We correlated the heteroplasmic levels of 20 mutations detected in common.
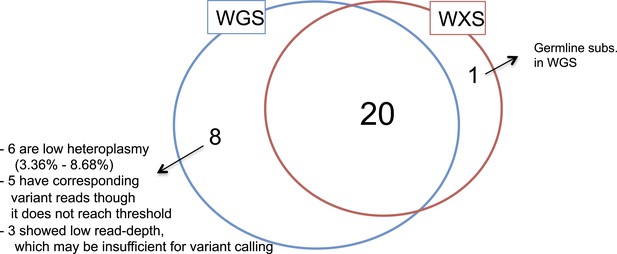
Validation of mtDNA somatic substitutions.
https://doi.org/10.7554/eLife.02935.007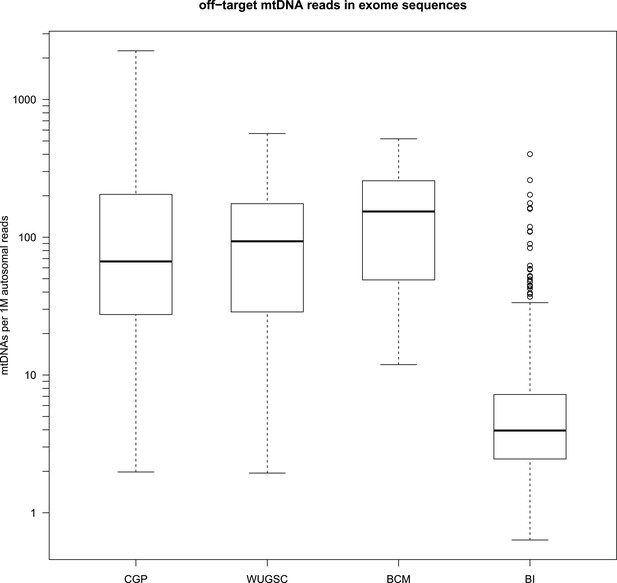
Amount of off-target mtDNA reads across four sequencing centers.
* CGP; Cancer Genome Project, Wellcome Trust Sanger Institute (n = 855), WUGSC; Washington University Genome Sequencing Center (n = 140), BCM; Baylor College of Medicine (n = 85), BI; Broad Institute (n = 435).
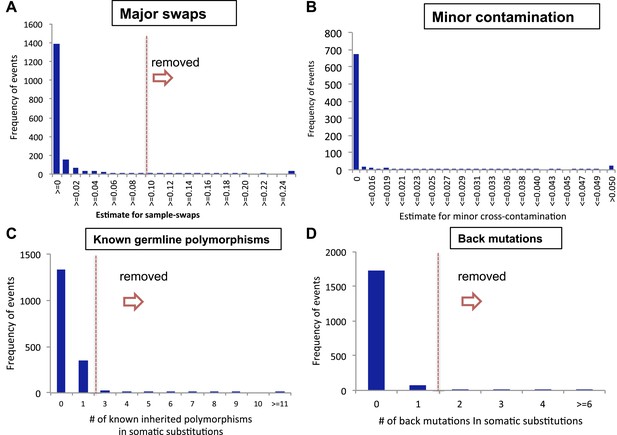
Filtering samples of potential DNA contaminations.
(A) A histogram presenting potential sample swaps in tumor–sample pairs. (B) A histogram presenting potential minor DNA cross-contamination in tumor samples. Cross-contamination levels were considered in filtering substitutions (see “Minor cross-contamination of DNA samples” section in Materials and Methods). (C) Histograms showing number of somatic substitutions overlapping with known inherited polymorphisms and (D) number of back mutations.
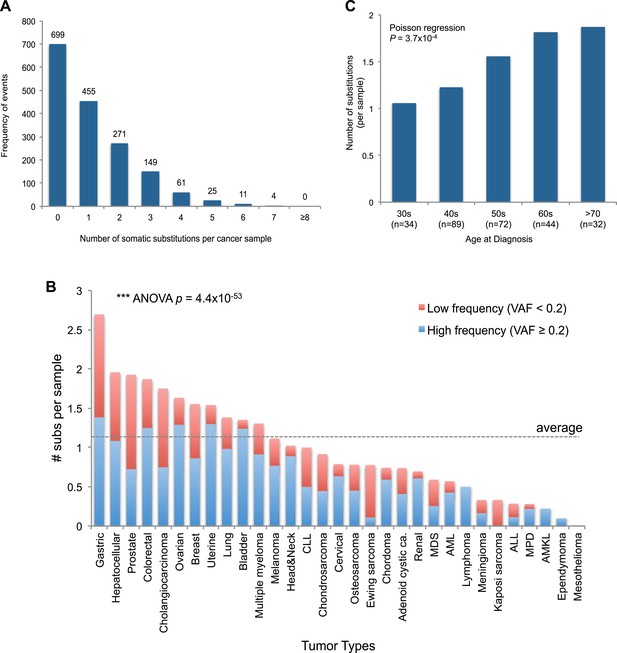
mtDNA somatic substitutions of human cancer.
(A) Number of somatic substitutions in a tumor sample. (B) Average number of somatic substitutions per sample across 31 tumor types. (C) Age of diagnosis and number of mtDNA somatic substitutions in breast cancers.
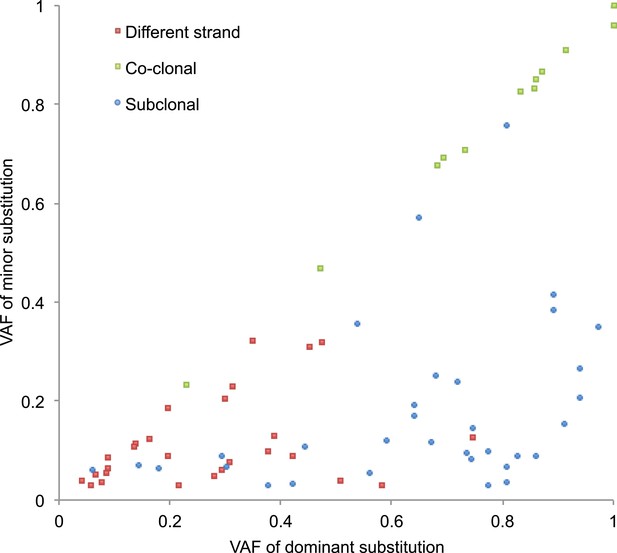
VAFs of phased somatic mtDNA substitutions.
This figure presents VAF pairs between co-clonal, sub-clonal, and different strand mtDNA substitutions. We expect similar VAFs for co-clonal pairs; lower VAF in sub-clonal mutations compared to clonal ones; and sum of a VAF pair is equal or less than 1.0.
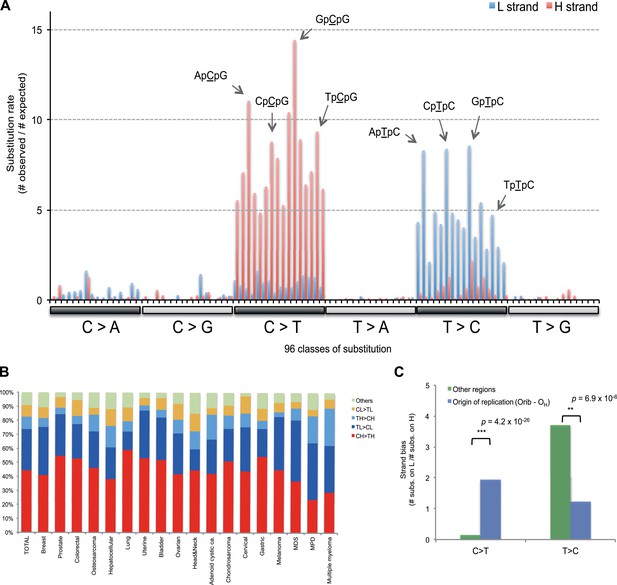
Replicative strand bias for mtDNA somatic substitutions.
(A) Replicative strand-specific substitution rate (# of observed/# of expected) by 96 trinucleotide context. Substitutions in a specific mtDNA segment (from Ori-b to OH) are not included, because they present a different substitutional signature. (B) Mutational signature across tumor types. Eighteen tumor types, which include at least 25 mtDNA mutations, were shown. (C) Inverted substitution signature in the Ori-b–OH.
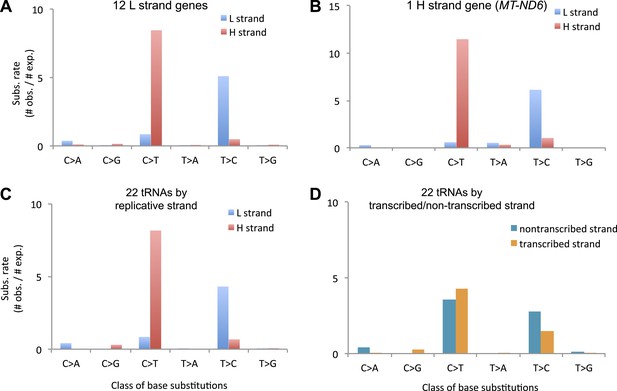
Replicative strand bias observed in mtDNA substitutions.
(A) Mutational signature of mtDNA somatic substitutions on the 12 L strand genes by replicative strand (L/H strand). It agrees very well with the background mutational signature. (Chi-square p = 0.99999). (B) Mutational signature of mtDNA somatic substitutions on the H strand gene (MT-ND6) by replicative strand. It is very close to the background very close to the expected background signature (Chi-square p = 0.027). If we consider signature by transcriptional strand, the signature difference is very clear (Chi-square p = 1 × 10−21). These suggest the strand bias not to be transcription-coupled, but replication coupled. (C) Mutational spectrum of mtDNA somatic substitutions on the 22 tRNA genes by replicative strand. Again, it agrees very well with the background mutational signature (Chi-square p = 0.71). (D) Mutational spectrum of mtDNA somatic substitutions on the 22 tRNA genes by non-transcribed (coding) and transcribed (non-coding) strand. Strand bias was greatly subsided because somatic substitutions on 14 L strand and 8 H strand tRNAs neutralize the strand bias (CH > TH and TL > CL) each other. As a result, this signature of tRNA mutations by transcriptional strand is significantly different from the background one (Chi-square p = 3.3 × 10−12). Taken all together, we concluded that the cause of strand bias is not transcription-coupled but is replicative.
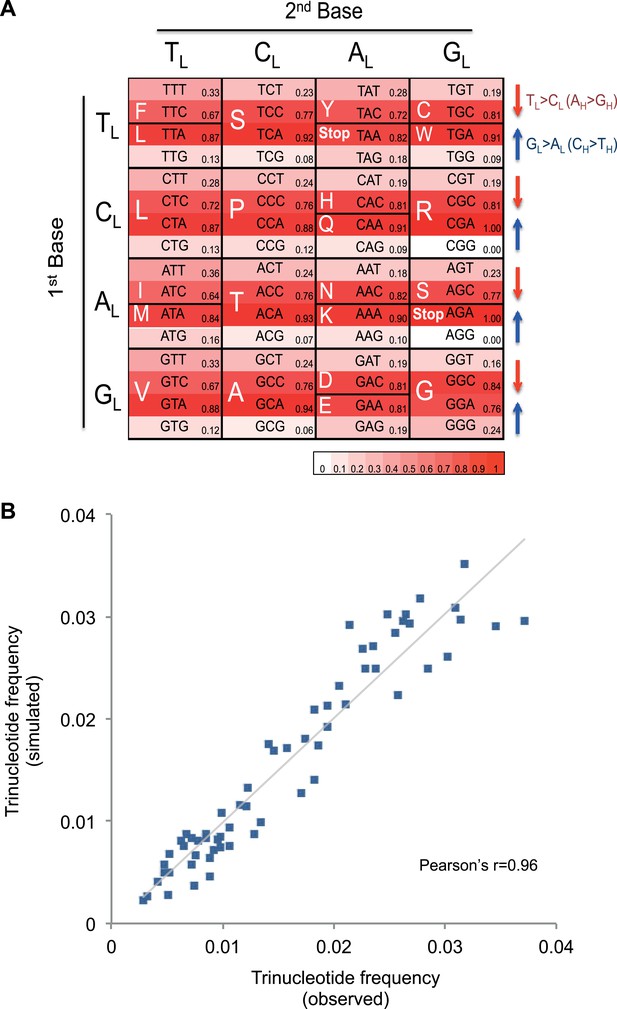
Mutational signature similar to processes shaping human mtDNA sequence over evolutionary time.
(A) Triplet codon depletion in human mtDNA by equivalent (CH > TH and TL > CL) mutational pressure. Relative frequency of each triplet codon within synonymous pairs (NNT–NNC or NNA–NNG) is shown by color. The arrows beside the box highlight the T > C (red) and G > A (blue) substitutional pressures on the L strand in germline mtDNA. (B) Correlation of triplet codon frequencies between from observed and from simulated evolutions of a random sequence mtDNA by the mtDNA somatic mutational signature with constraining mitochondrial protein sequences.
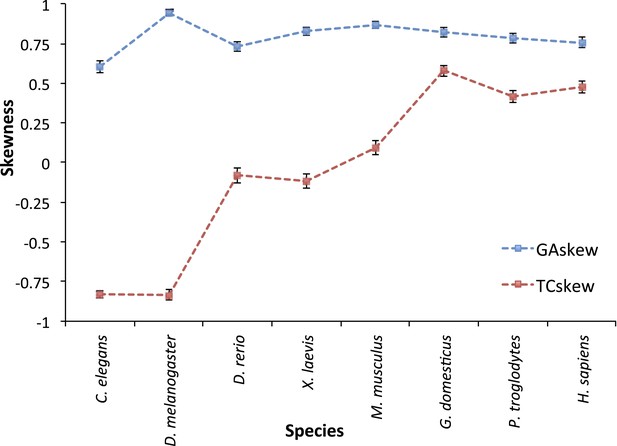
TC and GA skew for L strand mtDNA genes across 8 animal species.
C. elegans (a nematode) and D. melanogaster (fruit fly) mtDNA appears to have GL << AL (due to CH > TH mutational pressure) and CL >> TL (due to CL > TL mutational pressure) in the third base of triplet codon in L strand genes. Therefore they seem to have predominant C > T mutational pressure without strand bias. D. rerio (zebrafish), X. laevis (frog), and M. musculus (mouse) present GL << AL (due to CH > TH mutational pressure), but similar number of CL and TL. Therefore, mtDNA of these sequences is thought to have CH > TH, with strand bias. The existence of TL > CL is not clear. Finally, mtDNA of H. sapiens, P. troglodytes (Chimpanzee), and G. domesticus (Chicken) shows clear CH > TH and TL > CL as mentioned in the main manuscript. Interestingly, TL > CL seems to be slightly stronger in the mitochondria of chicken than that of human (or chimp). We suggest there would be some differences in the mechanism of mtDNA replication across the evolution tree.
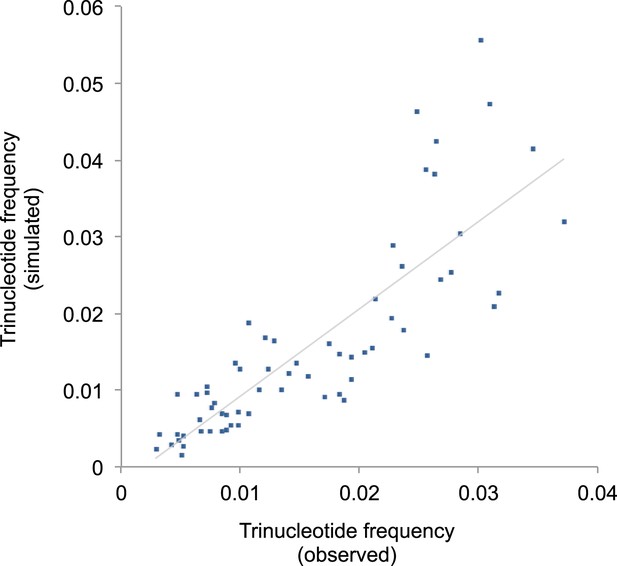
Correlation of triplet codon frequencies between from observed and from simulated evolutions under the mtDNA somatic mutational signature.
https://doi.org/10.7554/eLife.02935.016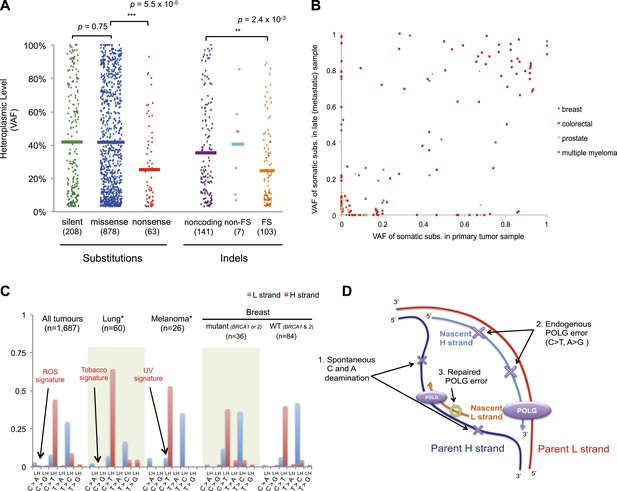
Selection and mutational process for mtDNA somatic substitutions.
(A) Truncating mutations (nonsense substitutions and frame-shifting (FS) coding indels) present significantly lower VAF. (B) Change of VAF of mtDNA somatic mutation between primary and metastatic (or late) cancer tissues. (C) Mutational signature for mtDNA across various tumor types. None of the three highlighted mechanisms or nuclear DNA double-strand breaks repair mechanism (BRCA) match with the mtDNA mutational signature. * Only substitutions in protein-coding genes considered. (D) A proposed model of mtDNA mutational process.
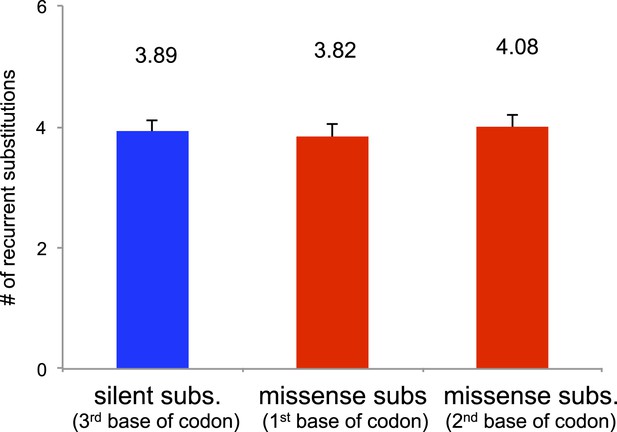
Number of recurrent substitutions between silent and missense substitutions.
100 sites were randomly selected from silent substitutions (at third base of triplet codon) and missense substitutions (at first and second base of triplet codon). No significant difference was observed among these three groups.
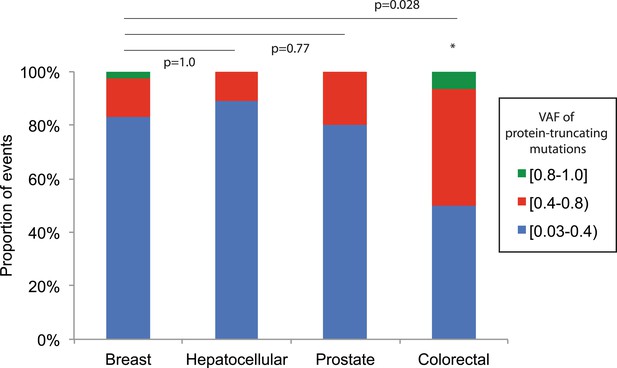
Comparison of VAF of protein-truncating mutations (nonsense substitution and indels) across tumor types.
Four tumor types with more than 10 protein-truncating mutations are shown. Fisher's exact were applied between breast and other tissue types.
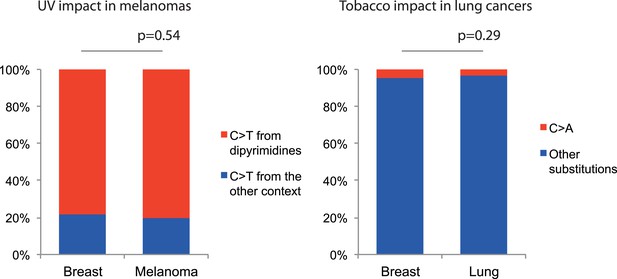
Negligible impacts of external mutagens (UV and tobacco smoking) to the somatic mtDNA mutations.
No evidence of UV and tobacco smoking was identified even in melanoma and lung cancers, respectively. (Left) We compared the proportion of C > T (and G > A) substitutions in the CpC (GpG) context (mutational signature for UV [Alexandrov et al., 2013]) between melanomas and breast cancers (controls). Because UV shows trivial impact to the nuclear DNA somatic mutations of breast cancers (Alexandrov et al., 2013), the vast majority of mtDNA C > T substitutions in the CpC context from breast cancers were not generated by UV. (Right) We compared the proportion of C > A (G > T) substitutions between lung and breast (control) cancers. C > A (G > T) substitutions are dominantly generated by tobacco smoking. Like UV, the impact of tobacco smoking to the somatic mutations of breast cancers is trivial (Alexandrov et al., 2013).
Tables
Summary statistics of mtDNA sequence data
| WGS | WXS | Average mt RD (WGS) | Average mt RD (WXS) | Total | WGS | WXS | Average mt RD (WGS) | Average mt RD (WXS) | Total | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Breast | 284 | 98 | 11594.3 | 52.7 | 382 | Meningioma | 0 | 12 | - | 42.5 | 12 |
| Colorectal | 1 | 75 | 34916.9 | 276.6 | 76 | Ependymoma | 1 | 9 | 10323.7 | 52.7 | 10 |
| Lung | 60 | 0 | 2798.1 | - | 60 | ||||||
| Prostate | 80 | 0 | 17810.6 | - | 80 | MPD | 12 | 138 | 1517.0 | 10.9 | 150 |
| Hepatocellular | 0 | 47 | - | 205.8 | 47 | MDS | 3 | 75 | 5648.7 | 44.5 | 78 |
| Melanoma | 13 | 13 | 513.9 | 353.5 | 26 | ALL | 64 | 6 | 886.6 | 35.9 | 70 |
| Gastric | 0 | 13 | - | 184.1 | 13 | CLL | 6 | 0 | 5002.2 | - | 6 |
| Cholangiocarcinoma | 0 | 8 | - | 143.9 | 8 | AML | 1 | 6 | 6783.6 | 27.4 | 7 |
| Mesothelioma | 0 | 6 | - | 106.3 | 6 | Multiple myeloma | 0 | 69 | - | 43.2 | 69 |
| Bladder | 54 | 0 | 646.2 | - | 54 | AMKL | 0 | 9 | - | 24.2 | 9 |
| Renal | 0 | 23 | - | 35.4 | 23 | Lymphoma | 0 | 4 | - | 99.5 | 4 |
| Ovarian | 0 | 38 | - | 58.9 | 38 | ||||||
| Uterine | 27 | 23 | 736.0 | 149.5 | 50 | Osteosarcoma | 38 | 90 | 9525.5 | 119.2 | 128 |
| Cervical | 0 | 52 | - | 85.2 | 52 | Chondrosarcoma | 0 | 47 | - | 99.1 | 47 |
| Adenoid cystic ca. | 1 | 60 | 714.7 | 75.6 | 61 | Ewing sarcoma | 0 | 27 | - | 69.5 | 27 |
| Head & Neck | 43 | 3 | 1369.1 | 18.8 | 46 | Kaposi sarcoma | 0 | 9 | - | 181.0 | 9 |
| Chordoma | 16 | 11 | 1240.0 | 82.1 | 27 | ||||||
| Total; 31 cancer types | 704 | 971 | 1675 | ||||||||
-
WGS, whole-genome sequencing; WXS, whole-exome sequencing; mt RD, mitochondrial read depth; MPD, myeloproliferative disease; MDS, myelodysplastic syndrome; ALL, acute lymphoblastic leukemia; CLL, chronic lymphoblastic leukemia; AML, acute myeloid leukemia; AMKL, acute megakaryoblastic leukemia.
Additional files
-
Supplementary file 1
Sequencing information of 1675 tumor–normal pairs.
- https://doi.org/10.7554/eLife.02935.021
-
Supplementary file 2
Catalogs of somatic mutations (substitutions and indels) and inherited polymorphisms identified in this study.
- https://doi.org/10.7554/eLife.02935.022
-
Supplementary file 3
List of phased somatic substitutions.
- https://doi.org/10.7554/eLife.02935.023
-
Supplementary file 4
dN/dS for 13 protein-coding genes in mitochondria.
- https://doi.org/10.7554/eLife.02935.024
-
Supplementary file 5
List of somatic substitution with higher recurrent rate than expected.
- https://doi.org/10.7554/eLife.02935.025
-
Supplementary file 6
Data accession numbers.
- https://doi.org/10.7554/eLife.02935.026