Centriolar remodeling underlies basal body maturation during ciliogenesis in Caenorhabditis elegans
Figures
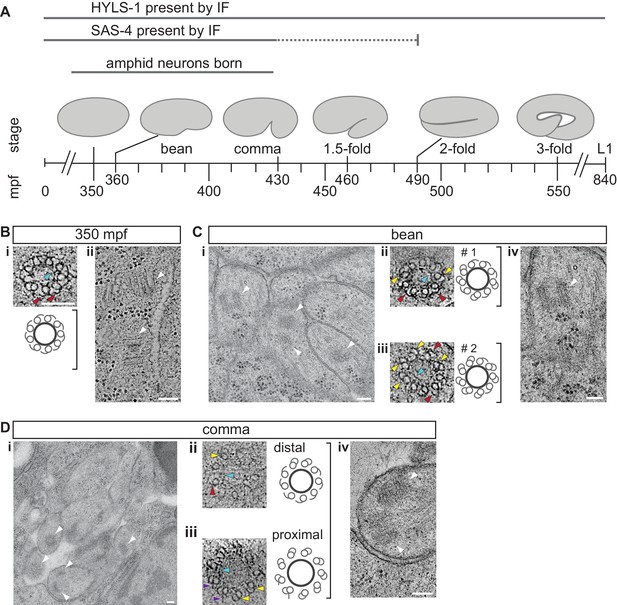
The centrioles of embryonic amphid sensory neurons remodel to initiate ciliogenesis.
(A) Timeline of embryonic development at 22°C (adapted from IntroFIG7 http://www.wormatlas.org/ver1/handbook/anatomyintro/anatomyintro.htm) showing select stages defined by morphology between fertilization (t = 0) and hatching (L1). The approximate time period during which amphid neurons are born is marked. The developmental stages during which core centriole components (e.g. SAS-4 and BB-associated protein HYLS-1 [Dammermann et al., 2009; Wei et al., 2016]) are detected by immunofluorescence (IF) in amphid neurons are marked. Dashed line indicates that the exact time of SAS-4 loss in amphid neurons is unknown. mpf – minutes post fertilization. (B–D) TEM images of the amphid channel in cross-section (Ci and Di), cross-section ET slices and schematics of a centriole in an unidentified cell of a 350 mpf embryo (Bi) and BBs in amphid neurons (Cii, Ciii, Dii, Diii), and ET slices showing centrioles/BBs in longitudinal orientation (Bii, Civ, Div) at the indicated stages of embryogenesis. Two different examples of bean-stage centrioles (#s 1 and 2) undergoing remodeling are shown in Cii and Ciii, respectively. Arrowheads: centrioles/BBs (white), dMTs (yellow), sMTs with hook appendages (red), central tube (light blue), putative nascent Y-links (purple). Scale bars: 100 nm.
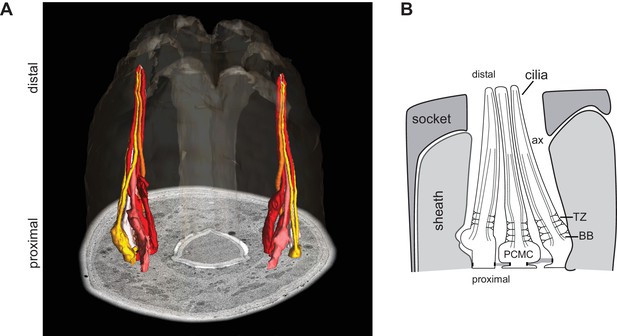
The cilia of a subset of amphid sensory neurons extend through a channel created by glial support cells.
(A, B) ssTEM 3D reconstructed model (A; adapted from Doroquez et al. [2014]) and schematic (B) showing a subset of amphid channel cilia. Surrounding sheath and socket glial cells are also shown in the schematic. Anterior is at top. BB - basal body, TZ - transition zone, ax - axoneme, PCMC – periciliary membrane compartment.
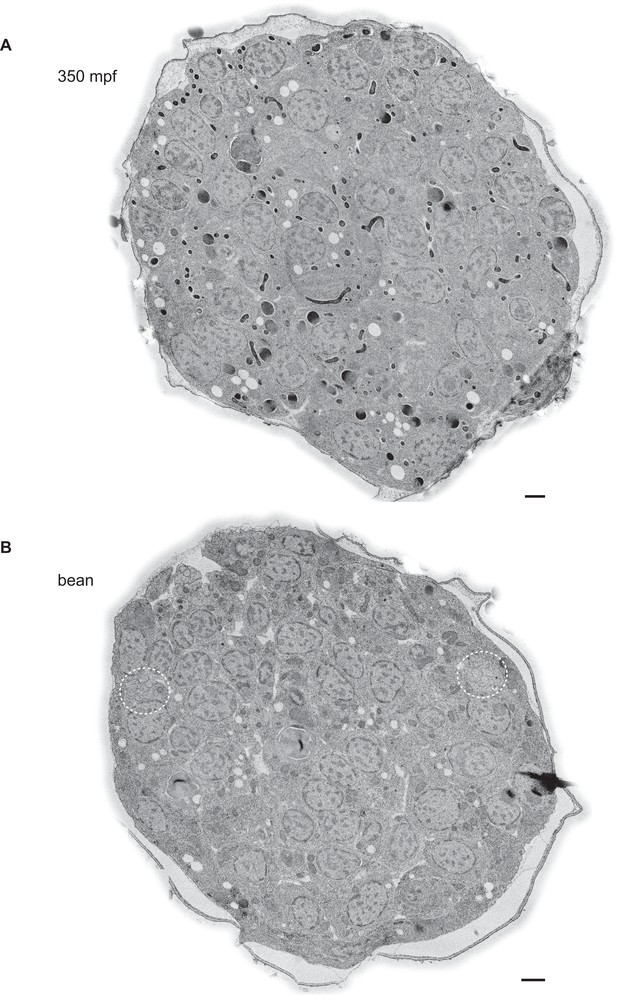
Example TEM cross-section images of C. elegans embryos at the 350 mpf and bean stages.
(A, B) TEM cross-sections of HPF-FS C. elegans embryos of the indicated stages; montages of overlapping images were recorded of 70-nm thick cross-sections; edges surrounding the cuticle were feather-cropped. The amphid channels are indicated by white dashed circles in B. Scale bars: 1 μm.
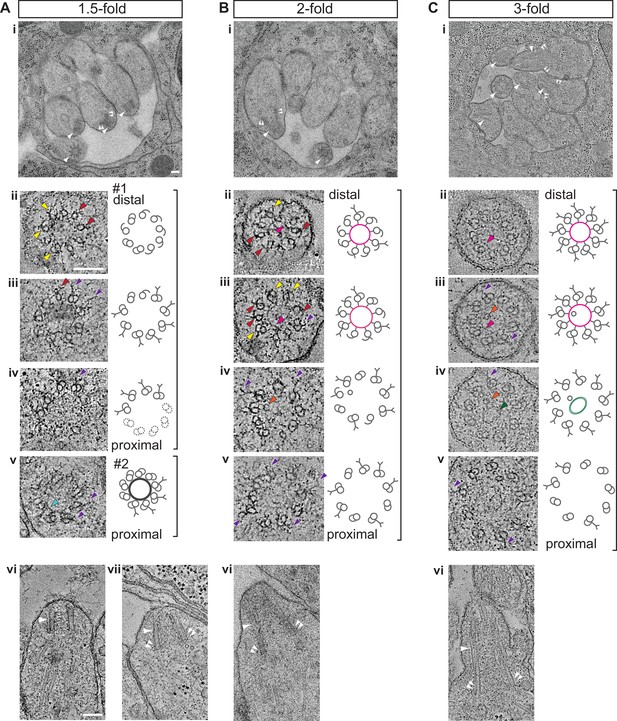
Degeneration of the central tube and dMT flaring at the ciliary base are observed by the 1.5-fold stage of embryogenesis.
(A–C) TEM images (Ai and Bi) and ET slice (Ci) of the amphid channel in cross-section, cross-section ssET images and schematics of BBs in amphid neurons (Aii–v, Bii–v, Cii–v), and ET slices (Avi, Avii, Cvi) and TEM image (Bvi) showing BBs/axonemes in longitudinal orientation at the indicated stages of embryogenesis. Cii–v show a subset of ET slices from the serial section tomogram. Examples of BBs/axonemes in 1.5-fold embryo with a largely degenerated central tube and an intact central tube are shown in Aiv and Av, respectively. Arrowheads: centrioles/BBs (large white), flared dMTs at cilia base (small white double), dMTs (yellow), sMTs with hook appendages (red), central tube (light blue), Y-links (purple), apical ring (pink), isMTs (orange), a vesicle (green). Each bracket delineates a single BB/axoneme with its proximal and distal regions marked accordingly. Scale bars: 100 nm.
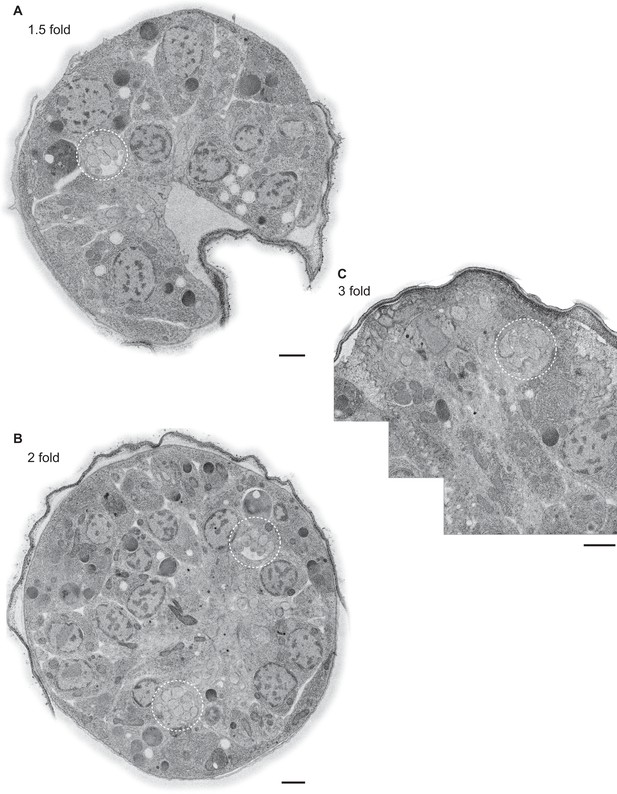
Example TEM cross-section images of C. elegans embryos at the 1.5-, two-, and three-fold stages.
(A–C) TEM cross-sections of HPF-FS C. elegans embryos of the indicated stages; montages of overlapping images were recorded of 70-nm thick cross-sections; edges surrounding the cuticle were feather-cropped. The amphid channels are indicated by white dashed circles. Scale bars: 1 μm.
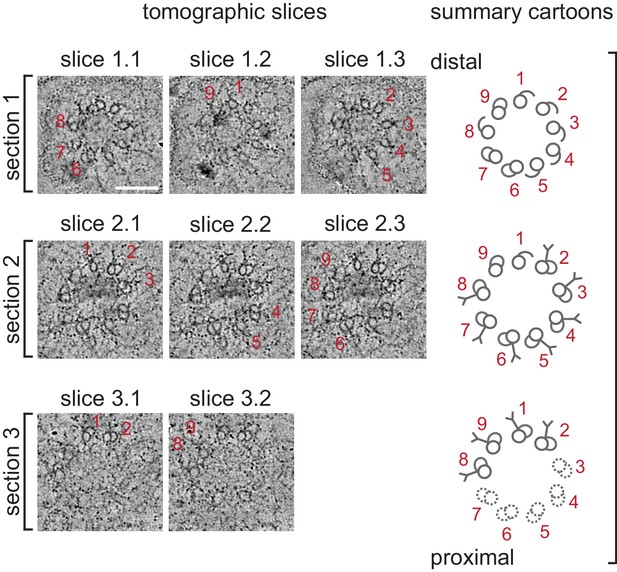
Selected ET slices of tomograms of a 1.5-fold BB/axoneme reconstructed from sequential plastic sections.
Three selected tomographic slices (left; ET slices 1–3) of three tomograms that were reconstructed from sequential plastic sections #1–3 from the same cilium (proximal and distal regions marked accordingly) from a 1.5-fold embryo. Thickness of each tomographic slice is 10 nm. The main structural features observed in each tomogram are highlighted and summarized in cartoons (right). dMTs are numbered.
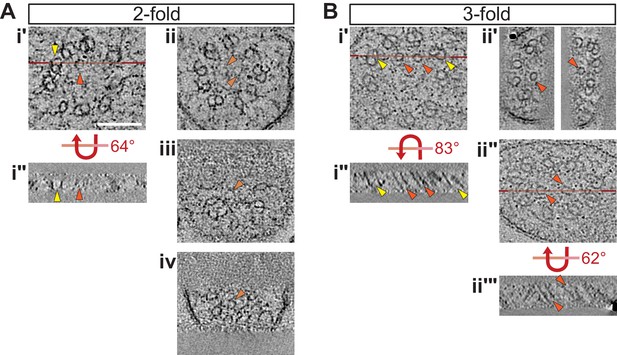
Selected ET slices of tomograms of two- and three-fold cilia showing isMTs.
(A, B) ET slices showing examples of cilia with fully formed isMTs in cross-section (Ai’, Bi’, Bii’ and Bii’) and in longitudinal view (Ai’’, Bi’’, and Bii’’’) at the indicated embryonic stages. Example ET slices showing cilia with very short/incompletely formed isMTs in cross-section are depicted in Aii–Aiv. Horizontal red lines in Ai’, Bi’, and Bii’’ indicate the location of the tomographic slices shown in Ai’’, Bi’’, and Bii’’’, respectively. Bii’’–Bii’’’ shows the same slice of a cilium, but in Bii’ more tomographic slices were binned and the projection angles of the right and left isMT adjusted to optimize their appearance, whereas Bii’’ is a thin tomographic overview slice to indicate the location of Bii’’’. Arrowheads: dMTs (yellow), fully formed isMTs (orange), incompletely formed isMTs (light orange). Scale bar: 100 nm.
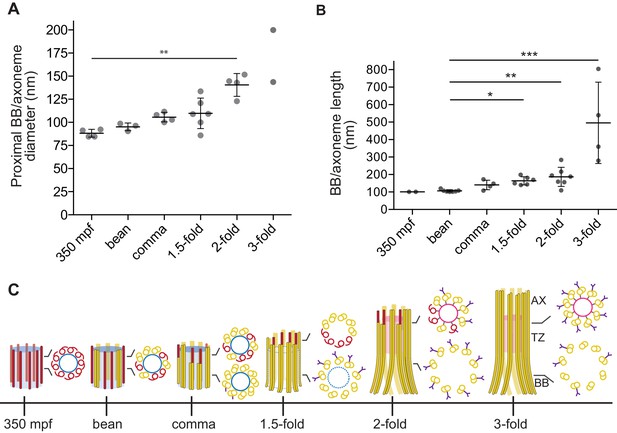
Quantification of BB/axoneme diameter and length in embryonic amphid sensory neurons.
(A) Quantification of the centriole/BB/axoneme diameter measured as the distance between centers of A-tubules. The first tomographic slice of a ssET sequence showing the entire proximal region of each BB/axoneme in cross-section was used for measurements. Each dot represents a measurement from an individual BB/axoneme in different neurons from the same embryo. Horizontal bars indicate mean. Errors are SD. ** indicate that marked data sets are different at p<0.01 (Kruskal-Wallis test with post-hoc correction for multiple comparisons). (B) Quantification of the centriole/BB/axoneme length at the indicated stages of embryonic development. Each dot represents a measurement from an individual BB/axoneme in different neurons from the same embryo. Horizontal bars indicate mean. Errors are SD. *, **, and *** indicate that marked data sets are different at p<0.05, 0.01, and 0.001, respectively (Kruskal-Wallis test with post-hoc correction for multiple comparisons). (C) Model summarizing key early ciliogenesis stages in the examined subset of C. elegans sensory neurons. BB – basal body; TZ – transition zone; AX – axoneme. Blue and pink circles indicate the central tube and apical ring, respectively. Model is based solely on our ability to visualize specific ciliary structures, and no assumptions are made regarding the presence or absence of proteins associated with these structures.





