Shared rhythmic subcortical GABAergic input to the entorhinal cortex and presubiculum
Figures
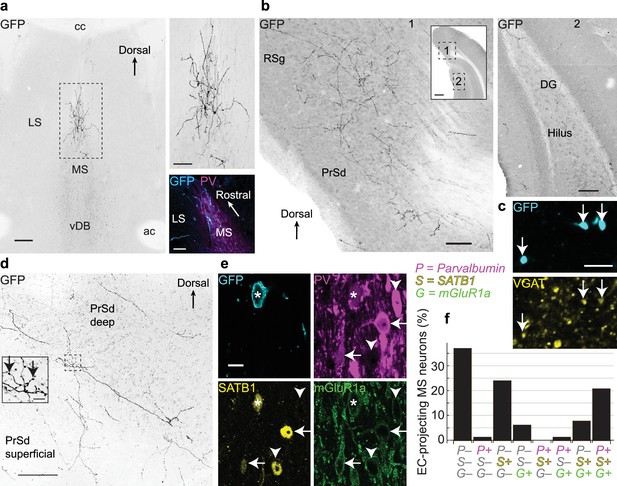
Medial septal GABAergic neurons terminating in the entorhinal cortex also innervate the dorsal presubiculum and retrosplenial cortex.
(a) Coronal section showing EC-projecting GFP-immunoreactive neurons restricted to the dorsal MS, following injection of PRV-hSyn-Cre into the caudo-dorsal EC and AAVDIO-EYFP into the MSDB (animal MS60). Top right, enlarged view of the boxed region. Bottom right, horizontal section (animal MS77) showing GFP-immunoreactive neurons (cyan) restricted to the rostral part of the dorsal MS, delineated by PV immunoreactivity (magenta). (b) Virally-labeled axon collaterals of EC-projecting MSDB neurons densely innervating extra-hippocampal regions (left, 1) and sparsely innervating the DG and CA1 (right, 2) from a single coronal section (animal MS60). Inset, locations of Figures 1 and 2 (boxes). (c) A subset of axon terminals from EC-projecting MSDB neurons in the RSg (GFP, cyan) are immunoreactive for VGAT (yellow, arrows) (animal MS66). (d) Coronal section of the PrSd (animal MS60) with axon collaterals and terminals from EC-projecting MSDB neurons. Inset, enlarged view of boxed region (arrows, axon terminals). (e) An EC-projecting medial septal neuron soma (asterisk, from animal MS66) immunoreactive for GFP (cyan, plasma membrane) was weakly immunoreactive for PV (magenta), SATB1 (yellow, nucleus) and mGluR1a (green, membrane). Note GFP-negative neurons with similar (arrows) or different (arrowheads) molecular profiles. (f) Quantification of PV (P), SATB1 (S) and mGluR1a (G) immunoreactivity for virally-labeled EC-projecting neurons located in the dorsal MS (data from six animals). Scale bars (μm): (a) 200 (left image), 100 (right images); (b) 100 (inset 500); (c) 5; (d) 100, inset 10; (e) 10. Image type: (a–b) Widefield epifluorescence, reverse contrast, 70 μm thick sections. (c) confocal image, single optical section, 0.31 μm thick; (d) confocal image, reverse contrast, maximum intensity z-projection, 35 sections, 30.96 μm thick; (e) confocal image, single optical section, 0.38 μm thick. LS, lateral septum; cc, corpus callosum; ac, anterior commissure.

Medial septal neurons terminating in the entorhinal cortex also innervate the dorsal presubiculum.
(a) Distribution of EYFP +axons (reverse contrast fluorescence) from the MSDB in the caudal EC across three 70 µm-thick coronal sections from the same animal (MS60). Left, three sections caudal of the injection site; middle, two sections caudal of the injection site. Boxed regions are enlarged below. Right, three sections rostral of the injection site, with the location of parasubiculum (PaS) and superficial layers of the lateral EC (lEC) marked by Wfs1 immunoreactivity (top). Blood vessel, b.v. Axons are distributed across all EC layers. (b) Left, the injection site of animal MS60 is marked by an asterisk. Right, location of EYFP+ axons from the MSDB in the dorsal EC (horizontal section, animal MS78). (c) Expanded view of a horizontal section (see Figure 1a) showing GFP-immunoreactive EC-projecting medial septal neurons (cyan) restricted to the rostro-dorsal MS (animal MS77). PV, magenta. (d) Coronal section of the temporal cortex (animal MS60, see Figure 1b) with virally-labeled axon collaterals of EC-projecting MSDB neurons innervating extra-hippocampal regions (PrSd, RSg). Top right, enlarged view of boxed region. V1, primary visual cortex; mEC, medial EC; SUB, subiculum. Image type: (a–d) Widefield epifluorescence, 70 μm thick sections. Images in (a), (b) (right) and (c) are montages. Scale bars (µm): (a) left: top 500, bottom 200; middle: top 500, bottom 100; right: 200; (b) 200; (c) 100; (d) 250, inset 100.
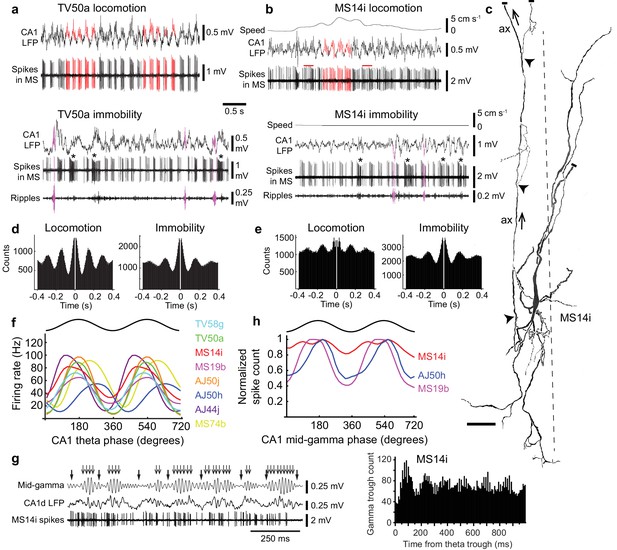
Firing patterns of medial septal neurons projecting to the EC.
(a) Top, rhythmic burst firing of EC-projecting medial septal neuron TV50a during locomotion. CA1 LFP, local field potential in SP. Red, examples of bursts of spikes associated with peaks of CA1 theta oscillations. Bottom, rhythmic burst firing during immobility with some spikes during some SWRs (magenta; 130–230 Hz filtered CA1 LFP). Examples of long accommodating bursts are shown below the asterisks. (b) Same as in (a) for another EC-projecting neuron, MS14i, which fires less rhythmic bursts at high rate during locomotion (top) and rhythmic bursts during immobility (bottom). Red bars, examples of bursts spanning two theta cycles. (c) Reconstruction of the soma, dendrites, and axon (ax; arrows: towards dorsal fornix) of MS14i in the tip of dorsal MS. Dashed line, midline. Local axon collaterals, which give rise to terminals, are marked by arrowheads. Cut ends not included in the reconstruction are marked by small bars. Scale bar, 50 μm. (d,e) Autocorrelograms of spikes during locomotion and immobility for TV50a (d) and MS14i (e). (f) Firing rate versus CA1 theta phase plots show preferential firing near the peak of theta during movement (locomotion and small movements) for eight labeled neurons (color-coded) projecting to extra-hippocampal regions. Black, sinusoid representation of two idealized cycles. Data are duplicated to represent rhythmicity. (g) Left, part of the time series from (a) during locomotion displaying several theta cycles in the unfiltered LFP (thick arrows mark theta troughs), and 55–80 Hz filtered LFP mid-gamma cycles (thin arrows mark high amplitude gamma troughs). Note that gamma troughs occur mostly at the peak of theta cycles. Right, event correlation of detected mid-gamma troughs relative to theta troughs from the MS14i recorded CA1d LFP. Note peak around 100 ms corresponding to the peak of theta cycles. (h) Normalized spike count versus CA1 mid-gamma phase for all significantly coupled EC-projecting neurons (see also Table 3).

Rhythmicity of identified orchid cells.
Autocorrelograms of spikes for identified orchid cells during different behavioral states. Note that cells fire rhythmically during all behavioral states.
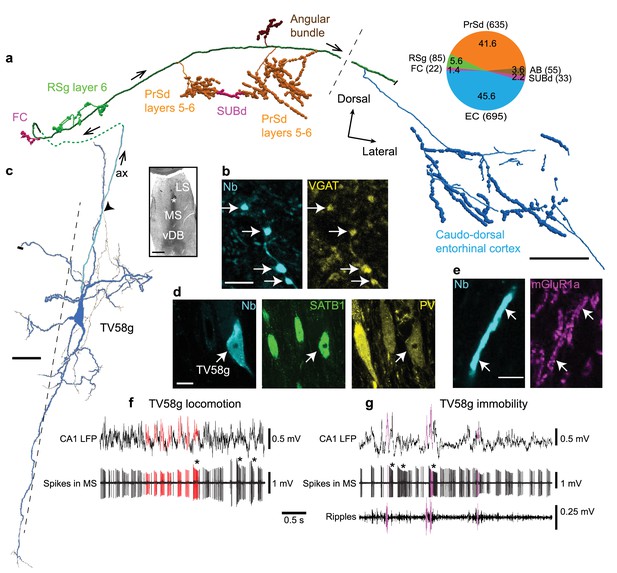
Firing patterns and cortical target regions of a GABAergic medial septal orchid cell.
(a) Reconstruction of part of the axon of neuron TV58g axon (cortical regions color-coded). Black dashed line, rostral (left) and caudal (right) innervation zones. Pie chart, number (parentheses) and proportion (%) of axonal varicosities. Projection axon enters the cortex via the dorsal fornix (green dashed line). (b) Presubicular terminals of TV58g (cyan) were VGAT+ (yellow, arrows). (c) Micrograph (left, asterisk) and reconstruction (right) of the soma and dendrites (blue) and axon (cyan, ax) of TV58g in the tip of dorsal MS. Black dashed line, midline. A local axon collateral is marked by an arrowhead. A cut dendrite is marked by a small bar. (d) The soma of TV58g (cyan, neurobiotin, arrow) was SATB1+ (green, nuclei) and PV+ (yellow). Note similar profiles of adjacent neurons. (e) Dendrites of TV58g (cyan) were mGluR1a+ (magenta, arrows). (f) Rhythmic burst firing during locomotion. CA1 LFP, local field potential in SP. Red, examples of detected peaks of CA1 theta oscillations and associated MS neuron spikes. Note preferential coupling of bursts to theta peak. (g) Rhythmic burst firing during immobility with some spikes during some SWRs (magenta; ripples, 130–230 Hz filtered CA1 LFP). Examples of accommodating bursts are shown below asterisks in (f) and (g). Nb, neurobiotin. FC, fasciola cinereum, AB, angular bundle, LS, lateral septum, vDB, vertical diagonal band. Images (z-thickness in µm and z-projection type): (d) 2.3, maximum, (b) 2.61, maximum, (e) 0.37, single optical section. Scale bars (µm): a, 500, b,e, 5, c, 50, inset, 500, d, 10.
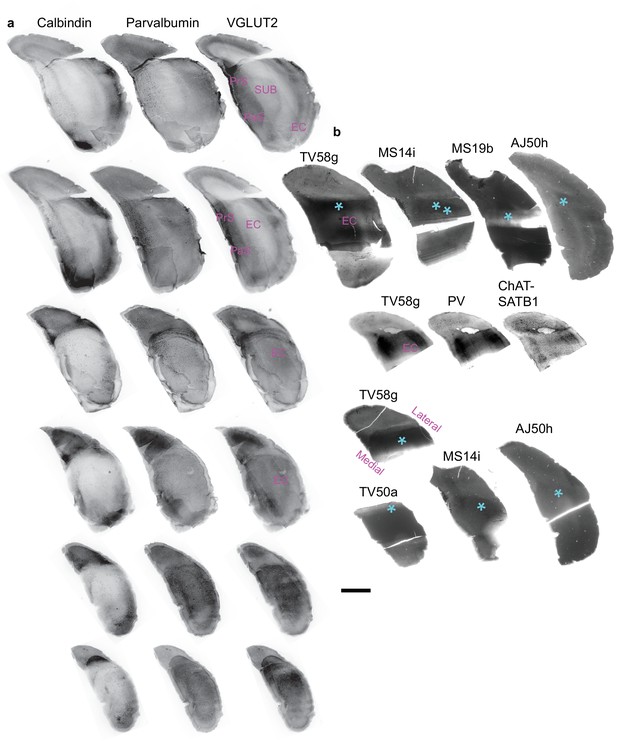

Location of the axons of orchid cells in the caudo-dorsal entorhinal cortex.
(a) Consecutive series of ‘reference’ coronal sections through the caudal temporal cortex in three columns (top to bottom;~70 µm thickness; one brain) immunoreacted for CB and PV and VGluT2 (three channels per section; widefield epifluorescence montages, reverse contrast). (b) Actual coronal sections of brains TV58, TV50, MS14, MS19 and AJ50 containing the axons of recorded and labeled orchid cells (not visible) aligned relative to reference sections in (a). Asterisks (cyan) mark exact positions of orchid cell axons and/or terminals in the EC. Two TV58 sections and one of TV50 have missing ventral and dorsal regions, respectively. Images are from diaminobenzidine (DAB)-based horseradish peroxidase (HRP) reacted sections except the second row (TV58g section) that shows widefield fluorescence of a streptavidin-conjugated fluorophore along with immunoreactivity for PV and double-immunoreacted for ChAT and SATB1, delineating layers. Sections from MS14 and MS19 have been flipped horizontally to appear in the same orientation as the others. PaS, parasubiculum. Scale bar, 1 mm.

Location of labeled orchid cell axon in the dorsal presubiculum (I).
Reference images of in situ hybridization expression from the Allen Brain Atlas at different rostro-caudal levels (top to bottom) of coronal mouse brain sections for Slc17a6 (a), Satb2 (b), Etv1 (c) and Nos1 (d) genes. Note strong Nos1 mRNA expression in PrS, but absence in the adjacent areas. Asterisks, filled circles and open circles (all cyan): examples of approximate locations of orchid cells TV58g, AJ50j and AJ50h axon terminals and/or thin collaterals. Granular retrosplenial cortex, RSg; dorsal subiculum, SUBd.

Location of labeled orchid cell axon in the dorsal presubiculum (II).
Reference images of in situ hybridization expression from the Allen Brain Atlas at different rostro-caudal levels (top to bottom) of coronal mouse brain sections for Slc17a6 (a) Satb2 (b) and Etv1 (c) genes for the delineation of the dorsal presubiculum. (d) Two representative light microscopic coronal sections of calbindin immunoreactivity (HRP-reaction) show delineation of layers 1–6. Layer two is a compact layer, and layer four is an interplexiform layer. Arrow, part of layer two enriched with CB-immunopositive neurons. Scale bars, 100 µm. (e) Representative sagittal sections of the three genes from (a–c). Asterisks, filled circles and open circles (all cyan): examples of approximate locations of orchid cells TV58g, AJ50j and AJ50h axon terminals and/or thin collaterals; the most caudal filled circle in (b) corresponds to the most caudal location of AJ50j projection axon. Entorhinal cortex, EC.
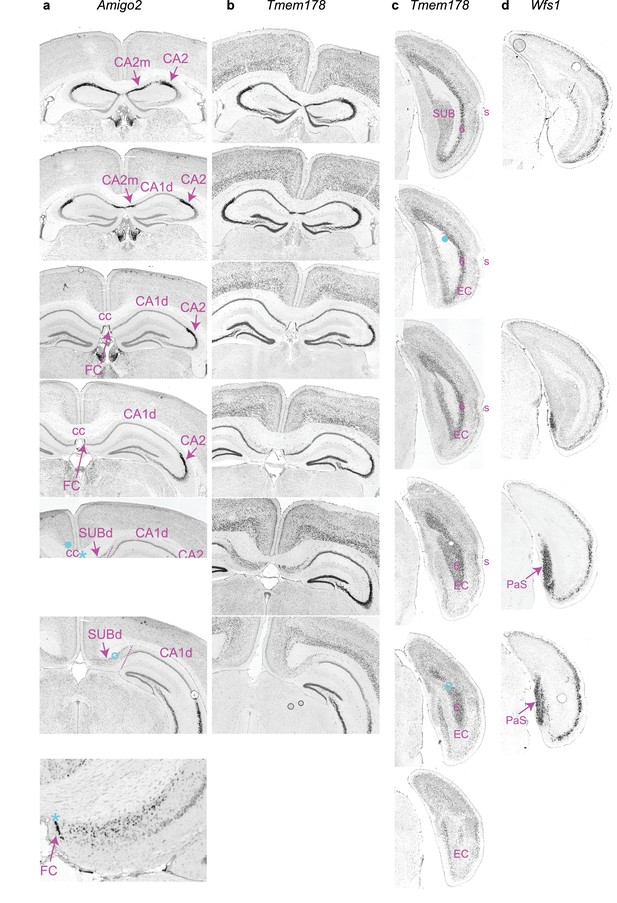
Delineation of synaptic target regions of orchid cells.
(a) Reference images of in situ hybridization expression for Amigo2 highlights CA2 and the FC at different levels of the septal pole of the hippocampus (coronal sections). Top to bottom, rostral to caudal examples. Note that the FC is caudal of medial CA2 (CA2m). Bottom image, enlarged view of innervated FC region. (b,c,) In situ hybridization data for Tmem178 delineates CA3, CA2 and FC (b) and layer 6 of EC (c) in coronal sections. (d), In situ hybridization data for Tmem178 delineates the PaS in caudal coronal sections. Magnification in a,b) are the same; others are different. Asterisks, filled circles and open circles (all cyan): examples of approximate locations of orchid cells TV58g, AJ50j and AJ50h axon terminals and/or thin collaterals; filled circle in c) corresponds to the most caudal location of AJ50j projection axon. Corpus callosum, CC; rhinal sulcus, S. All images obtained from the Allen Brain Atlas.
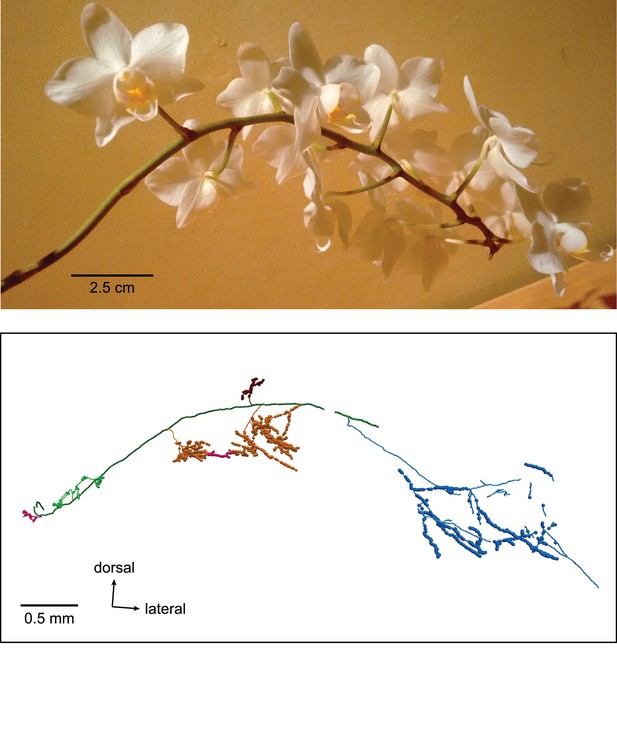
Resemblance of a medial septal axon to a moth orchid.
Top, Phalaenopsis orchid. Bottom, orchid cell cortical axon (TV58g).
Reconstruction of orchid cell TV58g.
Green arrow, dorsal; red arrow, lateral; blue arrow, caudal.
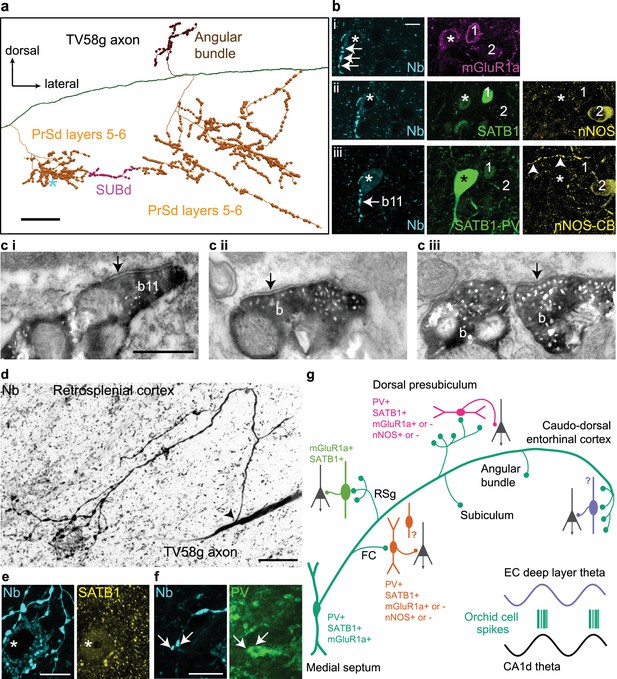
Postsynaptic GABAergic target neurons of a GABAergic orchid cell, TV58g.
(a) Selected collaterals and varicosities (see Figure 2a). Asterisk, location of target neuron in b. (b) Boutons (cyan, neurobiotin, arrows) in the PrSd (asterisk in a) are apposed to a neuron immunopositive for mGluR1a (i, top, magenta, somato-dendritic membrane), SATB1 (ii, middle, green, nucleus), and PV (iii, bottom, green) but lacking detectable immunoreactivity for both nNOS (ii, yellow) and CB (iii, yellow). Arrowheads in iii (bottom right) highlight a CB-immunoreactive process. Two neighboring PV-immunonegative neurons (1, 2) were not targeted. (c) Electron micrographs of synapses (arrows) made by boutons (b) of TV58g on interneurons in the PrSd with the proximal dendrite of a PV+ cell (i, b11 in b), (ii) the dendrite of an unknown interneuron and (iii) soma of another interneuron. (d) Main axon and collateral (arrow, branch point) along with terminals surrounding a neuron in layer 6 of RSg (reverse contrast, confocal microscopic image). The target neuron contains a high level of endogenous biotin (high background), revealed by the streptavidin-conjugated fluorophore signal. (e) The target neuron soma (asterisk) innervated by multiple terminals (cyan) from (d) showed nuclear immunoreactivity for SATB1 (yellow). (f) The dendrite of the target showed positive immunoreactivity for PV (green); arrows highlight two terminals in close apposition. Nb, neurobiotin. (g) Schematic summary of the data presented in Figures 3 and 4 and Tables 1 and 6. Green circles represent the relative proportions of cortical axon terminals from all orchid cells (see Table 1). Immunoreactivity of synaptic target neurons is color-coded based on innervated cortical region. Innervated cortical interneurons may target principal cells (dark gray) on the dendrites, soma or axon initial segment. Bottom right, two idealized theta cycles with representative orchid cell spike bursts at the peak of CA1d theta and EC theta (see Figure 6). Images (z-thickness in µm and z-projection type): (b) 3.63/4.3/4.76, (i/ii/iii), maximum, (d) 22.09 standard deviation, (e) 2.8 maximum, (f) 2.88 maximum. Scale bars: (a) 100 µm, (b,e,f) 10 µm, (c) 0.5 µm (applies to i/ii/iii), (d) 20 µm.
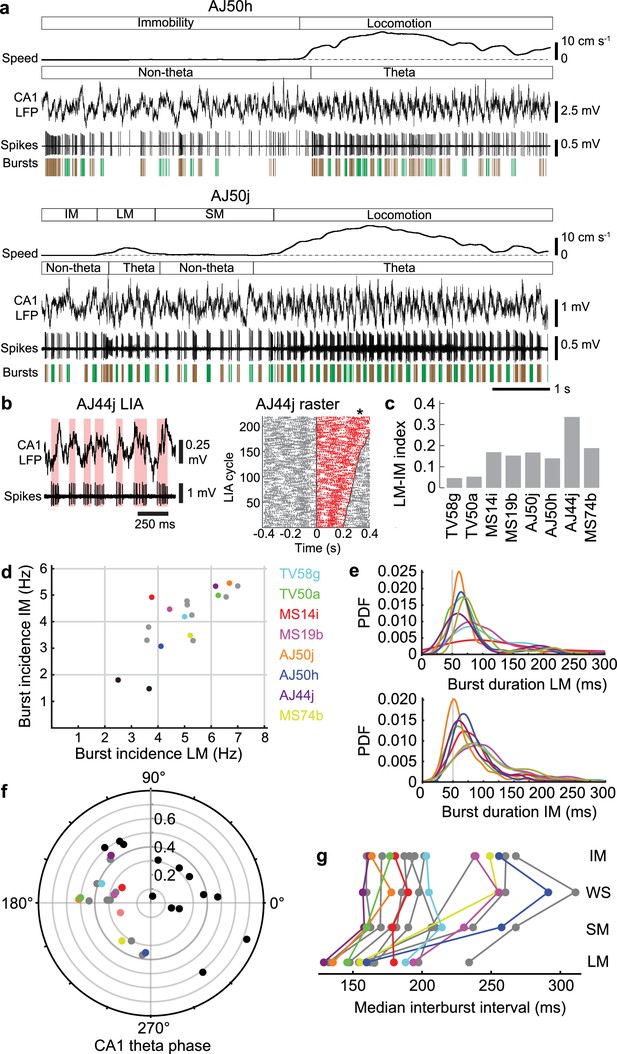
Orchid cell firing reflects behavioral states.
(a) Firing of orchid cells AJ50h (top) and AJ50j (bottom), recorded consecutively in the same session. Algorithm detected bursts are shown below spikes (sequential bursts marked alternately green and brown). Note periods of theta oscillations and non-theta periods during different behavioral states. (b) Left, rhythmic burst firing of orchid cell AJ44j during CA1 LIA (‘non-theta’ periods), with red bars highlighting bursts aligned mostly to the ascending/peak phase of LIA oscillations. Right, raster of spikes within (red) and outside (gray) zero-crossings of LIA cycles, ordered by cycle length (minimum interval 200 ms). Time 0 marks the falling transition of the LFP (zero crossing). Asterisk, position of the peak of the LFP during slower LIA cycles, corresponding to greater spike counts. (c), Mean firing rate index for different states (IM versus LM) for identified orchid cells. All neurons increase their firing from IM to LM. (d) Burst incidence during IM versus LM for identified orchid cells (color-coded, cell IDs on right side), putative orchid cells (gray) and septo-hippocampal cells (black). (e) Probability density functions (PDF) of burst durations of identified orchid cells (color-coded as in c) during LM (top) and IM (bottom), showing that peak burst durations are greater than 50 ms (gray line). (f) Preferred CA1 theta phase (degrees) of identified orchid cells (color-coded as in d), putative orchid cells (gray), and identified Teevra cells (black, cells from Joshi et al., 2017); vector length as radial axis (r). (g) Median interburst interval for identified and putative orchid cells during different behavioral states (points; some neurons lack particular states). LM, locomotion; SM, small movements; WS, whisking/sniffing without movement; IM, immobility.
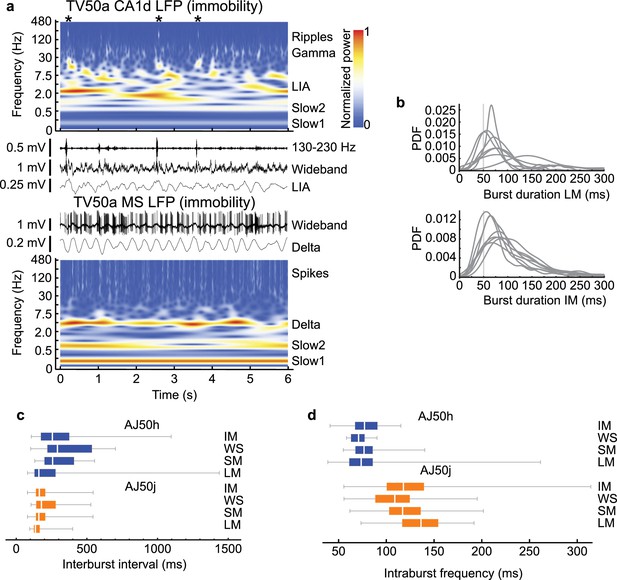
Burst firing characteristics during different behavioral states.
(a) Time-frequency plots and LFP time series during immobility for orchid cell TV50a. Top, CA1d LFP, bottom, MS LFP, same time period. Note LIA in CA1d LFP consisting of various slow frequencies (red/yellow), and ‘delta’ oscillation in MS LFP. ‘Slow1’ and ‘Slow2’ are other detected slower oscillations. LIA and delta low-pass filtered traces are shown below their respective wideband LFPs. The MS LFP is located very close to the soma of TV50a, hence high amplitude action potentials. Asterisks, location of ripples. (b) Probability density functions (PDF) of burst durations of putative orchid cells during LM (top) and IM (bottom), showing that peak burst durations are greater than 50 ms (light gray line). (c, d) Interburst intervals (c) and intraburst frequencies (d) during different behavioral states for identified orchid cells AJ50h (top) and AJ50j (bottom) recorded sequentially in the same animal. White bars, median. Boxes, quartiles. LM, locomotion; SM, small movements; WS, whisking/sniffing without movement; IM, immobility.
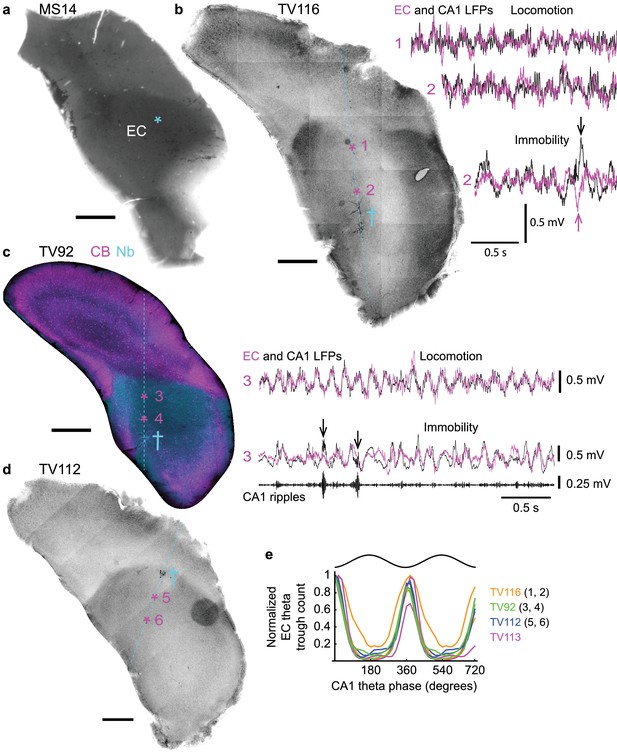
Phase relationship of LFPs in dorsal CA1 and entorhinal cortex.
(a) The position of axon terminals from orchid cell MS14i in the caudo-dorsal EC is marked by an asterisk (DAB-based HRP reacted, 70 µm-thick coronal section flipped horizontally). (b–d) Coronal sections from different animals (TV116, TV92, TV112) showing the EC at similar rostro-caudal levels to (a) with position of glass electrode tracts (cyan, dashed lines). Cross symbols mark sites of labeled neurons used to assess relative depth of LFP recording sites. (b) Left, reverse contrast fluorescence image (neurobiotin signal). Asterisks mark two LFP recording sites. Right, overlaid entorhinal (magenta) and CA1d (black) LFPs recorded at sites 1 and 2. Arrows, sharp waves. (c) Immunoreactivity for CB (magenta) and visualization of neurobiotin (Nb, cyan; widefield epifluorescence). Asterisks mark two LFP recording sites. Right, LFP recordings from site three during locomotion and immobility. Ripples, 130–230 Hz filtered CA1d LFP (also marked by arrows in the LFP). (e) Theta phase histogram for EC LFP theta (5–12 Hz) trough counts (normalized) relative to CA1d theta phase (n = 4 mice, two sites per mouse). Scale bars for all images: 500 µm.

Ripple-related activity of orchid cells and putative orchid cells.
(a-d) Firing probability density (top), spike raster (middle) and cumulative distribution functions (CDFs, bottom) of firing patterns during (red) and outside (gray) SWRs for four identified orchid cells. In the raster plot, continuous staggered lines delineate start and end of aligned ripples, horizontal lines delineate ripples within 0.5 s of an aligned ripple. The CDFs show the firing rate distributions during actual ripples (red curves) and for 1000 shuffled distributions (gray) across three behavioral states (immobility, whisking/sniffing, small movements); black curves, median shuffled distributions; dashed curves, 95% confidence intervals. When the red curve falls between the confidence intervals the cell fires during SWRs by chance. MS19b is inhibited during most SWRs, whereas TV58g is activated during SWRs with high firing rates. (e–i) Same as (a–d) but for putative orchid cells. MS17k is activated during SWRs, with SWRs mostly occurring on the second half of individual rhythmic bursts (e, top).
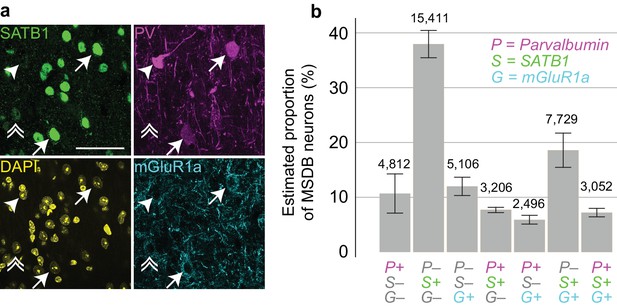
Proportions of MSDB neurons immunoreactive for PV, SATB1 and mGluR1a.
(a) Confocal microscopic z-projection (3 µm thick) of SATB1 (green), PV (magenta) and mGluR1a (cyan) immunoreactivity in the MS. Arrows, two PV+/SATB1+/mGluR1a+ neurons. Arrowhead, PV+/SATB1–/mGluR1a– neuron. Double arrowhead, PV–/SATB1+/mGluR1+ neuron. DAPI, yellow. Scale bar, 50 µm. (b) Stereological estimation of neuronal subpopulations in the MSDB immunoreactive for combinations of PV (P), SATB1 (S) and mGluR1a (G). Total estimated counts are shown above each bar. Error bars, standard error.
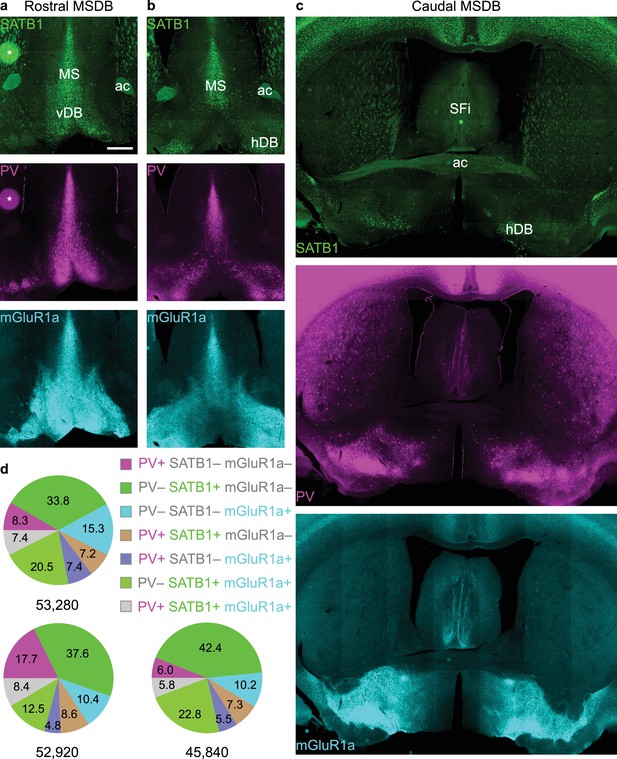
Stereological estimation of the number of neurons immunoreactive for PV and/or SATB1 and/or mGluR1a across the entire MSDB.
(a-c) Widefield epifluorescence montages of coronal sections of the MSDB at three different rostro-caudal levels. Nuclear SATB1 immunoreactivity (green) was used to define the borders of the MSDB. Asterisk marks a labeling artefact. PV, magenta; mGluR1a, cyan. (d) Pie charts of estimated proportions (%) of neurons across the entire MSDB tested for immunoreactivity for the three molecules (based on n = 444, 441 and 382 neurons counted from three mice). Total number of estimated neurons are shown below each pie chart. Scale bar (a–c) 500 µm. MS, medial septum; vDB, vertical Diagonal Band; ac, anterior commissure; hDB, horizontal Diagonal Band; SFi, septofimbrial nucleus.
Tables
Axon terminal distribution and firing patterns of identified orchid cells.
All projection axons traveled in the dorsal fornix. *Varicosities were sampled from different numbers of brain sections per labeled neuron. **Main axon observed dorsal of SUBd/PrSd, too weak to observe collaterals. ***Main axon faded away just rostral of caudo-dorsal EC. Abbreviations: c, fine axon collaterals observed; u, unknown/unavailable; n.o., not observed. Spike burst defined as >3 spikes with interspike intervals (ISIs) < 40 ms. Firing rates and burst incidence (Hz) are expressed as mean of 1 s bins ± s.d.; intraburst frequency (Hz) as mean ±s.d.; burst duration and interburst interval (ms) are expressed as median and interquartile range. Abbreviations: LM, locomotion; SM, small movements including limbs, tail, and shifts in posture; WS, high-frequency whisking and/or sniffing in the absence of other movements; IM, immobility. KS, Kolmogorov-Smirnov; SP, stratum pyramidale; SO, stratum oriens.
| Cell name | TV58g | TV50a | MS14i | MS19b | AJ50j | AJ50h | AJ44j | MS74b | |
|---|---|---|---|---|---|---|---|---|---|
| Target hemisphere | Right | Right | Left | Left | Left | Right | Right | Right | |
| Varicosities per region (n sampled)* | Medial septum | 61 | u | 146 | n.o. | n.o. | 51 | n.o. | u |
| Dentate gyrus | n.o. | n.o. | n.o. | n.o. | n.o. | n.o. | n.o. | u | |
| CA3 | n.o. | n.o. | n.o. | n.o. | n.o. | n.o. | u | u | |
| CA2 | n.o. | n.o. | n.o. | n.o. | n.o. | n.o. | u | u | |
| CA1 | n.o. | n.o. | (c) | n.o. | n.o. | n.o. | u | u | |
| SUBd | 33 | n.o. | 3 | 104 | n.o. | c | u | u | |
| FC | 22 | (c) | n.o. | n.o. | n.o. | n.o. | u | u | |
| RSg | 85 (L6) | n.o. | n.o. | n.o. | 51 | n.o. | u | u | |
| Angular Bundle | 55 | 11 | n.o. | n.o. | n.o. | n.o. | u | u** | |
| PrSd L1-2 | n.o. | 3 | 8 | 15 | c | n.o. | u | u | |
| PrSd L3 | n.o. | 10 | 31 | 149 | n.o. | c | u | u | |
| PrSd L5-6 | 635 | 66 | 45 | 34 | 27 | 29 | u | u | |
| PaS | n.o. | n.o. | n.o. | n.o. | n.o. | n.o. | u | u | |
| EC | 695 | c | 32 | 63 | (c)*** | 6 | u | u | |
| Labeling strength | +++ | ++ | ++ | ++ | ++ | ++ | + | + | |
| Behavioral states | Max speed (cm/s) | u | u | 6.3 | 18.8 | 17.2 | 16.5 | 12.0 | u |
| Firing rate LM | 48.4 ± 6.0 | 51.4 ± 10.6 | 67.8 ± 12.9 | 43.4 ± 9.3 | 65.3 ± 11.7 | 33.6 ± 9.6 | 76.7 ± 11.0 | 55.0 ± 11.0 | |
| Firing rate SM | 41.6 ± 5.3 | 47.6 ± 8.8 | 51.0 ± 12.7 | 33.3 ± 8.6 | 42.5 ± 9.7 | 27.7 ± 3.8 | 43.8 ± 9.9 | u | |
| Firing rate WS | 47.1 ± 9.3 | u | 47.9 ± 9.0 | 33.2 ± 8.0 | 34.7 ± 6.1 | 24.3 ± 4.9 | 37.7 ± 2.5 | 34.2 ± 14.9 | |
| Firing rate IM | 44.2 ± 12.0 | 46.3 ± 10.5 | 48.2 ± 10.0 | 31.9 ± 5.2 | 46.6 ± 9.9 | 25.4 ± 7.6 | 38.1 ± 6.8 | 37.6 ± 16.0 | |
| Burst incidence LM | 5.0 ± 1.0 | 6.3 ± 0.9 | 3.8 ± 1.7 | 4.4 ± 0.9 | 6.7 ± 1.0 | 4.1 ± 1.7 | 6.2 ± 1.9 | 5.2 ± 1.3 | |
| Burst incidence SM | 4.4 ± 0.5 | 5.7 ± 1.0 | 4.7 ± 1.1 | 3.6 ± 1.1 | 5.5 ± 1.3 | 3.3 ± 0.8 | 5.6 ± 1.2 | u | |
| Burst incidence WS | 4.4 ± 0.9 | u | 4.6 ± 1.1 | 4.4 ± 1.3 | 4.6 ± 1.0 | 2.8 ± 0.8 | 6.0 ± 1.0 | 2.9 ± 1.4 | |
| Burst incidence IM | 4.2 ± 1.2 | 5.0 ± 0.9 | 4.9 ± 0.9 | 4.5 ± 1.3 | 5.4 ± 1.0 | 3.1 ± 1.3 | 5.3 ± 1.1 | 3.5 ± 0.9 | |
| CA1d theta (5 – 12 Hz) | LFP measurement | SO/SP | SP | upper SP | SP | SP | SP | SP | SP |
| Preferred theta phase | 159° | 176° | 152° | 167° | 177° | 264° | 130° | 233° | |
| Mean vector length (r) | 0.37 | 0.50 | 0.23 | 0.27 | 0.51 | 0.35 | 0.44 | 0.34 | |
| Rayleigh P-value | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | |
| Total n spikes | 1081 | 2679 | 6659 | 5143 | 1743 | 1413 | 860 | 578 | |
| Spikes per cycle (mean ± s.d.) | 6.8 ± 3.1 | 6.7 ± 3.0 | 8.7 ± 4.2 | 7.2 ± 3.4 | 7.2 ± 3.1 | 4.6 ± 1.9 | 7.7 ± 3.3 | 8.1 ± 3.2 | |
Details of juxtacellular labeling.
Number of juxtacellular labeling attempts on the final recording day for each animal (neurobiotin is metabolized within 24 hr). Letters refer to individually recorded neurons. E.g. in animal TV58, recorded neurons ‘e’ and ‘g’ were selected for juxtacellular labeling; in animal TV50 only neuron ‘a’ was recorded and labeled. A ‘labeling attempt’ refers to the application of current pulses to a recorded neuron. Modulation refers to successful entrainment of action potentials to the duration of positive current pulses (Pinault, 1996). Strength and duration of modulation are also estimated in order to predict the labeling strength (Table 1).
| Animal name | Labeling attempts from single neurons | Modulated neurons | Recovered neurons |
|---|---|---|---|
| TV58 | 2 (e, g) | 1 (g, very strong, >5 min; rostral of e) | 1 (g) |
| TV50 | 1 (a) | 1 (a, strong, 77 s) | 1 (a, damaged) |
| MS14 | 1 (i) | 1 (i, strong, 1 min) | One strong (i); one weak, dorsal of i, dendrites cross with i, potentially gap-junction coupled |
| MS19 | 1 (b) | 1 (b, strong, 30 s) | 1 (b) |
| AJ50 | 4 (g-j) | 3 (g, strong, 6 s, lost; h strong 30 s, dorsal of g; j, strong, 2 min, rostral of h) | Two strong (h, j); one very weak; dorsal of h, potentially gap-junction coupled |
| AJ44 | 1 (j) | 1 (j, strong, 12 s) | 1 (j) |
| MS74 | 1 (b) | 1 (b, strong, 2 min) | 1 (b, damaged) |
| MS17 | 1 (k) | 1 (k, strong, 30 s, lost cell after) | 1 (k) |
Firing patterns of identified orchid cells.
Abbreviations: u, unknown/unavailable. Spike burst defined as > 3 spikes with ISIs < 40 ms. Intraburst frequency (Hz) is expressed as mean ± s.d.; burst duration and interburst interval (ms) as median and interquartile range. LM, locomotion; SM, small movements including limbs, tail, and shifts in posture; WS, high-frequency whisking and/or sniffing in the absence of other movements; IM, immobility. KS, Kolmogorov-Smirnov.
| Cell name | TV58g | TV50a | MS14i | MS19b | AJ50j | AJ50h | AJ44j | MS74b | |
|---|---|---|---|---|---|---|---|---|---|
| Behavioral states | Burst duration LM | 87.0, 79.8 | 66.8, 30.5 | 121.0, 193.0 | 96.2, 57.5 | 63.2, 22.6 | 70.1, 36.3 | 68.2, 48.9 | 80.1, 40.1 |
| Burst duration SM | 86.7, 47.5 | 71., 42.0 | 84.2, 56.7 | 85.7, 53.4 | 54.0, 23.8 | 65.2, 56.3 | 70.1, 38.3 | u | |
| Burst duration WS | 91.8, 77.1 | u | 91.7, 61.5 | 92.0, 56.8 | 59.2, 30.2 | 68.3, 35.5 | 52.4, 30.3 | 91.5, 91.9 | |
| Burst duration IM | 94.6, 65.5 | 70.2, 46.0 | 81.1, 48.9 | 100.0, 57.3 | 57.8, 33.3 | 73.9, 33.7 | 68.6, 42.7 | 97.0, 75.0 | |
| Two-sample KS test P-value burst duration LM-IM | 0.5928 | 0.0021 | <0.0001 | 0.6573 | 0.0003 | 0.3998 | 0.1947 | 0.0293 | |
| Interburst interval LM | 187.7, 80.9 | 146.0, 35.2 | 179.6, 162.5 | 193.5, 91.8 | 135.8, 41.3 | 160.0, 148.9 | 129.0, 52.1 | 155.3, 86.4 | |
| Interburst interval SM | 214.2, 81.2 | 159.9, 49.3 | 178.7, 99.1 | 230.1, 166.5 | 159.9, 66.1 | 257.6, 209.6 | 158.2, 67.6 | u | |
| Interburst interval WS | 205.1, 100.3 | u | 189.9, 105.1 | 255.4, 180.0 | 178.1, 137.8 | 291.7, 315.5 | 157.3, 73.9 | 254.5, 229.1 | |
| Interburst interval IM | 203.3, 94.5 | 177.3, 92.3 | 180.3, 87.5 | 238.3, 161.8 | 163.3, 66.2 | 255.8, 201.0 | 161.5, 61.7 | 249.0, 135.2 | |
| Two sample KS test P-value interburst interval LM-IM | 0.0725 | <0.0001 | 0.0039 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | |
| Intraburst frequency LM | 95.9 ± 22.1 | 111.4 ± 24.8 | 95.8 ± 24.4 | 83.7 ± 24.6 | 136.1, 27.2 | 75.9 ± 23.6 | 131.4 ± 30.9 | 95.2 ± 20.2 | |
| Intraburst frequency SM | 84.9 ± 19.5 | 105.7 ± 24.5 | 99.7 ± 30.0 | 82.8 ± 27.5 | 121.0, 27.2 | 79.2 ± 17.7 | 96.0 ± 24.5 | u | |
| Intraburst frequency WS | 96.6 ± 24.3 | u | 93.1 ± 24.3 | 80.7 ± 22.7 | 110.6, 27.4 | 71.8 ± 10.2 | 96.4 ± 13.6 | 75.6 ± 16.1 | |
| Intraburst frequency IM | 94.7 ± 22.8 | 113.1 ± 34.0 | 100.6 ± 28.0 | 77.6 ± 19.7 | 124.0, 32.6 | 78.1 ± 16.8 | 89.3 ± 18.2 | 79.3 ± 22.9 | |
| Two sample KS test P-value intraburst freq LM-IM | 0.3102 | 0.0693 | 0.0357 | 0.0180 | <0.0001 | 0.0602 | <0.0001 | <0.0001 | |
| CA1d mid-gamma (55 – 80 Hz) | Preferred gamma phase | u | u | 164° | 158° | u | 191° | u | u |
| Mean vector length (r) | u | u | 0.04 | 0.25 | u | 0.19 | u | u | |
| Rayleigh P-value | u | 0.0860 | 0.0008 | <0.0001 | 0.6225 | <0.0001 | 0.0799 | 0.4700 | |
| Total n spikes | u | 1240 | 3553 | 2052 | 1445 | 398 | 579 | 437 | |
| CA1d SWRs (130 – 230 Hz) | n SWRs | 34 | 40 | 57 | 58 | 0 | 0 | 0 | 16 |
| Mean rate inside (Hz) | 57.08 | 31.37 | 57.71 | 4.88 | u | u | u | u | |
| Lambda rate outside (Hz) | 44.82 | 49.14 | 48.96 | 33.16 | u | u | u | u | |
| Poisson P-value | 0.0181 | 0.0015 | 0.0238 | <0.0001 | u | u | u | u | |
Immunohistochemical profiles of orchid neurons.
Positive (+) or negative (-) immunoreactivity observed within subcellular domain: s, soma; n, nucleus; d, proximal dendrite; a, axon; t, axon terminals; u, unknown (unavailable or inconclusive). *Soma was not recovered; dendrites were beaded indicative of a cell damaged during labeling. **No axon was recovered.
| Cell name | TV58g | TV50a* | MS14i | MS19b | AJ50h | AJ50j | AJ44j | MS74b* | MS17k** | |
|---|---|---|---|---|---|---|---|---|---|---|
| Immunoreactivity | PV | ad+ | a+ | ds+ | a+ | a+ | a+ | a+ | a+ | ds+ |
| SATB1 | n+ | u | n+ | n+ | n+ | n+ | n+ | u | n+ | |
| mGluR1a | d+ | u | d+ | s+ | ds+ | ds+ | ds+ | u | ds+ | |
| VGAT | t+ | u | u | u | t+ | u | u | u | u | |
| NK1R | d- | u | u | s- | u | s- | s- | u | ds+ | |
| PCP4 | d+ | a+ | ds+ | u | u | u | u | u | u | |
| SMI32 | d+ | u | u | u | u | u | u | u | u | |
| Calbindin | d- | ad- | d- | ad- | u | u | u | a- | d- | |
| Calretinin | d- | ad- | d- | ad- | u | u | u | u | d- | |
| ChAT | sd- | a- | u | u | u | u | u | u | d- | |
| nNOS | sd- | u | u | u | u | u | u | u | u | |
Marker genes used to define sub-regions of the temporal cortex.
https://doi.org/10.7554/eLife.34395.018| Marker gene | Gene product | Expression profile | Reference |
|---|---|---|---|
| Amigo2 | Adhesion molecule with Ig like domain 2 | CA2, FC, temporal CA3 | http://mouse.brain-map.org/experiment/show/71250310 |
| Etv1 | Ets variant 1 (ER81) | SUBd, RSg | http://mouse.brain-map.org/experiment/show/72119595 |
| Nos1 | Neuronal nitric oxide synthase 1 | PrS | http://mouse.brain-map.org/experiment/show/75147762 |
| Satb2 | Special AT-rich sequence binding protein 2 | CA1, RSg, PrS | http://mouse.brain-map.org/experiment/show/73992708 |
| Slc17a6 | solute carrier family 17 member 6 (VGLUT2) | SUBd, RSg, PrS | http://mouse.brain-map.org/experiment/show/73818754 |
| Tmem178 | Transmembrane protein 178 | CA2, FC, CA3, EC layer 6 | http://mouse.brain-map.org/experiment/show/73992709 |
| Wfs1 | Wolfram syndrome one homolog (Wolframin) | CA1d, PaS | http://mouse.brain-map.org/experiment/show/74881161 |
Cortical target neurons of orchid cell TV58g.
Molecular profiles of presumed postsynaptic neurons, based on close apposition of medial septal terminals; +, detectable positive immunoreactivity or signal; –, undetectable immunoreactivity or signal in vicinity of immunopositive signals; u, unknown (unavailable or inconclusive). Parentheses indicate weak immunoreactivity or signal. Endo-biotin, endogenous biotin (detected with streptavidin-conjugated fluorophore) may indicate high metabolic activity due to a high density of mitochondria, such as in ‘fast spiking’ neurons.
| Target ID | Location | PV | SATB1 | mGluR1a | Endo-biotin | CCK | CB | nNOS | Figure |
|---|---|---|---|---|---|---|---|---|---|
| S45A | FC | – | + | + | – | – | – | – | |
| S45B | u | + | u | – | – | u | – | ||
| S45D | + | + | – | + | – | – – | + | ||
| S45G | + | + | – | + | – | + | |||
| S44A | RSg L6 | – | + | + | – | – | – | – | 4d-f |
| S44B | u | + | u | – | – | u | – | ||
| S38A | PrSd L5-6 | + | + | + | + | – | – | – | 4b |
| S38C | + | + | u | + | – | – | – | ||
| S38I | u | + | + | (+) | – | u | + | ||
| S38J | u | + | u | – | – | + | + | ||
| S36A | + | + | u | + | u | u | u | ||
| S34A | + | + | u | + | u | u | – | ||
| S34C | + | + | – | (+) | u | u | + | ||
| Total + | 7 | 13 | 4 | 8 | 0 | 1 | 5 | ||
Details of mouse behavior.
LM, locomotion; SM, small postural movements; WS, theta-frequency whisking and/or sniffing without small movements; IM, immobility. *LM periods for these recordings were classified based on high amplitude long-duration (>1 s) signals from the accelerometer with high theta power, and short-duration low amplitude accelerometer signals as SM; those periods outside movement were classed as IM, which may include some whisking/sniffing. Training: limited (limited exposure to head-fixation prior to recording the cell, unfamiliar environment); moderate (typical exposure, familiar with environment); fully trained (fully habituated, familiar with environment and head-fixation). Mouse activity is ordered from the most to least frequent.
| Animal name | Cell ID | Recording conditions | Training | Monitoring | Mouse activity |
|---|---|---|---|---|---|
| TV58 | TV58g | Circular treadmill | Moderate | Video, EMG, accelerometer | IM, WS, SM, LM |
| TV50 | TV50a | Circular treadmill | Moderate | Accelerometer, no video* | IM/(WS), LM, SM |
| MS14 | MS14i | Circular treadmill | Moderate | Video, wheel movement | IM, SM, WS, LM |
| MS19 | MS19b | Circular treadmill | Limited | Video, wheel movement | SM, LM, WS, IM |
| AJ50 | AJ50j | Circular treadmill | Fully trained | Video, wheel movement | IM, WS, SM, LM |
| AJ50h | Circular treadmill | Fully trained | Video, wheel movement | LM, IM, SM, WS | |
| AJ44 | AJ44j | Circular treadmill | Fully trained | Video, wheel movement | IM, SM, LM, WS |
| MS74 | MS74b | Frisbee (with netting) | Fully trained | Video, accelerometer | IM, LM, WS |
| TV77 | TV77q | Circular treadmill | Moderate | Video, wheel movement | IM, WS, SM, LM |
| TV78 | TV78d | Circular treadmill | Moderate | Video, wheel movement | IM/(WS), LM, SM. Whiskers not in focus in video. |
| TV78k | Circular treadmill | Moderate | Video, wheel movement | IM, SM, LM, WS | |
| TV78l | Circular treadmill | Moderate | Video, wheel movement | IM, SM, (LM), (WS) Whiskers not in focus in video. Slow LM. | |
| MS17 | MS17k | Circular treadmill | Moderate | Video, wheel movement | IM, LM, SM, WH |
| MS58 | MS58i | Running disc | Moderate | Accelerometer, no video* | IM/(WS), SM, LM |
| MS24 | MS24a | Circular treadmill | Moderate | Video, wheel movement | IM, LM, WS, SM |
| MS84 | MS84f | Frisbee (with paper) | Fully trained | IM, LM, SM, WS | |
| TV72 | TV72n | Circular treadmill | Limited | Video, wheel movement | IM, SM, WS, LM. 50% of LM was involuntary. |
| TV85 | TV85e | Running disc | Limited | Accelerometer, video | IM, WS, SM, LM. 66% of LM was involuntary. |
| TV86 | TV86b | Running disc | Moderate | Accelerometer, video | IM, WS, LM, SM |
Behavior-dependent firing patterns of putative orchid cells.
*Labeled soma observed in the dorsal MS; only a faint projection axon was recovered. **Destroyed soma observed in the dorsal MS; no axon was recovered. Abbreviations: u, unknown/unavailable. Spike burst defined as > 3 spikes with ISIs < 40 ms. Firing rates and burst incidence (Hz) are expressed as mean of 1 s bins ± s.d.; intraburst frequency (Hz) as mean ± s.d.; burst duration and interburst interval (ms) as median and interquartile range. LM, locomotion; SM, small movements including limbs, tail, and shifts in posture; WS, high-frequency whisking and/or sniffing in the absence of other movements; IM, immobility. KS, Kolmogorov-Smirnov.
| Cell name | MS17k* | MS58i* | MS24a | MS84f | TV78d | TV78k | TV85e | TV86b |
|---|---|---|---|---|---|---|---|---|
| Max speed (cm/s) | 9.9 | u | 16.8 | u | 14.3 | 4.5 | u | u |
| Firing rate LM | 51.6 ± 25.5 | 80.9 ± 20.2 | 48.7 ± 10.7 | 65.2 ± 7.5 | 31.5 ± 11.9 | 60.6 ± 12.5 | 87.6 ± 14.9 | 67.9 ± 9.1 |
| Firing rate SM | 51.8 ± 13.8 | 42.3 ± 9.1 | 50.1 ± 11.7 | 56.0 ± 10.9 | 41.6 ± 18.3 | 34.4 ± 12.1 | 47.3 ± 14.3 | 49.5 ± 6.4 |
| Firing rate WS | 46.7 ± 13.0 | 52.1 ± 14.7 | 44.2 ± 11.5 | 48.9 ± 11.6 | u | 30.3 ± 15.5 | 50.1 ± 10.1 | 55.1 ± 9.3 |
| Firing rate IM | 44.6 ± 13.8 | 52.1 ± 14.7 | 33.7 ± 8.8 | 48.4 ± 9.2 | 39.5 ± 9.8 | 33.6 ± 9.6 | 58.0 ± 13.0 | 37.0 ± 9.7 |
| Burst incidence LM | 3.3 ± 2.2 | 6.6 ± 1.0 | 5.3 ± 1.4 | 5.1 ± 1.5 | 3.6 ± 1.3 | 3.6 ± 1.0 | 7.0 ± 1.4 | 5.3 ± 1.2 |
| Burst incidence SM | 4.7 ± 1.0 | 4.9 ± 1.4 | 4.0 ± 0.9 | 4.6 ± 1.4 | 4.0 ± 1.1 | 3.3 ± 0.9 | 5.2 ± 1.5 | 5.0 ± 0.0 |
| Burst incidence WS | 4.7 ± 1.3 | u | 3.4 ± 1.0 | 4.8 ± 1.3 | u | 2.8 ± 1.0 | 5.4 ± 1.1 | |
| Burst incidence IM | 4.3 ± 1.2 | 4.9 ± 1.1 | 3.3 ± 1.2 | 4.8 ± 1.1 | 3.8 ± 1.0 | 3.3 ± 1.0 | 5.3 ± 1.1 | |
| Burst duration LM | 86.7, 115.8 | 70.2, 22.0 | 57.4, 36.8 | 86.4, 111.8 | 76.9, 39.4 | 156.3, 101.6 | 64.1, 34.0 | 89.6, 62.7 |
| Burst duration SM | 90.8, 84.8 | 67.3, 47.2 | 107.0, 84.0 | 99.2, 77.6 | 90.0, 57.6 | 112.9, 70.2 | 62.5, 25.9 | 101.2, 77.0 |
| Burst duration WS | 82.9, 74.0 | 76.7, 57.4 | 97.3, 65.1 | 96.7, 93.0 | u | 118.3, 50.0 | 62.1, 43.4 | 96.5, 70.3 |
| Burst duration IM | 84.4, 67.7 | 76.7, 57.4 | 94.1, 73.0 | 94.3, 69.9 | 95.3, 72.1 | 112.6, 72.3 | 65.9, 45.1 | 75.8, 43.4 |
| Two sample KS test P-value burst duration LM-IM | 0.3902 | 0.0024 | <0.0001 | 0.0294 | <0.0001 | <0.0001 | 0.5547 | 0.0045 |
| Interburst interval LM | 166.6, 155.2 | 133.1, 31.1 | 155.6, 96.5 | 147.4, 115.9 | 197.8, 163.9 | 233.8, 155.6 | 133.0, 32.1 | 163.1, 91.8 |
| Interburst interval SM | 190.7, 135.2 | 169.8, 117.3 | 236.9, 138.7 | 178.1, 86.8 | 206.6, 136.8 | 267.4, 163.5 | 163.1, 73.2 | 181.8, 84.8 |
| Interburst interval WS | 197.9, 95.7 | u | 259.9, 194.5 | 189.1, 92.9 | u | 311.2, 299.0 | 159.7, 79.0 | 185.4, 95.3 |
| Interburst interval IM | 201.8, 117.9 | 170.7, 100.1 | 260.7, 186.0 | 187.2, 88.1 | 238.1, 136.9 | 267.6, 137.6 | 159.9, 91.8 | 194.4, 113.2 |
| Two sample KS test P-value inter-burst interval LM-IM | 0.0484 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.0130 | <0.0001 | <0.0001 |
| Intraburst frequency LM | 103.4 ± 35.6 | 141.9, 24.7 | 144.3, 46.8 | 105.1, 23.8 | 83.3, 22.9 | 81.8, 17.9 | 150.2, 32.8 | 119.4, 32.4 |
| Intraburst frequency SM | 94.2 ± 28.5 | 101.9, 26.1 | 104.4, 35.8 | 89.3, 27.0 | 87.4, 22.8 | 68.0, 14.6 | 117.9, 28.2 | 96.3, 24.6 |
| Intraburst frequency WS | 95.6 ± 38.8 | u | 97.8, 25.9 | 88.9, 24.2 | u | 70.8, 17.8 | 118.2, 26.9 | 105.0, 26.8 |
| Intraburst frequency IM | 95.0 ± 27.3 | 101.9, 30.3 | 82.5, 27.4 | 86.9, 22.7 | 89.4, 27.2 | 72.4, 19.8 | 135.5, 39.0 | 96.3, 25.4 |
| Two sample KS test P-value intraburst frequency LM-IM | 0.0914 | <0.0001 | <0.0001 | <0.0001 | 0.0449 | <0.0001 | 0.0003 | <0.0001 |
Network state-dependent firing patterns of putative orchid cells.
*Labeled soma observed in the dorsal MS; only a faint projection axon was recovered. **Destroyed soma observed in the dorsal MS; no axon was recovered. Abbreviations: u, unknown/unavailable.
| Cell name | MS17k* | MS58i** | MS24a | MS84f | TV78d | TV78k | TV85e | TV86b | |
|---|---|---|---|---|---|---|---|---|---|
| CA1d theta (5 – 12 Hz) | LFP measurement | SP | SO/SP | SO/SP | Alveus | SP | SP | SP | SP |
| Pref. theta phase | 176° | 133° | 178° | 177° | 261° | 163° | 160° | 243° | |
| Mean vector length (r) | 0.29 | 0.42 | 0.43 | 0.31 | 0.38 | 0.26 | 0.41 | 0.30 | |
| Rayleigh P-value | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | |
| Total n spikes | 727 | 2869 | 3287 | 4624 | 2154 | 1291 | 2792 | 1292 | |
| Spikes per cycle (mean ± s.d.) | 9.0 ± 3.7 | 7.6 ± 3.9 | 6.8 ± 3.9 | 8.8 ± 2.7 | 5.3 ± 3.4 | 7.5 ± 4.1 | 7.3 ± 4.4 | 9.6 ± 3.6 | |
| CA1d mid-gamma (55 – 80 Hz) | Pref. gamma phase | u | u | 155° | u | 144° | u | 207° | u |
| Mean vector length (r) | u | u | 0.3210 | u | 0.12 | u | 0.04 | u | |
| Rayleigh P-value | 0.1919 | 0.6089 | <0.0001 | 0.0870 | <0.0001 | 0.3506 | 0.0417 | 0.4627 | |
| Total n spikes | 1003 | 1032 | 284 | 1227 | 871 | 1457 | 2000 | 1274 | |
| CA1d SWRs (130 – 230 Hz) | n SWRs | 24 | 6 | 55 | 0 | 24 | 46 | 19 | 42 |
| Mean rate inside (Hz) | 69.35 | u | 27.47 | u | 3.75 | 19.67 | u | 4.48 | |
| Lambda rate outside (Hz) | 46.33 | u | 36.92 | u | 40.71 | 31.92 | u | 42.31 | |
| Poisson P-value | 0.0004 | u | 0.0176 | u | <0.0001 | 0.0068 | u | <0.0001 | |
Firing patterns of identified septo-hippocampal neurons.
(Related to Figure 5) Abbreviations: u, unknown/unavailable. Spike burst defined as > 3 spikes with ISIs < 40 ms. Firing rates and burst incidence are expressed as mean of 1 s bins ± s.d.; intraburst frequency as mean ± s.d.; burst duration and interburst interval as median and interquartile range. LM, locomotion; SM, small movements including limbs, tail, and shifts in posture; WS, high-frequency whisking and/or sniffing in the absence of other movements; IM, immobility.
| Cell name | TV77q | TV78l | |
|---|---|---|---|
| Behavioral states | Firing rate LM (Hz) | 38.4 ± 12.2 | 25.3 ± 1.7 |
| Firing rate SM (Hz) | 22.5 ± 7.6 | 16.7 ± 5.8 | |
| Firing rate WS (Hz) | 16.0 ± 7.5 | 23.3 ± 8.6 | |
| Firing rate IM (Hz) | 15.6 ± 8.1 | 21.6 ± 6.2 | |
| Burst incidence LM (Hz) | 3.7 ± 1.4 | 2.5 ± 0.6 | |
| Burst incidence SM (Hz) | 2.6 ± 0.9 | 1.5 ± 1.2 | |
| Burst incidence WS (Hz) | 1.5 ± 1.1 | 1.3 ± 1.2 | |
| Burst incidence IM (Hz) | 1.5 ± 1.2 | 1.8 ± 1.2 | |
| Burst duration LM (ms) | 69.7, 73.6 | 85.9, 96.2 | |
| Burst duration SM (ms) | 75.9, 40.5 | 64.9, 19.6 | |
| Burst duration WS (ms) | 74.3, 38.6 | 109.3, 102.4 | |
| Burst duration IM (ms) | 69.0, 48.1 | 86.3, 60.5 | |
| Interburst interval LM (ms) | 196.6, 206.7 | 537.7, 412.8 | |
| Interburst interval SM (ms) | 332.0, 250.9 | 457.3, 418.8 | |
| Interburst interval WS (ms) | 427.4, 588.3 | 427.3, 214.8 | |
| Interburst interval IM (ms) | 445.5, 615.9 | 393.4, 384.6 | |
| Intraburst frequency LM (Hz) | 85.3 ± 27.1 | 69.8 ± 35.9 | |
| Intraburst frequency SM (Hz) | 78.1 ± 28.3 | 65.6 ± 15.9 | |
| Intraburst frequency WS (Hz) | 82.9 ± 27.2 | 69.3 ± 13.3 | |
| Intraburst frequency IM (Hz) | 85.7 ± 31.1 | 64.1 ± 15.6 | |
| CA1d theta (5 – 12 Hz) | LFP measurement | SP | SP |
| Preferred theta phase | 230° | u | |
| Mean vector length (r) | 0.27 | u | |
| Rayleigh P-value | <0.0001 | 0.2819 | |
| Total n spikes | 1206 | 505 | |
| Spikes per cycle (mean ± s.d.) | 3.1 ± 2.7 | u | |
| CA1d SWRs(130 – 230 Hz) | n SWRs | 50 | 61 |
| Mean rate inside (Hz) | 6.20 | 0.65 | |
| Lambda rate outside (Hz) | 16.05 | 21.73 | |
| Poisson P-value | 0.0025 | <0.0001 | |
Specificity information for primary antibodies.
Rb, rabbit; Gt, goat; Ms, mouse; Gp, guinea pig; Ck, chicken.
| Molecule | Host | Dilution | Source | Specificity information | RRID |
|---|---|---|---|---|---|
| Calbindin (CB) | Rb | 1:5000 | Swant, CB-38 (lot 5.5) | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_2721225 |
| Calretinin (CR) | Rb | 1:500-1:1000 | Swant, 7699/3 hr (lot 18299) | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_10000321 |
| Calretinin (CR) | Gt | 1:1000 | Swant, CG1 | Table 1 in (Unal et al., 2015) | RRID:AB_10000342 |
| Choline acetyltransferase (ChAT) | Gt | 1:500 | Chemicon, AB144P | Table 1 in (Unal et al., 2015) | RRID:AB_2079751 |
| GFP | Ck | 1:500 | Aves Labs, GFP-1020 | Table 1 in (Unal et al., 2015) | RRID:AB_10000240 |
| Metabotropic glutamate receptor 1a (mGluR1a) | Gp | 1:500-1:1000 | Frontier Institute, mGluR1a-GP-Af660 | Characterized by Tanaka et al. 2000 Eur. J. Neurosci. 12, 781–792 | RRID:AB_2531897 |
| Neuronal nitrogen oxide synthase (nNOS) | Rb | 1:1000 | EMD Millipore, AB5380 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_91824 |
| Parvalbumin (PV) | Rb | 1:1000 | Swant, PV-28 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_2315235 |
| Parvalbumin (PV) | Gt | 1:1000 | Swant, PVG-214, lot 3.6 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_2313848 |
| Parvalbumin (PV) | Gp | 1:5000 | Synaptic Systems, 195 004, lot 5 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_2156476 |
| Purkinje cell protein 4 (PCP4) | Rb | 1:1000 | Santa Cruz, sc-74816, lot G0814 | Characterized by San Antonio et al. 2014 J. Comp. Neurol. 522, 1333–1354 | RRID:AB_2236566 |
| Pro-cholecystokinin (pro-CCK) | Rb | 1:500 | April 2005 gift (similar to Frontier Institute, CCK-pro-Rb-Af350) | Supplementary Table 2 in (Viney et al., 2013) | |
| SATB1 (N-14) | Rb | 1:200 | Abcam, ab70004 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_1270545 |
| SATB1 (N-14) | Gt | 1:200-1:250 | Santa Cruz, sc-5989 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_2184337 |
| SMI32 (neurofilament H non-phosphorylated) | Ms | 1:1000 | Covance, SMI-32R lot 14835102 | Similar to immunoreactivity characterized in primates by Campbell and Morrison 1989 J. Comp. Neurol. 282, 191–205 and in rats by Ouda et al. 2012 Brain Struc. Func. 217, 19–36 | RRID:AB_509997 |
| Substance P receptor (NK1R) | Rb | 1:500 | Millipore, AB5060, lot LV1525037 | Characterized by Shigemoto et al. 1993 Neurosci. Lett. 153, 157–160 | RRID:AB_2200636 |
| Vesicular GABA transporter (VGAT) | Gp | 1:500 | Synaptic Systems, 131 004 | Supplementary Table 2 in (Viney et al., 2013) | RRID:AB_887873 |
| Vesicular glutamate transporter 2 (VGLUT2) | Gp | 1:2000 | Synaptic Systems, 135 404 lot 135404/16 | Similar to immunoreactivity characterized in mouse hippocampus by Herzog et al. 2006 J. Neurochem. 99, 1011–1018 | RRID:AB_887884 |
| Wfs1 | Rb | 1:500 | Proteintech, 11558–1-AP | Table 1 in (Unal et al., 2015) | RRID:AB_2216046 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.34395.031





