Highly synergistic combinations of nanobodies that target SARS-CoV-2 and are resistant to escape
Figures
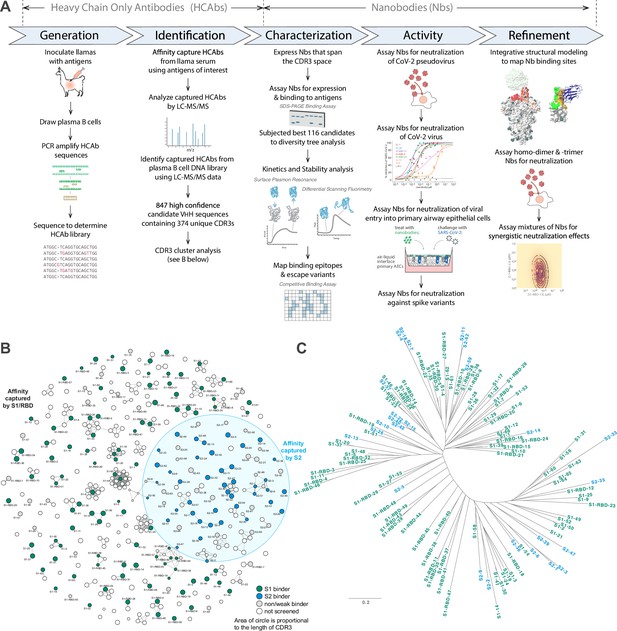
Approach.
(A) Schematic of our strategy for generating, identifying, and characterizing large, diverse repertoires of nanobodies that bind the spike protein of SARS-CoV-2. The highest quality nanobodies were assayed for their ability to neutralize SARS-CoV-2 pseudovirus, SARS-CoV-2 virus, and viral entry into primary human airway epithelial cells. We also measured the activities of homodimers/homotrimers and mixtures. (B) A network visualization of 374 high-confidence CDR3 sequences identified from the mass spectrometry workflow. Nodes (CDR3 sequences) were connected by edges defined by a Damerau–Levenshtein distance of no more than 3, forming 183 isolated components. A thicker edge indicates a smaller distance value, that is, a closer relation. (C) Dendrogram showing sequence relationships between the 116 selected nanobodies, demonstrating that the repertoire generally retains significant diversity in both anti-S1 (green) and anti-S2 (blue) nanobodies, albeit with a few closely related members. Scale, 0.2 substitutions per residue.
-
Figure 1—source data 1
Nanobody sequences.
- https://cdn.elifesciences.org/articles/73027/elife-73027-fig1-data1-v3.xlsx
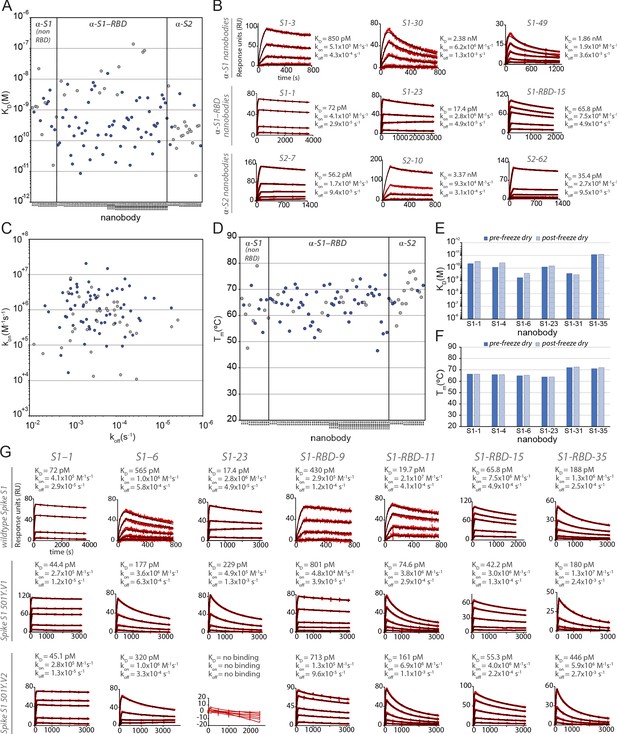
Biophysical characterization of anti-SARS-CoV-2 spike nanobodies.
(A) Each nanobody plotted against their affinity (KD) for their antigen separated into three groups based on their binding region on SARS-CoV-2 spike protein. The data points highlighted in blue correspond to nanobodies that neutralize. The majority of nanobodies have high affinity for their antigen with KDs below 1 nm. 10 nanobodies are not included in this plot as they were unable to be analyzed successfully using surface plasmon resonance (SPR). (B) SPR sensorgrams for each of the three targets on SARS-CoV-2 spike protein of our nanobody repertoire, showing three representatives for each binding region. (C) The association rate of each nanobody (kon) versus the corresponding dissociation rate (koff). The majority of our nanobodies have fast association rates (~10+5–10+7 M–1 s–1), with many surpassing the kon of high-performing monoclonal antibodies (~10+4–10+5 M–1 s–1). (D) Each nanobody plotted against their Tm as measured by differential scanning fluorimetry (DSF), revealing all but two nanobodies fall within a Tm range between 50 and 80°C, where the bulk of our nanobodies have a Tm ≥ 60°C. No data could be collected for two nanobodies, and 10 nanobodies exhibited two dominant peaks in the thermal shift assay and were not included in this plot (a full summary of this data can be seen in Tables 1–3). The KD (E) and Tm (F) of six nanobodies were assessed pre- and post-freeze-drying, revealing no significant change in affinity or Tm after freeze-drying. (G) SPR sensorgrams comparing the kinetic and affinity analysis of seven nanobodies against wild-type spike S1 (Wuhan strain), spike 20I/S1 501Y.V1 (alpha variant), and 20H/spike S1 501Y.V2 (beta variant).
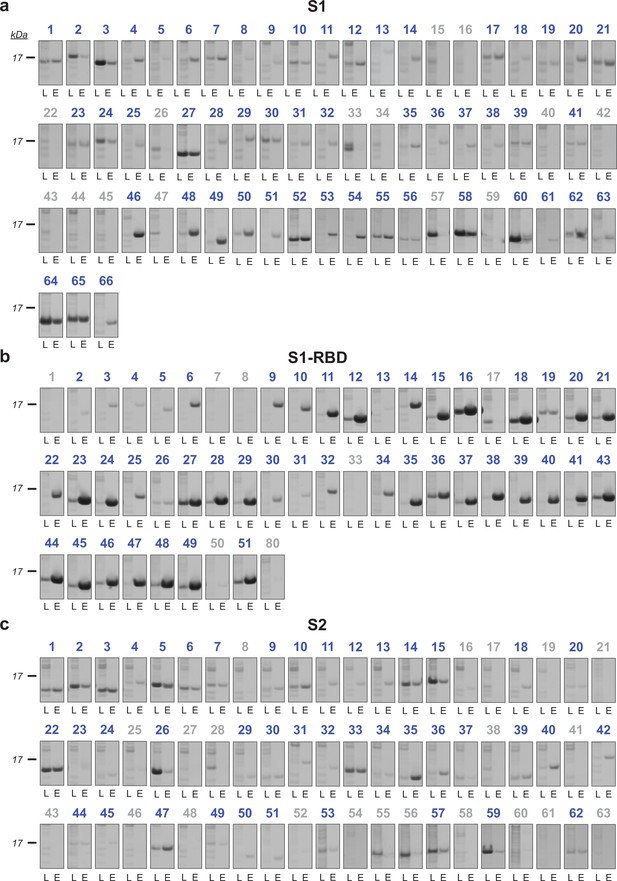
Binding of nanobody candidates to immobilized antigen.
All nanobody candidates identified by (a) S1, (b) S1-RBD, or (c) S2 purification were expressed in bacterial periplasm, which was bound to the respective immobilized antigen protein. After washes, loaded input (L) and elution (E) samples were analyzed by Coomassie stained SDS-PAGE. Positive binders (blue) displayed a nanobody band in the elution, while negative candidates (gray) had none.
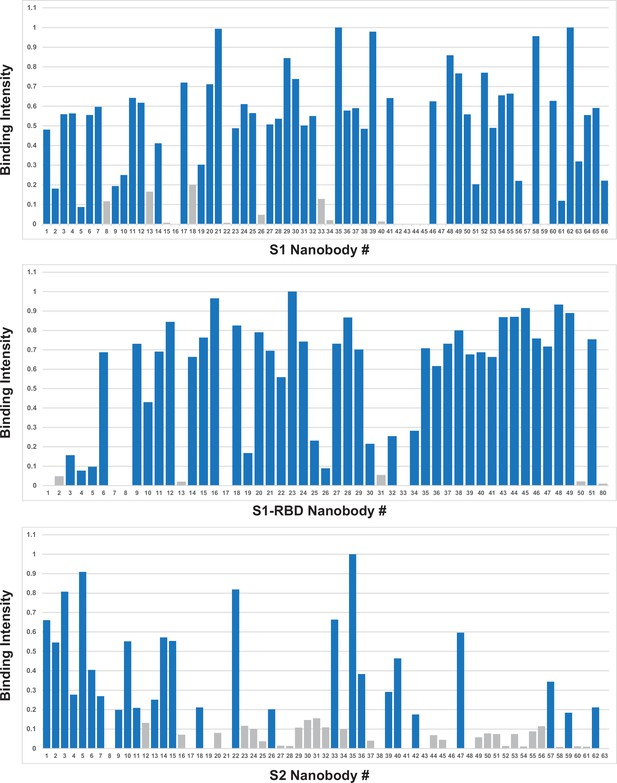
Quantified antigen binding of nanobody candidates.
All nanobody candidates were expressed in bacterial periplasm, which was bound to immobilized S1, S1-RBD, or S2 antigen protein. Bound nanobody was quantified by Coomassie staining after SDS-PAGE. Binding intensity against each antigen was normalized to the maximum observed binding among all nanobodies. Candidates with >20% maximum activity (blue) were selected for follow-up, while others (gray) were generally discarded.
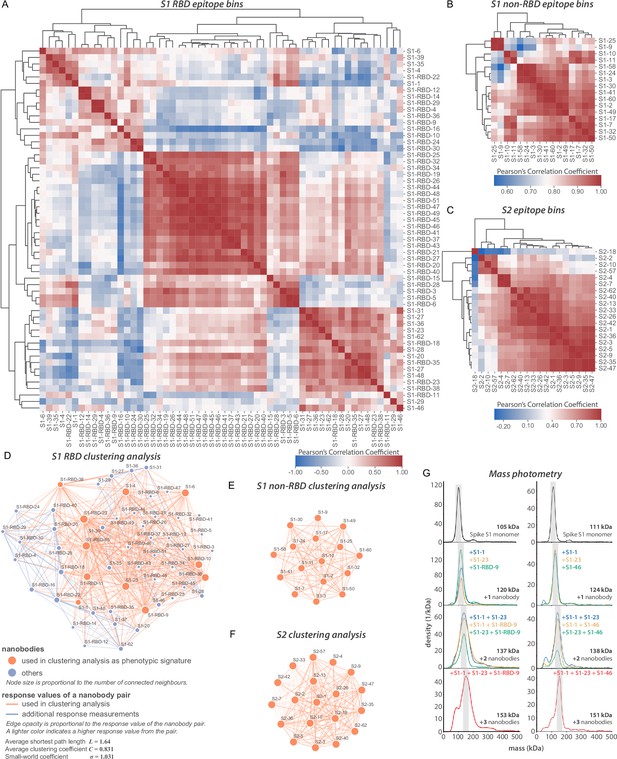
Epitope characterization of nanobodies against SARS-CoV-2 spike.
(A) Major epitope bins are revealed by a clustered heat map of Pearson’s correlation coefficients computed from the response values of nanobodies binding to the spike RBD in pairwise cross-competition assays on a biolayer interferometer. Correlated values (red) indicate that the two nanobodies respond similarly when measured against a panel of 11 RBD nanobodies that bind to distinct regions of the RBD. A strong correlation score indicates binding to a similar/overlapping region on the RBD. Anticorrelated values (blue) indicate that a nanobody pair responds divergently when measured against nanobodies in the representative panel and indicate binding to distinct or non-overlapping regions on the RBD. (B) As in (A), but for 16 S1 non-RBD-binding nanobodies. (C) As in (A), but for 19 S2-binding nanobodies. (D) A network visualization of anti-S1-RBD nanobodies. Each node is a nanobody and each edge is a response value measured by biolayer interferometry from pairwise cross-competition assays. Orange nodes represent 11 nanobodies used as a representative panel for clustering analysis in (A). Blue nodes represent the other nanobodies in the dataset. The average shortest distance between any nanobody pair in the dataset is 1.64. An average clustering coefficient of 0.831 suggests that the measurements are well distributed across the dataset. The small world coefficient of 1.031 indicates that the network is more connected than to be expected from random, but the average path length is what you would expect from a random network, together indicating that the relationship between nanobody pairs not actually measured can be inferred from the similar/neighboring nanobodies. (E, F) As in (D) but for S1 non-RBD and S2 nanobodies, respectively. These are complete networks with every nanobody measured against the others in the dataset. (G) Mass photometry (MP) analysis of spike S1 monomer incubated with different anti-spike S1 nanobodies. Two examples of an increase in mass as spike S1 monomers (black line) are incubated with 1–3 nanobodies. The accumulation in mass upon addition of each different nanobody on spike S1 monomer is due to each nanobody binding to non-overlapping space on spike S1, an observation consistent with Octet binning data. As a control, using MP, each individual nanobody was shown to bind spike S1 monomers on its own (data not shown).
-
Figure 3—source data 1
Normalized response values from epitope binning of nanobodies.
- https://cdn.elifesciences.org/articles/73027/elife-73027-fig3-data1-v3.xlsx
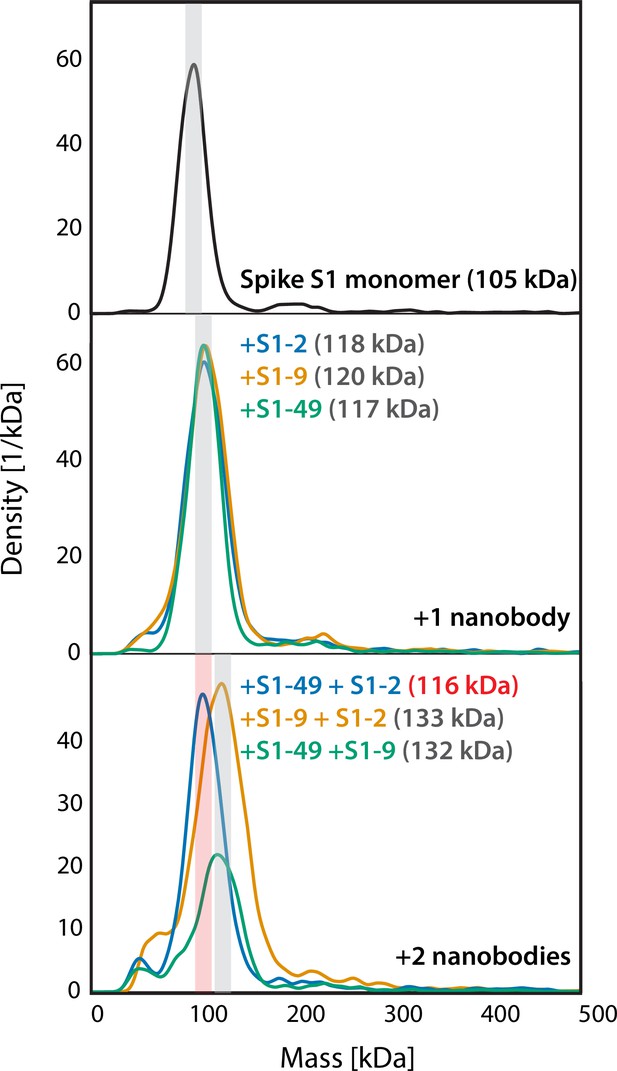
Mass photometry (MP) of non-RBD S1 nanobodies.
MP analysis of spike S1 monomer incubated with different non-RBD-binding anti-spike S1 nanobodies. There is either an accumulation in mass upon addition of each different nanobody on spike S1 monomer that is due to each nanobody binding to non-overlapping space on spike S1 or not, suggesting overlapping epitopes. These results, coupled with surface plasmon resonance (SPR) binning data, assisted the binning analysis for non-RBD anti-S1 nanobodies.
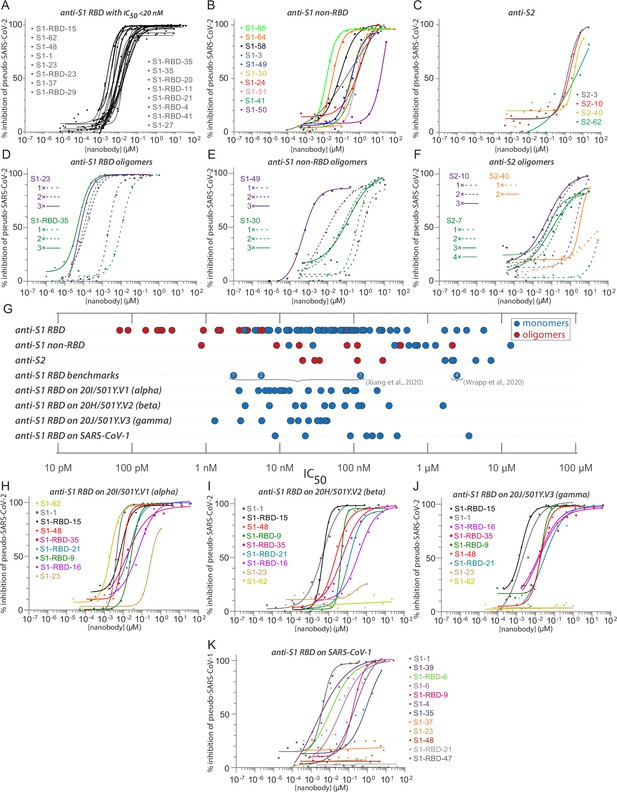
Diverse and potent nanobody-based neutralization of SARS-CoV-2.
Nanobodies targeting the S1-RBD, S1 non-RBD, and S2 portions of spike effectively neutralize lentivirus pseudotyped with various SARS-CoV spikes and their variants from infecting ACE2 expressing HEK293T cells. (A) Of the 116 nanobodies, monomers that neutralize SARS-CoV-2 pseudovirus with IC50 values 20 nM and lower are displayed. (B) Representative nanobodies targeting the non-RBD portions of S1 and (C) the S2 domain of SARS-CoV-2 neutralize SARS-CoV-2 pseudovirus. (D–F) Oligomerization of RBD, S1 non-RBD, and S2 nanobodies significantly increases neutralization potency. (G) Summary scatter plot of all nanobody IC50s across the major domains of SARS-CoV-2 spike and where tested, across SARS-CoV-2 variant 20H/501Y.V2 and SARS-CoV-1. Representative published nanobodies were also tested in our neutralization assays and show similar potency towards SARS-CoV-2 pseudovirus. From Xiang et al., 2020: (1) Nb-21 (IC50 2.4 nM); (2) Nb-34 (IC50 5.6 nM); and (3) Nb-93 (IC50 123 nM). From Wrapp, 2020a: (4) VHH-72 (IC50 2.5 μM). (H–K) Representative SARS-CoV-2 RBD targeting nanobodies cross-neutralize the 20I/501Y.V1/alpha variant with H69-, V70-, Y144- amino acid deletions and N501Y, A570D, D614G, P681H, T716I, S982A, and D1118H amino acid substitutions in spike (H); the 20H/501Y.V2/beta variant with L18F, D80A, K417N, E484K, and N501Y amino acid substitutions in spike; (I) 20J/501Y.V3/gamma variant with L18F, T20N, P26S, D138Y, R190S, K417T, E484K, N501Y, D614G, H655Y, T1027I, and V1176F amino acid substitutions in spike (J); and SARS-CoV-1 spike pseudotyped lentivirus (K). In all cases, n ≥ 2 biological replicates of each nanobody monomer/oligomer with a representative biological replicate with n = 4 technical replicates per dilution are displayed. See also Table 1, Table 2, Table 4, and Table 5.
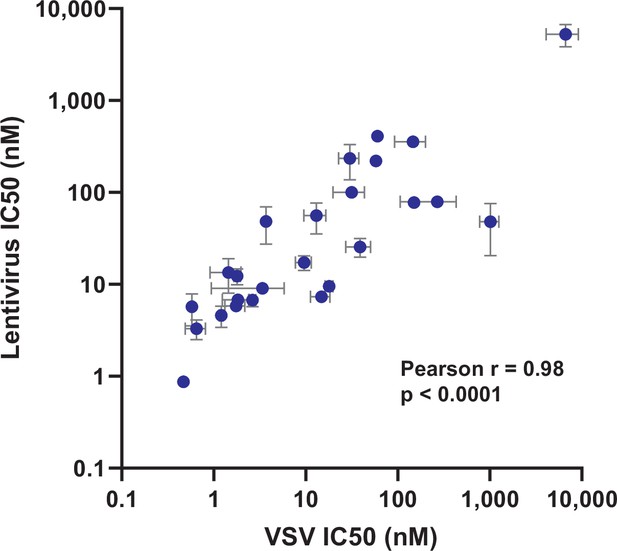
Correlation between pseudovirus assays.
IC50s of the 25 nanobodies tested in both lentiviral and VSV-based pseudovirus assays with wild-type SARS-CoV-2 spike were plotted to compare the values obtained from each system, showing a strong correlation.
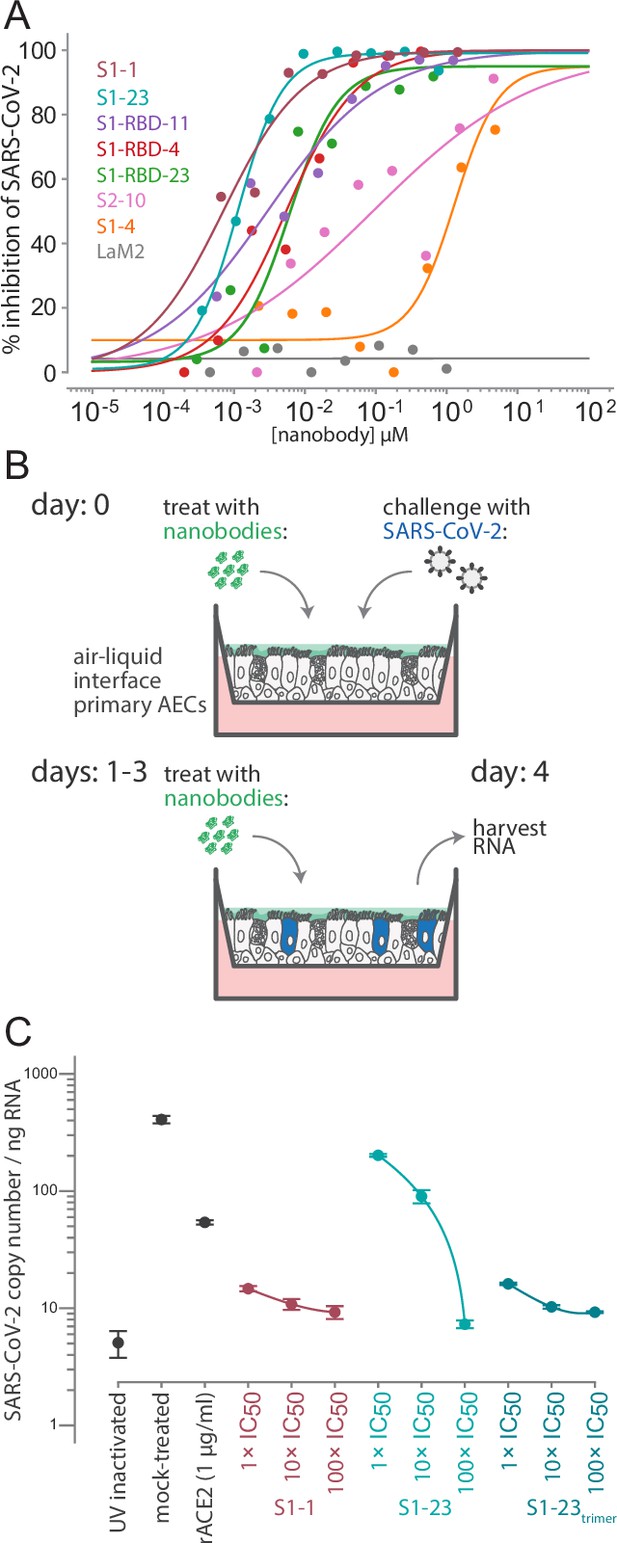
Authentic SARS-CoV-2 neutralization by anti-spike nanobodies.
(A) Nanobodies neutralize the authentic SARS-CoV-2 virus with similar kinetics as the SARS-CoV-2 pseudovirus. Neutralization curves are plotted from the results of a focus-forming reduction neutralization assay with the indicated nanobodies. Serial dilutions of each nanobody were incubated with SARS-CoV-2 (MOI 0.5) for 60 min and then overlaid on a monolayer of Vero E6 cells and incubated for 24 hr. LaM2, an anti-mCherry nanobody (Fridy et al., 2014a), was used as a non-neutralizing control. After 24 hr, cells were collected and stained with anti-spike antibodies and the ratio of infected to uninfected cells was quantified by flow cytometry. (B) A schematic of an air-liquid interface (ALI) culture of primary human airway epithelial cells (AECs) as a model for SARS-CoV-2 infection. Cells were incubated with nanobodies and then challenged with SARS-CoV-2 (MOI 0.5). After daily treatment with nanobodies for three more days, the cultures are harvested to isolate RNA and quantify the extent of infection. (C) Potent neutralization of authentic SARS-CoV-2 in AECs. The AECs were infected with the indicated concentrations of anti-SARS-CoV-2 spike nanobodies. The infected cultures were maintained for 5 days with a daily 1 hr incubation of nanobodies before being harvested for RNA isolation and determination of the SARS-CoV-2 copy number by qPCR. SARS-CoV-2 copy number was normalized to total RNA measured by spectrophotometry. Mock-treated samples exposed to infectious and UV-inactivated SARS-CoV-2 virions served as positive and negative controls. Recombinant soluble angiotensin converting enzyme 2 (rACE2) was used as a positive treatment control. The indicated nanobodies were used at 1, 10, and 100× their IC50 values determined in pseudovirus neutralization assays (Table 1 and Table 4).
-
Figure 5—source data 1
Neutralization data from authentic SARS-CoV-2 experiments.
- https://cdn.elifesciences.org/articles/73027/elife-73027-fig5-data1-v3.xlsx
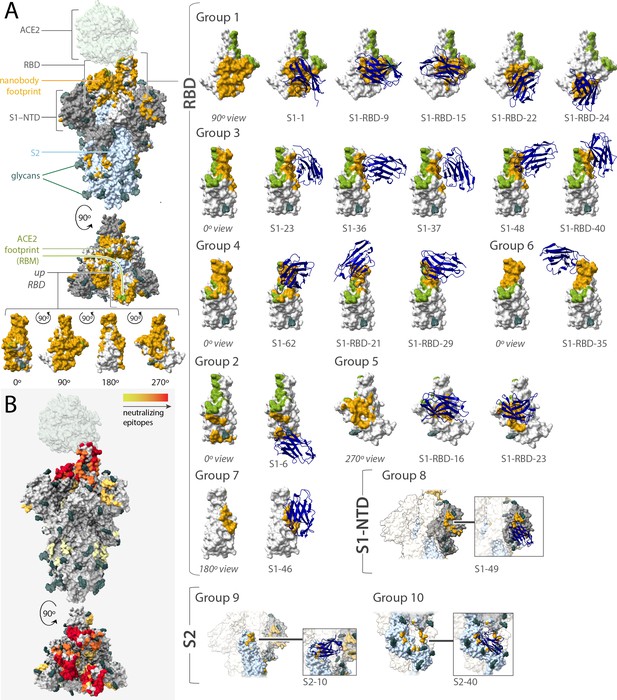
Epitope coverage of the 21 structural models of anti-spike SARS-CoV-2 nanobodies and neutralization potential of each epitope.
(A) The structure of SARS-CoV-2 full spike (PDB ID: 6VYB) solved via cryo-EM with one RBD in the up position overlaid with the crystal structure of RBD bound to ACE2 (PDB ID: 6M0J). Key elements of spike are colored as follows: RBD (white), S1-NTD (gray), and S2 (light blue). All 21 modeled nanobody footprints are colored gold on full spike with the ACE2 footprint (RBM) colored green. Full coverage of the 18 anti-RBD modeled nanobody footprints on RBD is seen in four different orientations. All 21 nanobodies are categorized into 10 groups based on their footprint on spike where groups 1–7 are anti-RBD nanobodies; group 8 contains an anti-S1-NTD nanobody and groups 9 and 10 contain anti-S2 nanobodies. (B) Heatmap of neutralizing epitopes on the structure of SARS-CoV-2 full spike (PDB ID: 6VYB). Epitopes are colored from pale yellow (epitopes with weak neutralization against SARS-CoV-2) to dark red (strong neutralization against SARS-CoV-2).
-
Figure 6—source data 1
PDB files of structural models of anti-spike SARS-CoV-2 nanobodies.
- https://cdn.elifesciences.org/articles/73027/elife-73027-fig6-data1-v3.zip

Mapping of spike substitutions in rVSV/SARS-CoV-2/GFP escape mutants obtained in the presence of the corresponding nanobody.
(A) Mapped on to the structure of SARS-CoV-2 spike trimer in complex with one ACE2 molecule (PDB ID: 7KNB, used for all SARS-CoV-2 spike trimer representations) is the position of neutralization-resistant amino acid substitutions (in red), also known as ‘escape mutants’ that were generated in response to cultivation of rVSV/SARS-CoV-2/GFP in the presence of each nanobody, and were subsequently shown to confer resistance to the same nanobody. (B) Structure of the SARS-CoV-2 RBD (PDB ID: 6M0J) showing the positions of amino acid residues (in green) that form the ACE2 binding site, for reference. (C) Structure of the SARS-CoV-2 RBD (PDB ID: 6M0J) showing the positions (in red) of the location of substitutions that confer resistance for each nanobody tested in two orientations 90° apart. For structure pairs of S1-RBD-16 and S1-RBD-23 escape mutants, the rotation is 90° from the structure to the left in the pair. (D) The location of four key non-RBD escape mutants S172G, Y170H, T315I, and S982R resulting from assays performed with the anti-S1 non-RBDs S1-49, S1-3, S1-30, and anti-S2 (S2-10) nanobodies, respectively. (E) Infectious rVSV/SARS-CoV-2/GFP yield (IU/ml) following two passages in the presence of the indicated individual nanobodies or nanobody combinations, at 100 IC50 of the individual nanobodies, or 50 IC50 of each of the nanobodies in the combinations. Each data point represents an independent titer measurement. Red open circles represent virus escapes while blue circles represent nanobody combinations for which no escapes (titer = 0) were detected. The location of two escape mutants K378Q (F) and Y508H (G) mapped onto SARS-CoV-2 spike trimer for the two corresponding nanobodies S1-RBD-9 and S1-RBD-15, respectively, revealing an exposed putative nanobody binding site on RBD when in the ‘up’ position that is hidden when RBD is in the ‘down’ position. (H) A close-up of escape mutant S172G on each monomer of SARS-CoV-2 spike trimer revealing a larger crevice between the NTD of spike S1 and RBD when the RBD is in the ‘down’ position compared to the ‘up’ position. (I) Three orientations of SARS-CoV-2 spike trimer revealing the position in all three orientations of escape mutant S982R revealing the putative binding site for nanobody S2-10 is accessible regardless of whether the RBD is the ‘up’ or ‘down’ position.
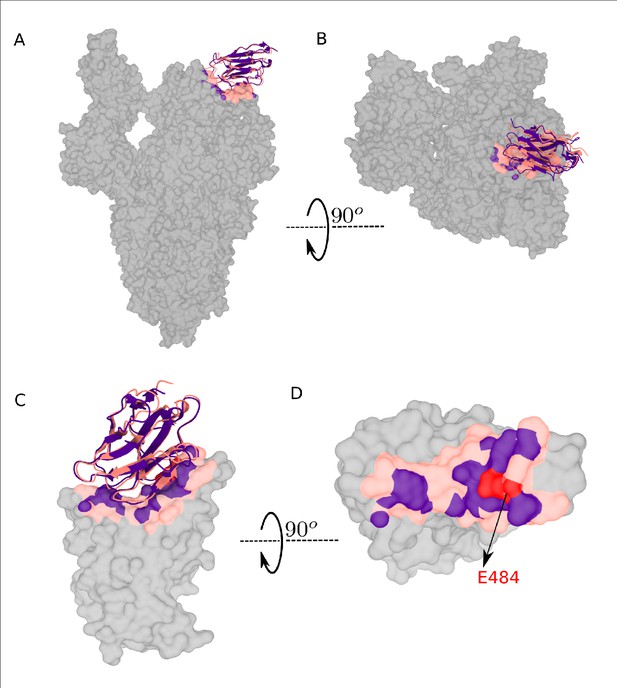
Comparison between RBD-Nb21 interface modeled with Integrative Modeling Platform (IMP) (red) with cryo-EM structure (7N9B) of the co-complex (indigo, taken from Sun et al., 2021).
IMP modeling was done using the 6M0J.E structure of the RBD (residues 333–526), which was subsequently aligned with the corresponding region of the down RBD from 7N9B.A, to compare the modeled and experimental binary complexes. (A) and (B) show the binding modes and the epitopes on the whole spike while (C) and (D) focus on just the 333–526 region of the RBD. Based on the observation that the point mutation E484K/Q significantly inhibits Nb21 binding (Sun et al., 2021), we treat E484 as an effective escape mutant for Nb21 to benchmark our computational model developed in IMP. Both the epitope (defined as all RBD residues within 6 Å of any Nb21 residue) and the binding mode (i.e., relative orientation) of Nb21 agree between the IMP model and the cryo-EM structure to within 2.1 Å backbone RMSD.
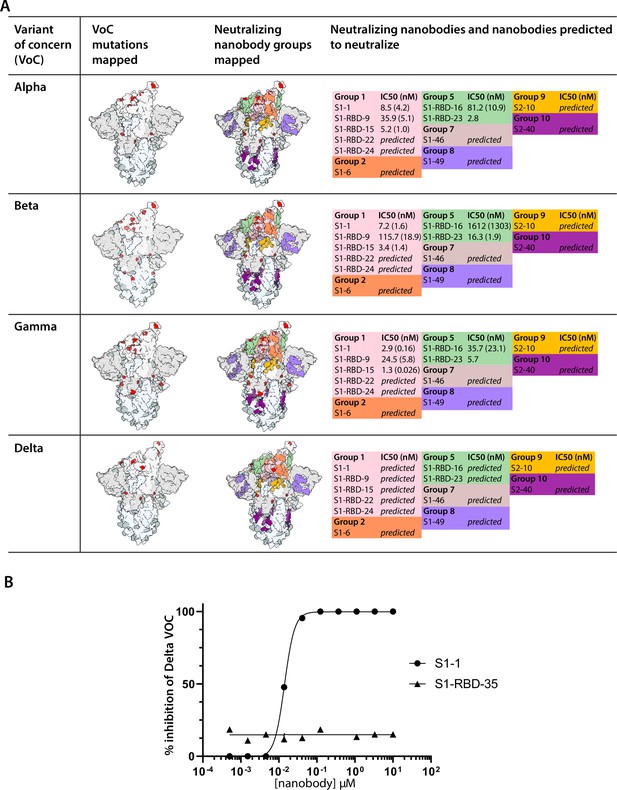
Nanobody groups resistant or predicted to be resistant against SARS-CoV-2 variants of concern (VOC).
(A) Color-coded and mapped on the structure of SARS-CoV-2 spike trimer with one RBD in the ‘up’ conformation (PDB ID: 7BNN) are the footprints of eight nanobody groups determined, which are mapped relative to the mutations found on four key SARS-CoV-2 VOC. Summarized are the nanobodies that make up each group and where applicable their degree of neutralization. The nanobody footprints are taken from the integrative modeling of these nanobodies on SARS-CoV-2 spike trimer (see Figure 6). (B) Neutralization of authentic SARS-CoV-2 isolate WA-1 and an isolate of the delta variant of concern (B.1.617.2) in a plaque reduction neutralization test assay. Serial dilutions of S1-1 and S1-RBD-35 were incubated with each SARS-CoV-2 variant for 60 min and then overlaid on a monolayer of Vero E6 cells and incubated for 90 min. Following infection, the cell cultures incubated for 48 hr before the number of visualized plaques were counted to determine the half maximal inhibitory concentration of each nanobody.
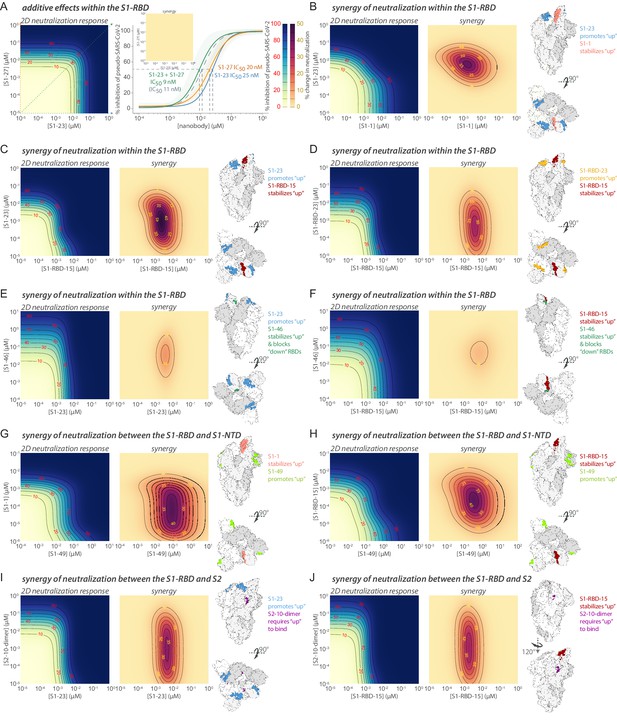
Synergistic neutralization of spike with nanobody cocktails.
(A) An example of additive effects between two anti-SARS-CoV-2 spike nanobodies. S1-23 and S1-27 were prepared in a two-dimensional serial dilution matrix and then incubated with SARS-CoV-2 pseudovirus for 1 hr before adding the mixture to cells. After 56 hr, the expression of luciferase in each well was measured by addition of Steady-Glo reagent and read out on a spectrophotometer. The left panel shows a heatmap of pseudovirus neutralization by a two-dimensional serial dilution of combinations of S1-23 and S1-27. Lines and red numbers demarcate the % inhibition, that is, inhibitory concentration where X% of the virus is neutralized, e.g., IC50. Dark blue regions are concentrations that potently neutralize the pseudovirus, as per the heatmap legend. The right panel shows neutralization curves (with 90% confidence interval bands) and the calculated IC50 of each nanobody alone, or in a 1:1 combination was determined along with a calculated IC50 based on the theoretical additive mixture model of the pair (curve with dotted gray line). The inset shows a difference (synergy) map calculated as the difference between the parameterized 2D neutralization response and that expected in a null model of only additive effects. Here, no difference is observed. (B) S1-1 synergizes with S1-23 in neutralizing SARS-CoV-2 pseudovirus. The left panel shows the heatmap of pseudovirus neutralization observed by a two-dimensional serial dilution of combinations of S1-1 and S1-23. The middle panel shows a heatmap mapping the synergy of neutralization observed for this pair. The lines bounding the darker purple areas demarcate regions in the heatmap where the observed neutralization is greater than additive by the indicated percentages (yellow numbers), as per the heatmap legend. The right panel shows two representations of spike with the accessible S1-1 (salmon) and S1-23 (steel blue) epitopes (PDB ID: 6VYB). (C–J) Examples of synergy between nanobodies binding the S1-RBD, or between the S1-RBD and S1-NTD or S2 domains of spike. The layout is as found in (B), but comparing S1-RBD-15 with S1-23 (C), S1-RBD-15 with S1-RBD-23 (D), S1-23 with S1-46 (E), S1-RBD-15 with S1-46 (F), S1-49 with S1-1 (G), S1-49 with S1-RBD-15 (H), S1-23 with S2-10-dimer (I), and S1-RBD-15 with S2-10-dimer (J).
-
Figure 7—source data 1
Neutralization data from synergy experiments.
- https://cdn.elifesciences.org/articles/73027/elife-73027-fig7-data1-v3.xlsx
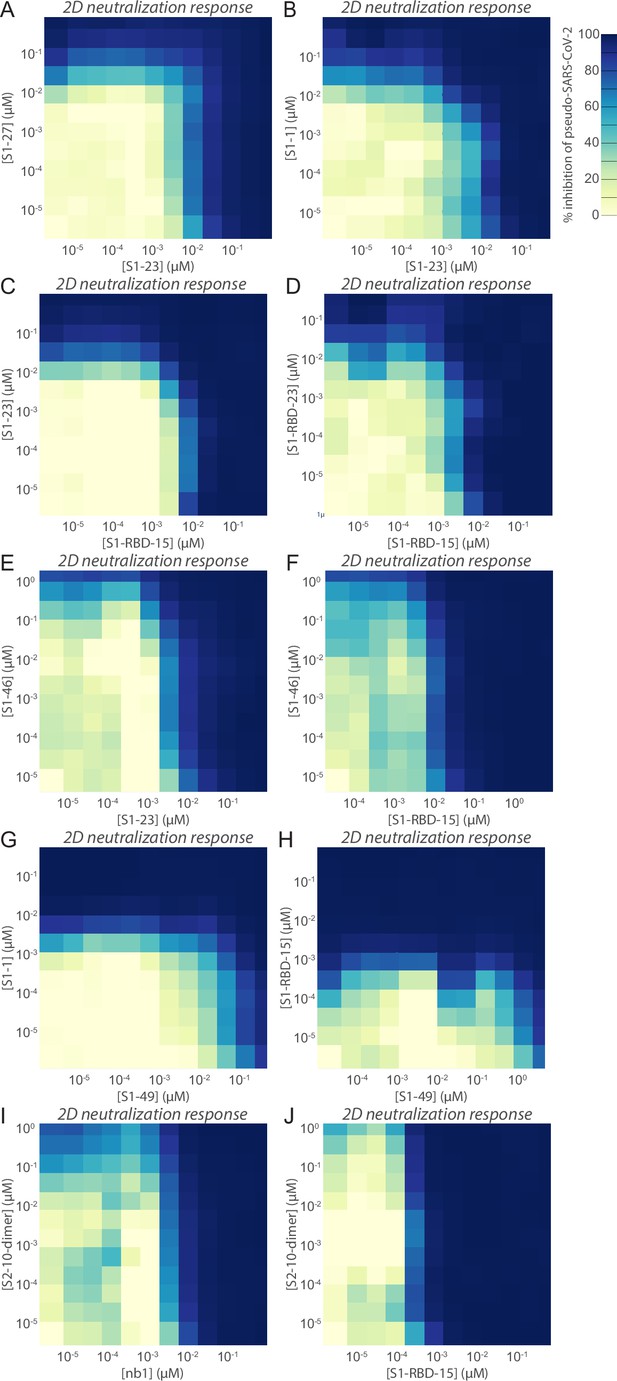
Heatmaps of nanobody synergy.
Neutralization of pseudovirus harboring the SARS-CoV-2 spike by pairs of nanobodies. The normalized % neutralization is visualized on a 2D grid of nanobody concentrations where each nanobody has been titrated in the background of the other tested nanobody. Nanobody concentrations are indicated on their respective axes for (A) S1-23 and S1-27, (B) S1-23 and S1-1, (C) S1-RBD-15 and S1-23, (D) S1-RBD-15 and S1-RBD-23, (E) S1-23 and S1-46, (F) S1-RBD-15 and S1-46, (G) S1-49 and S1-1, (H) S1-49 and S1-RBD-15, (I) S1-23 and S2-10-dimer, and (J) S1-RBD-15 and S2-10-dimer.
Tables
S1 nanobody characterization; related to Figures 2 and 4.
Nanobodies against S1 were determined to bind RBD or non-RBD epitopes by their affinity for recombinant full-length S1 and/or S1 RBD protein. Binding kinetics against these two recombinant proteins were determined by surface plasmon resonance (SPR), with on rates, off rates, and KDs determined by Langmuir fits to binding sensorgrams unless otherwise noted. Nanobody melting temperatures (Tm) were determined by differential scanning fluorimetry (DSF). Nanobodies were assayed for neutralization activity against a SARS-CoV-2 spike pseudotyped HIV-1 virus (PSV), with IC50s calculated from neutralization curves. Standard error of the mean (s.e.m.) is reported when available.
| ID | Epitope | S1 Kon(M–1 s–1) | S1 Koff(s–1) | S1 KD (M) | RBD Kon(M–1 s–1) | RBD Koff(s–1) | RBD KD (M) | Tm (°C) | SARS-CoV-2 PSV IC50 (s.e.m.)(nM) |
|---|---|---|---|---|---|---|---|---|---|
| S1-1 | RBD | 4.14E+05 | 2.98E-05 | 7.20E-11 | 6.50E+05 | 5.98E-07 | 9.20E-12 | 66.5 | 6.7 (1.0) |
| S1-2 | Non-RBD | 1.59E+06 | 1.88E-03 | 1.18E-09 | No interaction detected | 66.5 | NA | ||
| S1-3 | Non-RBD | 5.08E+05 | 4.32E-04 | 8.51E-10 | No interaction detected | 64 | 1030 (666) | ||
| S1-4 | RBD | 1.25E+06 | 1.06E-04 | 8.46E-11 | 1.26E+06 | 1.26E-04 | 1.37E-10 | 66 | 41.5 (3.7) |
| S1-5 | RBD | 1.33E+05 | 1.15E-03 | 8.61E-09 | – | 65.25 | NA | ||
| S1-6 | RBD | 1.02E+06 | 5.75E-04 | 5.65E-10 | 5.92E+05 | 3.69E-04 | 6.22E-10 | 65 | 56.1 (20.7) |
| S1-7 | Non-RBD | 7.59E+05 | 9.90E-04 | 1.30E-09 | No interaction detected | 60.5 | NA | ||
| S1-9 | Non-RBD | 9.51E+051.25E+05 | 4.28E-071.57E-04 | 4.50E-13*1.25E-09 | – | 47.5 | NA | ||
| S1-10 | Non-RBD | 8.35E+04 | 1.82E-03 | 2.19E-08 | – | 64 | NA | ||
| S1-11 | Non-RBD | – | – | 60 | NA | ||||
| S1-12 | RBD | 2.90E+05 | 8.92E-04 | 3.07E-09 | 2.33E+05 | 2.24E-04 | 9.63E-10 | 68 | NA |
| S1-14 | RBD | 1.08E+06 | 1.10E-03 | 1.02E-09 | 5.37E+05 | 7.99E-04 | 1.49E-09 | 57.5 | 135.8 (36.4) |
| S1-17 | Non-RBD | – | – | 65 | 1271 (888) | ||||
| S1-19 | RBD | 1.30E+063.55E+04 | 8.86E-032.41E-04 | 6.81E-09*6.81E-09 | – | 64.5 | 139 (9.6) | ||
| S1-20 | RBD | 1.48E+07 | 4.37E-03 | 2.95E-10 | – | 69 | 51.8 (3.7) | ||
| S1-21 | RBD | 4.77E+06 | 1.58E-04 | 3.31E-11 | 1.22E+06 | 2.45E-04 | 2.00E-10 | 70.5 | 226 (158) |
| S1-23 | RBD | 2.82E+06 | 4.91E-05 | 1.74E-11 | 1.09E+06 | 1.07E-04 | 9.78E-11 | 64 | 5.7 (2.2) |
| S1-24 | Non-RBD | 6.49E+05 | 2.89E-04 | 4.45E-10 | No interaction detected | 71.5 | 724 (144) | ||
| S1-25 | Non-RBD | 2.15E+05 | 3.39E-05 | 1.57E-10 | No interaction detected | 58 | NA | ||
| S1-27 | RBD | 3.15E+06 | 4.52E-04 | 1.43E-10 | 2.89E+06 | 6.30E-04 | 2.18E-10 | 54 | 19.5 (4.9) |
| S1-28 | RBD | 1.38E+06 | 7.97E-04 | 5.76E-10 | 1.79E+06 | 1.03E-03 | 5.77E-10 | 66 | 66.0 (10.9) |
| S1-29 | RBD | 2.39E+05 | 1.01E-03 | 4.21E-09 | 1.73E+05 | 8.89E+04 | 5.12E-09 | 61.5 | NA |
| S1-30 | Non-RBD | 6.21E+05 | 1.48E-03 | 2.38E-09 | No interaction detected | 57 | 717 (388) | ||
| S1-31 | RBD | 2.17E+06 | 5.63E-04 | 2.59E-10 | 1.94E+06 | 9.37E-04 | 4.84E-10 | 72 | 78.7 (3.5) |
| S1-32 | Non-RBD | 2.73E+05 | 4.66E-04 | 1.71E-09 | No interaction detected | 79 | NA | ||
| S1-35 | RBD | 2.46E+06 | 2.11E-05 | 8.60E-12 | 2.70E+06 | 9.77E-05 | 3.62E-11 | 70.5 | 12.5 (0.1) |
| S1-36 | RBD | 2.28E+06 | 3.92E-04 | 1.72E-10 | 7.87E+06 | 1.72E-03 | 2.18E-10 | 63 | 48.5 (21.1) |
| S1-37 | RBD | 4.03E+06 | 2.75E-04 | 6.82E-11 | 4.14E+06 | 2.09E-04 | 5.05E-11 | 65 | 6.8 (0.7) |
| S1-38 | RBD | 5.34E+06 | 1.12E-03 | 2.10E-10 | – | 64 | 66.1 (2.9) | ||
| S1-39 | RBD | 2.14E+06 | 8.11E-04 | 3.79E-10 | 1.68E+06 | 1.06E-03 | 6.30E-10 | 55 | 111 (4.0) |
| S1-41 | Non-RBD | 8.73E+05 | 1.38E-03 | 1.58E-09 | No interaction detected | 62.5 | 679 (53.4) | ||
| S1-46 | RBD | 1.68E+05 | 2.94E-04 | 1.75E-09 | 2.22E+05 | 1.70E-04 | 7.66E-10 | 68 | 312 (14.0) |
| S1-48 | RBD | 2.61E+06 | 6.22E-05 | 2.39E-11 | 1.66E+06 | 1.64E-04 | 9.85E-11 | 60.5 | 5.82 (0.5) |
| S1-49 | Non-RBD | 1.94E+06 | 3.63E-03 | 1.87E-09 | – | 49, 74‡ | 356 (32.8) | ||
| S1-50 | Non-RBD | 3.33E+053.34E-03 | 1.39E-023.94E-04 | 4.40E-09† | No interaction detected | 66 | 13(11) | ||
| S1-51 | RBD | 9.28E+04 | 4.22E-04 | 4.54E-09 | 3.77E+06 | 2.01E-03 | 5.33E-10 | 56 | 555.8 (52.5) |
| S1-52 | RBD | 4.22E+05 | 3.13E-04 | 7.74E-09 | 4.53E+04 | 1.94E-04 | 4.36E-09 | 57.5 | 3343 (291) |
| S1-53 | RBD | 1.40E+062.36E+04 | 8.46E-032.19E-04 | 6.05E-099.27E-09 | – | 51.5 | 2466 (939) | ||
| S1-54 | RBD | 1.13E+06 | 6.58E-05 | 5.84E-11 | 2.55E+04 | 2.88E-04 | 1.13E-08 | 69 | 1699 (1554) |
| S1-55 | RBD | 3.98E+063.53E+04 | 5.41E-035.31E-06 | 1.36E-09*1.51E-10 | 5.03E+051.84E+04 | 1.11E-021.82E-04 | 2.21E-08*9.89E-09 | 54.5 | 5725 (3372) |
| S1-56 | RBD | 1.46E+044.45E-03 | 2.99E-037.90E-05 | 3.57E-09† | 2.21E+03 | 1.05E-04 | 4.73E-08 | 54 | NA |
| S1-58 | Non-RBD | 5.73E+05 | 1.66E-04 | 2.90E-10 | – | 53.5 | 940 (795) | ||
| S1-60 | Non-RBD | 3.30E+054.61E+04 | 5.24E-063.67E-03 | 1.59E-11*9.58E-08 | – | 62 | NA | ||
| S1-61 | RBD | 9.87E+058.23E+02 | 1.81E-021.10E-04 | 1.84E-08*1.34E-07 | 4.46E+04 | 1.88E-04 | 4.21E-09 | 60 | NA |
| S1-62 | RBD | 2.68E+06 | 9.51E-05 | 3.54E-11 | 3.30E+06 | 6.30E-05 | 2.08E-11 | 71.5 | 3.3 (0.8) |
| S1-63 | RBD | 1.09E+063.39E+04 | 1.12E-021.67E-04 | 1.02E-08*4.94E-09 | 5.10E+04 | 2.23E-04 | 4.37E-09 | 65 | NA |
| S1-64 | Non-RBD | 6.97E+05 | 1.58E-04 | 2.26E-10 | – | 66 | 16.4 (11.7) | ||
| S1-65 | Non-RBD | 1.06E+06 | 1.67E-04 | 1.57E-10 | – | 60 | 7.3 (6.0) | ||
| S1-66 | Non-RBD | 4.66E+05 | 2.74E-04 | 5.87E-10 | – | 59 | NA | ||
| S1-RBD-3 | RBD | 8.81E+057.36E+04 | 1.76E-021.13E-03 | 2.00E-08*1.53E-08 | – | 72 | 384 (18.7) | ||
| S1-RBD-4 | RBD | 2.02E+06 | 1.64E-04 | 8.09E-11 | 2.83E+06 | 8.16E-04 | 2.89E-10 | 64.5 | 17.5 (1.98) |
| S1-RBD-5 | RBD | 1.94E+06 | 1.63E-04 | 8.38E-11 | 7.21E+06 | 1.05E-03 | 1.45E-10 | 64 | 174 (3.3) |
| S1-RBD-6 | RBD | 1.55E+06 | 1.63E-04 | 1.05E-10 | 3.48E+06 | 1.13E-03 | 3.24E-10 | 66.5 | 77.2 (21.8) |
| S1-RBD-9 | RBD | – | 2.85E+05 | 1.23E-04 | 4.30E-10 | 69 | 235 (97.5) | ||
| S1-RBD-10 | RBD | – | – | – | 52.9 | ||||
| S1-RBD-11 | RBD | 2.22E+07 | 2.94E-04 | 1.32E-11 | 2.06E+07 | 4.06E-04 | 1.97E-11 | 65 | 13.5 (5.5) |
| S1-RBD-12 | RBD | – | 1.10E+04 | 3.39E-05 | 3.10E-09 | 67 | NA | ||
| S1-RBD-14 | RBD | – | 1.33E+04 | 3.34E-04 | 2.51E-08 | 65 | NA | ||
| S1-RBD-15 | RBD | 5.37E+06 | 1.50E-04 | 2.79E-11 | 7.52E+06 | 4.95E-04 | 6.58E-11 | 59.5, 80‡ | 4.6 (1.2) |
| S1-RBD-16 | RBD | – | 1.68E+04 | 6.25E-05 | 3.73E-09 | 61 | 79.2 (4.2) | ||
| S1-RBD-18 | RBD | 2.28E+06 | 6.25E-04 | 2.74E-10 | 4.43E+06 | 1.27E-03 | 2.87E-10 | 69.5 | 67.2 (1.9) |
| S1-RBD-19 | RBD | – | – | 60 | 2124 (1451) | ||||
| S1-RBD-20 | RBD | 2.37E+06 | 2.23E-04 | 9.43E-11 | 3.05E+06 | 7.91E-04 | 2.59E-10 | 49, 70‡ | 12.4 (1.1) |
| S1-RBD-21 | RBD | 3.50E+06 | 1.31E-03 | 3.73E-10 | 3.15E+06 | 1.71E-03 | 5.45E-10 | 48.5, 70.5‡ | 17.3 (3.1) |
| S1-RBD-22 | RBD | 9.34E+05 | 2.28E-04 | 2.44E-10 | 9.24E+05 | 4.42E-04 | 4.78E-10 | 57.5 | 100 (0.1) |
| S1-RBD-23 | RBD | – | 2.89E+06 | 4.61E-05 | 1.59E-11 | 61 | 7.31 (0.4) | ||
| S1-RBD-24 | RBD | 1.61E+06 | 1.40E-03 | 8.65E-10 | 2.12E+06 | 1.22E-03 | 5.75E-10 | 46, 67‡ | 221 (4) |
| S1-RBD-25 | RBD | – | 8.41E+04 | 1.16E-02 | 1.38E-07 | – | NA | ||
| S1-RBD-26 | RBD | 1.06E+05 | 4.58E-06 | 4.32E-11 | 2.15E+05 | 1.33E-05 | 6.19E-11 | 66 | 241 (81.4) |
| S1-RBD-27 | RBD | – | 6.19E+06 | 1.24E-02 | 2.00E-09 | 71 | 163 (71.4) | ||
| S1-RBD-28 | RBD | 1.80E+06 | 4.27E-04 | 2.38E-10 | 1.80E+06 | 4.27E-04 | 2.38E-10 | 64.5 | 32.7 (3.1) |
| S1-RBD-29 | RBD | – | 5.36E+05 | 1.35E-03 | 2.51E-09 | 74 | 9.53 (1.0) | ||
| S1-RBD-30 | RBD | 2.15E+06 | 6.66E-05 | 3.10E-11 | 3.77E+06 | 4.82E-04 | 1.28E-10 | 65 | 25.0 (3.6) |
| S1-RBD-32 | RBD | – | 1.05E+05 | 7.90E-03 | 7.52E-08 | 65 | NA | ||
| S1-RBD-34 | RBD | – | 5.71E+04 | 4.88E-03 | 8.54E-08 | 64 | NA | ||
| S1-RBD-35 | RBD | 8.01E+05 | 1.68E-04 | 2.10E-10 | 1.33E+06 | 2.50E-04 | 1.88E-10 | 57, 68‡ | 12.3 (2.4) |
| S1-RBD-36 | RBD | – | – | 71 | NA | ||||
| S1-RBD-37 | RBD | – | 3.60E+05 | 8.88E-04 | 2.47E-09 | 71 | 523 (93.4) | ||
| S1-RBD-38 | RBD | – | 1.12E+06 | 9.84E-04 | 8.79E-10 | 68.5 | 84.6 (22.7) | ||
| S1-RBD-39 | RBD | – | 4.92E+05 | 7.77E-05 | 1.58E-10 | 67.5 | 90.4 (8.9) | ||
| S1-RBD-40 | RBD | – | 7.47E+05 | 2.77E-05 | 3.71E-11 | 70 | 25.6 (5.9) | ||
| S1-RBD-41 | RBD | – | 4.37E+05 | 1.39E-04 | 3.17E-10 | – | 17.0 | ||
| S1-RBD-43 | RBD | – | 6.21E+05 | 1.82E-04 | 2.92E-10 | 68 | 33.6 (1.3) | ||
| S1-RBD-44 | RBD | – | 1.91E+05 | 6.97E-05 | 3.65E-10 | 57.5 | 93.4 | ||
| S1-RBD-45 | RBD | – | 4.43E+05 | 4.14E-05 | 9.30E-11 | 53 | 22.6 | ||
| S1-RBD-46 | RBD | – | 4.69E+05 | 5.79E-04 | 1.23E-09 | 75.5 | 48.0 | ||
| S1-RBD-47 | RBD | – | 2.11E+05 | 6.45E-04 | 3.06E-09 | 53.5 | 127 (11.6) | ||
| S1-RBD-48 | RBD | – | 1.05E+05 | 2.42E-04 | 2.30E-09 | 58, 63‡ | 68.7 (14.2) | ||
| S1-RBD-49 | RBD | 3.24E+05 | 3.24E-04 | 1.00E-09 | 3.15E+05 | 5.34E-04 | 1.69E-09 | 66.5 | 37.6 (5.6) |
| S1-RBD-51 | RBD | – | 3.77E+06 | 2.01E-03 | 5.33E-10 | 52, 61‡ | 70.9 (29.3) | ||
-
*Curves were fit to a heterogeneous ligand model. Respective Kon, Koff, and KD values are shown for each component.
-
†Curves were fit to a two-state reaction model. Respective Kon, Koff, and KD values are shown for each binding state.
-
‡Two peaks were observed in the melting curve. Tms for both are reported.
-
–, not determined; NA, no activity.
S2 nanobody characterization; related to Figures 2 and 4.
Binding kinetics of S2 nanobodies were determined by surface plasmon resonance (SPR) using recombinant S2 protein, with on rates, off rates, and KDs determined by Langmuir fits to binding sensorgrams unless otherwise noted. Nanobody melting temperatures (Tm) were determined by differential scanning fluorimetry (DSF). Nanobodies were assayed for neutralization activity against a SARS-CoV-2 or SARS-CoV-1 spike pseudotyped HIV-1 virus (PSV), with IC50s calculated from neutralization curves with standard error of the mean (s.e.m.).
| ID | Kon(M–1 s–1) | Koff(s–1) | KD (M) | Tm (°C) | SARS-CoV-2 PSV IC50 (s.e.m.) (nM) |
|---|---|---|---|---|---|
| S2-1 | 6.32E+04 | 3.79E-04 | 6.00E-09 | 65.5 | NA |
| S2-2 | 1.26E+06 | 9.35E-05 | 7.41E-11 | 64.5 | 4460 (901) |
| S2-3 | 2.62E+05 | 7.21E-05 | 2.76E-10 | 65 | 2234 (751) |
| S2-4 | 2.44E+06 | 2.62E-04 | 1.08E-10 | 56 | NA |
| S2-5 | 9.35E+05 | 2.74E-04 | 2.93E-10 | 66 | NA |
| S2-6 | – | – | – | 61.5 | NA |
| S2-7 | 1.66E+06 | 9.36E-05 | 5.62E-11 | 61 | NA |
| S2-9 | 9.29E+05 | 2.32E-04 | 2.50E-10 | 64.5 | NA |
| S2-10 | 9.31E+04 | 3.13E-04 | 3.37E-09 | 59, 64.5* | 5269 (1418) |
| S2-11 | 7.94E+06 | 1.12E-03 | 1.41E-10 | 69.5 | NA |
| S2-13 | 7.02E+05 | 1.05E-04 | 1.49E-10 | 64.5 | NA |
| S2-14 | 3.16E+06 | 1.28E-03 | 4.07E-10 | 72.5 | NA |
| S2-15 | – | – | – | 70 | NA |
| S2-18 | 1.63E+06 | 4.87E-04 | 2.99E-10 | 47, 54.5* | NA |
| S2-22 | – | – | – | – | NA |
| S2-26 | 4.45E+05 | 8.15E-05 | 1.83E-10 | 76.5 | NA |
| S2-33 | 3.68E+05 | 5.58E-05 | 2.33E-10 | 70 | NA |
| S2-35 | 2.36E+05 | 4.72E-05 | 2.00E-10 | 77 | NA |
| S2-36 | 4.39E+06 | 3.69E-04 | 8.41E-11 | 74 | NA |
| S2-39 | – | – | – | 58 | NA |
| S2-40 | 5.08E+04 | 7.16E-05 | 1.41E-09 | 69.5 | 1712 (828) |
| S2-42 | 5.12E+05 | 3.77E-06 | 7.36E-12 | 69 | NA |
| S2-47 | 3.86E+05 | 1.14E-04 | 2.96E-10 | 40, 65* | NA |
| S2-57 | 2.33E+06 | 7.18E-04 | 3.08E-10 | 67 | NA |
| S2-59 | – | – | – | 37.5, 60* | NA |
| S2-62 | 1.65E+06 | 1.12E-04 | 6.81E-11 | 64, 77.5* | 7177 (5801) |
-
*Two peaks were observed in the melting curve. Tms for both are reported.
-
–, not determined; NA, no activity.
Nanobody binding activity against spike S1 variants; related to Figure 2.
Binding kinetics against wild-type spike S1 or two variants of concern were determined by surface plasmon resonance (SPR), with on rates, off rates, and KDs determined by Langmuir fits to binding sensorgrams.
| ID | Spike S1 variant | Kon(M–1 s–1) | Koff(s–1) | KD (M) |
|---|---|---|---|---|
| S1-1 | WT (Wuhan 2019) | 4.14E+05 | 2.98E-05 | 7.20E-11 |
| 20I/501Y.V1 | 2.71E+05 | 1.21E-05 | 4.44E-11 | |
| 20H/501Y.V2 | 2.78E+05 | 1.25E-05 | 4.51E-11 | |
| S1-6 | WT (Wuhan 2019) | 1.02E+06 | 5.75E-04 | 5.65E-10 |
| 20I/501Y.V1 | 3.55E+06 | 6.27E-04 | 1.77E-10 | |
| 20H/501Y.V2 | 1.03E+06 | 3.29E-04 | 3.20E-10 | |
| S1-23 | WT (Wuhan 2019) | 2.82E+06 | 4.91E-05 | 1.74E-11 |
| 20I/501Y.V1 | 5.96E+06 | 1.36E-03 | 2.29E-10 | |
| 20H/501Y.V2 | NA | NA | NA | |
| S1-RBD-9 | WT (Wuhan 2019) | 2.85E+05 | 1.23E-04 | 4.30E-10 |
| 20I/501Y.V1 | 4.84E+04 | 3.88E-05 | 8.01E-10 | |
| 20H/501Y.V2 | 1.34E+05 | 9.55E-05 | 7.13E-10 | |
| S1-RBD-11 | WT (Wuhan 2019) | 2.22E+07 | 2.94E-04 | 1.32E-11 |
| 20I/501Y.V1 | 3.84E+06 | 2.87E-04 | 7.46E-11 | |
| 20H/501Y.V2 | 6.85E+06 | 1.10E-03 | 1.61E-10 | |
| S1-RBD-15 | WT (Wuhan 2019) | 5.37E+06 | 1.50E-04 | 2.79E-11 |
| 20I/501Y.V1 | 2.99E+06 | 1.26E-04 | 4.22E-11 | |
| 20H/501Y.V2 | 4.02E+06 | 2.22E-04 | 5.53E-11 | |
| S1-RBD-35 | WT (Wuhan 2019) | 8.01E+05 | 1.68E-04 | 2.10E-10 |
| 20I/501Y.V1 | 1.33E+07 | 2.40E-03 | 1.80E-10 | |
| 20H/501Y.V2 | 5.94E+06 | 2.65E-03 | 4.46E-10 |
-
NA, no activity.
Characterization of oligomerized spike nanobodies; related to Figure 4.
Nanobody oligomers (1–4 nanobody repeats) were assayed for neutralization activity against a SARS-CoV-2 spike pseudotyped HIV-1 virus (PSV), with IC50s calculated from neutralization curves. Standard error of the mean (s.e.m.) is reported where replicates were available. Epitopes were determined by relative affinity for recombinant S1 or S1 RBD protein.
| ID | Epitope | SARS-CoV-2 PSV IC50 (s.e.m.)(nM) | SARS-CoV-1 PSV IC50 (s.e.m.) (nM) |
|---|---|---|---|
| S1-1 | RBD | 6.7 (1.0) | 8.6 (7.2) |
| S1-1dimer | RBD | 4.9 (0.1) | – |
| S1-1trimer | RBD | 5.7 (0.1) | – |
| S1-23 | RBD | 5.7 (2.2) | – |
| S1-23dimer | RBD | 0.22 (0.05) | NA |
| S1-23trimer | RBD | 0.089 (0.019) | NA |
| S1-RBD-35 | RBD | 12.3 (2.4) | NA |
| S1-RBD-35dimer | RBD | 0.15 (0.11) | – |
| S1-RBD-35trimer | RBD | 0.068 (0.043) | NA |
| S1-3 | S1 non-RBD | 1,030 (666) | 4161 |
| S1-3dimer | S1 non-RBD | 429 | 513 |
| S1-3trimer | S1 non-RBD | 411 | – |
| S1-30 | S1 non-RBD | 717 (388) | – |
| S1-30dimer | S1 non-RBD | 18.3 | 662 |
| S1-30trimer | S1 non-RBD | 77.5 (3.6) | – |
| S1-7 | S1 non-RBD | NA | – |
| S1-7dimer | S1 non-RBD | NA | – |
| S1-7trimer | S1 non-RBD | NA | – |
| S1-17 | S1 non-RBD | 1271 (888) | – |
| S1-17dimer | S1 non-RBD | 2144 | – |
| S1-49 | S1 non-RBD | 356 (32.8) | NA |
| S1-49dimer | S1 non-RBD | 9.1 (1.2) | NA |
| S1-49trimer | S1 non-RBD | 0.87 (0.08) | NA |
| S2-7 | S2 | NA | NA |
| S2-7dimer | S2 | 246 | 3516 |
| S2-7trimer | S2 | 112 | – |
| S2-7tetramer | S2 | 29.7 | – |
| S2-10 | S2 | 5269 (1418) | – |
| S2-10dimer | S2 | 48.0 (27.5) | – |
| S2-10trimer | S2 | 34.3 | – |
-
–, not determined; NA, no activity.
Nanobody neutralization activity against spike variants; related to Figure 4.
Nanobodies were assayed for neutralization activity against a pseudotyped HIV-1 virus (PSV) expressing SARS-CoV-1 or SARS-CoV-2 wild-type or variant spike, with IC50s calculated from neutralization curves. Standard error of the mean (s.e.m.) is reported where replicates were available.
| ID | Epitope | SARS-CoV-2 PSV IC50 (s.e.m.)(nM) | SARS-CoV-1 PSV IC50 (s.e.m.) (nM) | SARS-CoV-2 20H/501Y.V2 PSV IC50 (s.e.m.) (nM) | SARS-CoV-2 20I/501Y.V1 PSV IC50 (s.e.m.) (nM) | SARS-CoV-2 20J/501Y.V3 PSV IC50 (s.e.m.) (nM) |
|---|---|---|---|---|---|---|
| S1-1 | RBD | 6.7 (1.0) | 8.6 (6.4) | 7.2 (1.6) | 8.5 (4.2) | 2.9 (0.2) |
| S1-3 | Non-RBD | 1030 (666) | 3598 (563) | – | – | – |
| S1-4 | RBD | 41.5 (3.7) | 179 | – | – | – |
| S1-6 | RBD | 56.1 (20.7) | 227 (205) | – | – | – |
| S1-17 | Non-RBD | 1271 (888) | NA | – | – | – |
| S1-20 | RBD | 51.8 (3.7) | NA | NA | 13.5 | NA |
| S1-23 | RBD | 5.7 (2.2) | NA | NA | 78.3 | NA |
| S1-24 | Non-RBD | 868 | NA | – | – | – |
| S1-27 | RBD | 19.5 (4.9) | NA | – | – | – |
| S1-30 | Non-RBD | 717.8 (387) | NA | – | – | – |
| S1-31 | RBD | 78.7 (3.5) | NA | – | – | – |
| S1-35 | RBD | 12.5 (0.1) | 386.8 (350) | – | – | – |
| S1-36 | RBD | 48.5 (21.1) | NA | – | – | – |
| S1-37 | RBD | 6.8 (0.7) | NA | NA | 13.5 | NA |
| S1-39 | RBD | 111 (4.0) | 22.1 (18.5) | – | – | – |
| S1-41 | Non-RBD | 679 (53.4) | NA | – | – | – |
| S1-48 | RBD | 5.82 (0.5) | NA | 20.9 (1.4) | 7.3 (1.5) | 14.2 (3.5) |
| S1-49 | Non-RBD | 356 (32.8) | NA | – | – | – |
| S1-51 | RBD | 555.8 (52.5) | NA | – | – | – |
| S1-58 | Non-RBD | 940 (795) | NA | – | – | – |
| S1-62 | RBD | 3.3 (0.8) | – | NA | 6.4 (4.5) | NA |
| S1-RBD-6 | RBD | 77.2 (21.8) | 89.7 (4.2) | 75.8 | 78.3 | 43.5 |
| S1-RBD-9 | RBD | 235 (97.5) | 149.4 (54.1) | 115.7 (18.9) | 35.9 (5.1) | 24.5 (5.8) |
| S1-RBD-11 | RBD | 13.5 (5.50) | NA | 40.4 (2.5) | 100.6 | 138.6 |
| S1-RBD-15 | RBD | 4.6 (1.2) | NA | 3.4 (1.4) | 5.2 (1.0) | 1.3 (0.03) |
| S1-RBD-16 | RBD | 79.2 (4.2) | – | 1612 (1303) | 81.2 (10.9) | 35.7 (23.1) |
| S1-RBD-20 | RBD | 12.4 (1.1) | NA | NA | 49.6 | NA |
| S1-RBD-21 | RBD | 17.3 (3.1) | NA | 128.5 (16.8) | 24.3 (0.7) | 40.7 (7.0) |
| S1-RBD-23 | RBD | 7.3 (0.4) | NA | 16.3 (1.9) | 2.8 | 5.7 |
| S1-RBD-27 | RBD | 163 (71.4) | NA | NA | 99.8 | NA |
| S1-RBD-29 | RBD | 9.5 (1.0) | NA | – | – | – |
| S1-RBD-35 | RBD | 12.3 (2.4) | NA | 51.2 (4.2) | 11.5 (1.1) | 17.7 (4.0) |
| S1-RBD-37 | RBD | 523 (93.4) | NA | NA | – | – |
| S1-RBD-40 | RBD | 25.6 (3.4) | NA | 299.8 (200) | 10.5 (0.2) | 32.2 (7.1) |
| S1-RBD-47 | RBD | 127 (11.6) | NA | NA | 206 (123) | NA |
| S1-RBD-48 | RBD | 68.7 (14.2) | NA | NA | 106.6 (65.6) | NA |
| S2-2 | S2 | 4460 (901) | NA | – | – | – |
| S2-3 | S2 | 2234 (751) | 6277 | – | – | – |
| S2-40 | S2 | 1712 (828) | NA | – | – | – |
| S2-62 | S2 | 7177 (5801) | 1954 (364) | – | – | – |
-
–, not determined; NA, no activity.
Nanobody neutralization activity against SARS-CoV-2; related to Figure 5.
Nanobodies were assayed for neutralization activity against authentic SARS-CoV-2, with IC50s calculated from neutralization curves.
| ID | Epitope | SARS-CoV-2 IC50(nM) |
|---|---|---|
| S1-1 | RBD | 1.1 |
| S1-4 | RBD | 1310 |
| S1-23 | RBD | 0.7 |
| S1-RBD-4 | RBD | 5.4 |
| S1-RBD-11 | RBD | 3.0 |
| S1-RBD-23 | RBD | 6.1 |
| S2-10 | Non-RBD | 91.2 |
| LaM2 | Non-target ctrl | NA |
-
NA, no activity.
Nanobody neutralization of rVSV/SARS-CoV-2 and selected resistant mutants; related to Figure 6.
Neutralization assays were carried out using rVSV/SARS-CoV-2 and 293T/ACE2cl.22 target cells treated with the denoted nanobodies. Pseudovirus with either wild-type or variant spike (with escape mutants selected using the corresponding nanobody) was used. Escape mutants and IC50s are listed. Amino acid substitutions contributing to loss of neutralization activity are indicated in bold.
| Nanobody | Epitope | rVSV/SARS-CoV-2 variant | IC50 (nM)± s.e.m. |
|---|---|---|---|
| S1-1 | RBD | wt | 2.63 ± 0.23 |
| Y369N | 122 ± 3.0 | ||
| G404E | 40.8 ± 1.01 | ||
| S1-6 | RBD | wt | 13.0 ± 3.47 |
| D574N*, Q792H, Q992H | 587 ± 31.1 | ||
| S371P, H66R, N969T | 202 ± 29.9 | ||
| S1-23 | RBD | wt | 0.58 ± 0.02 |
| F490S, E484K, Q493K | > 1000 | ||
| Q493R, G252R | > 1000 | ||
| E484K† | > 1000 | ||
| S1-36 | RBD | wt | 3.69 ± 0.14 |
| W64R,L452F | 262 ± 10.1 | ||
| W64R,F490L,I931G | 870 ± 202 | ||
| W64R, F490S | >1000 | ||
| S1-37 | RBD | wt | 1.83 ± 0.59 |
| W64R, F490S | >1000 | ||
| S1-48 | RBD | wt | 1.75 ± 0.43 |
| Y449H, F490S, Q787R | >1000 | ||
| S494P | >1000 | ||
| S1-62 | RBD | wt | 0.65 ± 0.16 |
| E484K | >1000 | ||
| S1-3trimer | S1 non-RBD | wt | 60.0 |
| W64R, Y170H, V705M | >1000 | ||
| W64R, Y170H, Q787H | >1000 | ||
| S1-30trimer | S1 non-RBD | wt | 150 |
| T315I | 2400 | ||
| S1-49 | S1 non-RBD | wt | 146 ± 53.8 |
| S172G | >1000 | ||
| S1-49dimer | S1 non-RBD | wt | 3.38 ± 2.44 |
| S172G | >1000 | ||
| S1-49trimer | S1 non-RBD | wt | 0.47 ± 0.00 |
| S172G | >1000 | ||
| S2-10 | S2 | wt | 6649 ± 2,545 |
| W64R, S982R | >100,000 | ||
| S2-10dimer | S2 | wt | 1015 ± 236 |
| W64R, S982R | >40,000 | ||
| S1-RBD-9 | RBD | wt | 30.2 ± 7.43 |
| T259K, K378Q | >1000 | ||
| W64R, K378Q | >1000 | ||
| K378Q | >1000 | ||
| S1-RBD-11 | RBD | wt | 1.44 ± 0.53 |
| F486S | >1000 | ||
| T478R | >1000 | ||
| T478I | >1000 | ||
| S1-RBD-15 | RBD | wt | 1.21 ± 0.06 |
| Y508H | 549 ± 36.9 | ||
| S1-RBD-16 | RBD | wt | 268 ± 162 |
| N354S | >1000 | ||
| S1-RBD-21 | RBD | wt | 9.61 ± 1.90 |
| F486L | >1000 | ||
| Y489H | >1000 | ||
| S1-RBD-22 | RBD | wt | 31.5 ± 11.8 |
| K378Q | >1000 | ||
| S1-RBD-23 | RBD | wt | 14.8 ± 3.55 |
| L452R | >1000 | ||
| H245R, S349P, H1083Y | >1000 | ||
| S1-RBD-24 | RBD | wt | 58.0 ± 0.00 |
| P384Q | >1000 | ||
| K378Q† | >1000 | ||
| S1-RBD-29 | RBD | wt | 18.0 ± 1.80 |
| E484G | >1000 | ||
| E484K | >1000 | ||
| S1-RBD-35 | RBD | wt | 1.80 ± 0.15 |
| T478I | >1000 | ||
| F486L† | 306 ± 17.2 | ||
| Y489H† | 57.4 ± 4.5 | ||
| S1-RBD-40 | RBD | wt | 38.9 ± 11.7 |
| W64R, F490S | > 500 |
-
*
Residue 574 is outside the structurally covered region of the RBD (residues 333–526) and, therefore, was not used in the Integrative Modeling Platform modeling.
-
†
Variant was separately identified by selection against a different nanobody.
Cross-linked residues used in integrative modeling; related to Figure 6.
The indicated nanobodies were bound to RBD, NTD, or the spike ectodomain and cross-linked with disuccinimidyl suberate (DSS). Cross-linked complexes were excised from SDS-PAGE gels, reduced, alkylated, and digested with either trypsin or chymotrypsin. Peptides were extracted and analyzed by mass spectrometry. Cross-linked residues (listed) were identified using pLink, and spectra were manually validated to eliminate false positives.
| Nanobody | Nanobody residue # | Spike construct | 7KRQ residue number |
|---|---|---|---|
| S1-49 | 49 | NTD | 187 |
| S1-49 | 70 | NTD | 187 |
| S1-49 | 70 | NTD | 41 |
| S1-49 | 70 | NTD | 182 |
| S1-49 | 81 | NTD | 97 |
| S1-49 | 81 | NTD | 187 |
| S1-49 | 81 | NTD | 182 |
| S1-49 | 81 | NTD | 41 |
| S1-1 | 47 | RBD | 458 |
| S1-1 | 47 | RBD | 462 |
| S1-1 | 47 | RBD | 417 |
| S1-1 | 69 | RBD | 458 |
| S1-1 | 69 | RBD | 444 |
| S1-1 | 69 | RBD | 417 |
| S1-1 | 80 | RBD | 417 |
| S1-1 | 80 | RBD | 386 |
| S1-1 | 80 | RBD | 444 |
| S1-1 | 80 | RBD | 458 |
| S1-1 | 80 | RBD | 417 |
| S1-1 | 91 | RBD | 444 |
| S1-1 | 105 | RBD | 386 |
| S1-23 | 47 | RBD | 458 |
| S1-23 | 47 | RBD | 444 |
| S1-23 | 47 | RBD | 462 |
| S1-23 | 69 | RBD | 444 |
| S1-23 | 69 | RBD | 462 |
| S1-23 | 91 | RBD | 417 |
| S1-23 | 91 | RBD | 444 |
| S1-23 | 91 | RBD | 417 |
| S1-46 | 47 | RBD | 458 |
| S1-46 | 69 | RBD | 386 |
| S1-46 | 69 | RBD | 458 |
| S1-46 | 80 | RBD | 458 |
| RBD-9 | 47 | RBD | 444 |
| RBD-9 | 69 | RBD | 386 |
| RBD-9 | 69 | RBD | 444 |
| RBD-9 | 80 | RBD | 458 |
| RBD-9 | 114 | RBD | 444 |
| RBD-35 | 47 | RBD | 458 |
| RBD-35 | 47 | RBD | 462 |
| RBD-35 | 62 | RBD | 417 |
| RBD-35 | 62 | RBD | 458 |
| RBD-35 | 68 | RBD | 458 |
| RBD-35 | 68 | RBD | 444 |
| S2-10 | 69 | Spike ectodomain | 964 |
| S2-10 | 80 | Spike ectodomain | 835 |
| S2-10 | 115 | Spike ectodomain | 854 |
| S2-10 | 115 | Spike ectodomain | 964 |
| S2-40 | 69 | Spike ectodomain | 814 |
| S2-40 | 69 | Spike ectodomain | 786 |
| S2-40 | 69 | Spike ectodomain | 790 |
Nanobody synergy of neutralization activity; related to Figure 7.
Parameters from modeling the synergy observed for the indicated nanobody pairs. Multidimensional synergy of combinations (MuSyC), equivalent dose, and Bivariate Response to Additive Interacting Doses (BRAID) models were used to determine if statistically significant synergy was evident from the neutralization response in a 2D grid of nanobody concentrations.
| First nanobody | Second nanobody | (Nanobody #1) experimental range | (Nanobody #2) experimental range | h1 | h2 | C1 (nM) | C2 (nM) | alpha12 | alpha21 | a12 | a21 | Kappa |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| S1-23 | S1-27 | 4.1 pM to 717 nM, 0 µM | 3.4 pM to 600 nM, 0 µM | 1.80 | 1.46 | 25.0 | 20.0 | n.s. | n.s. | n.s. | n.s. | n.s |
| S1-1 | S1-23 | 2.2 pM to 387 nM, 0 µM | 4.1 pM to 717 nM, 0 µM | 0.93 | 1.51 | 4.2 | 18.6 | 21.4 | 31.8 | Synergy | Synergy | Synergy |
| S1-RBD-15 | S1-23 | 3 pM -to 527 nM, 0 µM | 4.1 pM to 717 nM, 0 µM | 1.32 | 1.42 | 3.7 | 10.3 | 300.0 | 10.2 | Synergy | Synergy | Synergy |
| S1-RBD-15 | S1-RBD-23 | 3 pM to 527 nM, 0 µM | 1.8 pM to 325 nM, 0 µM | 1.44 | 1.00 | 4.4 | 7.9 | 523 | 2.9 | Synergy | n.s. | Synergy |
| S1-23 | S1-46 | 4.1 pM to 717 nM, 0 µM | 7.55 pM to 1.34 µM, 0 µM | 1.76 | 0.75 | 3.3 | 273.0 | 50.1 | 1.0 | Synergy | Antagonism | Synergy |
| S1-RBD-15 | S1-46 | 39.7 pM to 7.04 µM, 0 µM | 7.55 pM to 1.34 µM, 0 µM | 1.10 | 0.75 | 4.9 | 164 | 10.7 | 0.9 | Synergy | Antagonism | Synergy |
| S1-23 | S2-10-dimer | 4.1 pM to 717 nM, 0 µM | 5.1 pM to 897 nM, 0 µM | 1.48 | 1.14 | 5.4 | 45.4 | 4232.0 | 2.8 | Synergy | n.s. | Synergy |
| S1-49 | S1-1 | 2.1 pM to 367 nM, 0 µM | 2.2 pM to 387 nM, 0 µM | 0.77 | 1.62 | 152.0 | 1.7 | 1,147 | 18.1 | Synergy | Synergy | Synergy |
| S1-49 | S1-RBD-15 | 2.1 pM to 367 nM, 0 µM | 39.7 pM to 7.04 µM, 0 µM | 0.85 | 1.20 | 629.0 | 2.2 | 243.5 | 20.3 | n.s. | Synergy | Synergy |
| S1-RBD-15 | S2-10-dimer | 3 pM to 527 nM, 0 µM | 5.1 pM to 897 nM, 0 µM | 1.57 | 1.00 | 1.9 | 56 | 4,110 | 2.2 | Synergy | Synergy | Synergy |
-
n.s., not significant; h1, h2, Hill slope; C1, C2, IC50 (nM);alpha12, alpha21, synergistic/antagonistic fold change of potency from MuSyC model; a12, a21, equivalent dose model synergy/antagonism; kappa, BRAID model synergy/antagonism.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | ArticExpress(DE3) | Agilent | Cat# 230192 | Competent cells, enabling efficient high-level expression of heterologous proteins. |
| Strain, strain background (vesicular stomatitis virus) | rVSV/SARS-CoV-2/GFP; WT2E1 | Schmidt et al., 2020 | Recombinant chimeric VSV/SARS-CoV-2 reporter virus.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-Y369N | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-G404E | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-D574N, E484K, Q493K | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-S371P, H66R, N969T | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-F490S, E484K, Q493K | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-Q493R, G252R | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, L452F | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-H245R, H1083Y | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, F490L, I931G | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, F490S | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-Y449H, F490S, Q787R | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-S494P | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-S172G | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-E484K | Schmidt et al., 2020 | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, S982R | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-T259K, K378Q | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, K378Q | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-F486S | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-T478R | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-T478I | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-Y508H | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-N354S | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-F486L | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-Y489H | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-K378Q | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-L452R | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-H245R, S349P, H1083Y | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-P384Q | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-E484G | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, Y170H, V705M | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-W64R, Y170H, Q787H | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (vesicular stomatitis virus) | 2E1-T315I | This study | Mutant rVSV/SARS-CoV-2/GFP derivative.Inquiries should be addressed to P. Bieniasz. | |
| Strain, strain background (betacoronavirus) | SARS-CoV-2, Isolate USA-WA1/2020 | BEI Resources | NR-52281 | Wild-type SARS-CoV-2. |
| Strain, strain background (betacoronavirus) | SARS-CoV-2, Isolate USA-WA (B.1.617.2) | R.Coler | Delta | SARS-CoV-2 delta variant of concern. |
| Biological sample (Lama glama) | Bone marrow aspirates | Capralogics | From two male llamas immunized with SARS-CoV-2 spike S1, RBD, and S2. | |
| Biological sample (L. glama) | Sera | Capralogics | From two male llamas immunized with SARS-CoV-2 spike S1, RBD, and S2. | |
| Cell line (Homo sapiens) | 293T/17 | ATCC | CRL-11268 | Human kidney epithelial cells. |
| Cell line (H. sapiens) | 293/ACE2cl.22 | Schmidt et al., 2020 | 293T cells expressing human ACE2 (single-cell clone). | |
| Cell line (H. sapiens) | 293T-ACE2 | Cawford et al., 2020 | BEI NR-52511 | 293T cells expressing human ACE2 (single-cell clone). |
| Cell line (H. sapiens) | Primary human airway epithelial cells | This study | Air-liquid interface culture system.Inquiries should be addressed to J. Debley. | |
| Cell line (Cercopithecus aethiops) | VERO C1008 [Vero 76, clone E6, Vero E6] | ATCC | CRL-1586 | Monkey kidney epithelial cells. |
| Cell line (C. aethiops) | TMPRSS2+ Vero E6 | R.Coler | Vero E6 cells expressing human TMPRSS2. | |
| Antibody | Anti-COVID-19 and SARS-CoV S glycoprotein [CR3022] (human monoclonal) | Absolute Antibody | Cat# Ab01680-10.0 | Flow cytometry (1:1000). |
| Antibody | Anti-human IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 (goat polyclonal) | Invitrogen | Cat# A-11013 | Flow cytometry (1:2000). |
| Recombinant DNA reagent | pET21-pelB | Fridy et al., 2014a | Expression plasmid for expressing nanobodies. | |
| Recombinant DNA reagent | pET21-pelB-SARS-CoV-2 Nanobody | This study; Fridy et al., 2014a | See Table 1 and Table 2 | SARS-CoV-2 nanobody expression plasmids.Inquiries should be addressed to M. Rout. |
| Recombinant DNA reagent | pcDNA3.1+ SARS-1-S-C9 | T.Gallagher | SARS-CoV-1 spike expression plasmid. | |
| Recombinant DNA reagent | pcDNA3.1+ SARS-2-S-C9 WUHAN-1 | T.Gallagher | SARS-CoV-2 spike expression plasmid. | |
| Recombinant DNA reagent | pcDNA3.1+ SARS-2-B.1.1.7 | NIAID | SARS-CoV-2 spike expression plasmid for alpha variant. | |
| Recombinant DNA reagent | pHDM-SARS-CoV-2-Spike-B.1.351 | L.Stamatatos | SARS-CoV-2 spike expression plasmid for beta variant. | |
| Recombinant DNA reagent | pcDNA3.1+ SARS-2 P.1 | NIAID | SARS-CoV-2 spike expression plasmid for gamma variant. | |
| Recombinant DNA reagent | psPAX2 | D.Trono/Addgene | Plasmid #12260 | Second-generation lentiviral packaging plasmid. |
| Recombinant DNA reagent | pHAGE-CMV-Luc2-IRES-ZsGreen-W | J.Bloom / Cawford et al., 2020 | NR-52516 | Lentiviral backbone plasmid that uses a CMV promoter to express luciferase followed by an IRES and ZsGreen. |
| Sequence-based reagent (primer) | 6N_CALL001 | This study | PCR and sequencing primer | NNNNNNGTCCTGGCTGCTCTTCTACAAGG |
| Sequence-based reagent (primer) | 6N_CALL001B | This study | PCR and sequencing primer | NNNNNNGTCCTGGCTGCTCTTTTACAAGG |
| Sequence-based reagent (primer) | 6N_VHH_SH_rev | This study | PCR and sequencing primer | NNNNNNCTGGGGTCTTCGCTGTGGTGC |
| Sequence-based reagent (primer) | 6N_VHH_LH_rev | This study | PCR and sequencing primer | NNNNNNGTGGTTGTGGTTTTGGTGTCTTGGG |
| Peptide, recombinant protein | Spike S1 (Wuhan Str.) | Sino Biological | Cat# 40591-V08H | For determining KDs. |
| Peptide, recombinant protein | Spike RBD | Sino Biological | Cat# 40592-VNAH | For determining KDs. |
| Peptide, recombinant protein | SARS-CoV-2 (2019-nCoV) Spike RBD-mFc Recombinant Protein | Sino Biological | Cat# 40592-V05H | For epitope mapping. |
| Peptide, recombinant protein | Spike S2 | Sino Biological | Cat# 40590-V08B | For immunization and for determining KDs. |
| Peptide, recombinant protein | Spike S1 (501Y.V1) | Sino Biological | Cat# 40591-V08H12 | For determining KDs. |
| Peptide, recombinant protein | Spike S1 (501Y.V2) | Sino Biological | Cat# 40591-V08H10 | For determining KDs. |
| Peptide, recombinant protein | SARS-CoV-2 Spike S1, Sheep Fc-Tag | The Native Antigen Co. | Cat# REC31806 | For immunization. |
| Peptide, recombinant protein | SARS-CoV-2 Spike S2, Sheep Fc-Tag | The Native Antigen Co. | Cat# REC31807 | For immunization. |
| Peptide, recombinant protein | Protein M | Grover et al., 2014 | Used to deplete light-chain containing IgGs. | |
| Peptide, recombinant protein | Thyroglobulin | Sigma-Aldrich | Cat# A8531-1V | Used to calibrate mass photometer. |
| Peptide, recombinant protein | Bovine serum albumin | Sigma-Aldrich | Cat# T9145-1VL | Used to calibrate mass photometer. |
| Peptide, recombinant protein | Beta-amylase | Sigma-Aldrich | Cat# A8781-1VL | Used to calibrate mass photometer. |
| Peptide, recombinant protein | FabRICATOR (IdeS) | Genovis | Cat# A0-FR1-050 | Protease to cleave VHH domain from the HCAb. |
| Commercial assay, kit | ProteOn Amine Coupling Kit | Bio-Rad | Cat# 1762410 | Used to couple VHH domain to beads. |
| Commercial assay, kit | TruSeq Nano DNA Low Throughput Library Prep Kit | Illumina | Cat# 20015964 | Used to sequence VHH. |
| Commercial assay, kit | Steady-GLO | Promega | Cat# E2520 | Used in pseudovirus assay. |
| Software, algorithm | Llama-Magic | Fridy et al., 2014a, | https://github.com/FenyoLab/llama-magic | For identifying nanobody sequences. |
| Software, algorithm | IMP, the Integrative Modeling Platform | Russel et al., 2012 | https://integrativemodeling.org | For integrative structural modeling. |
| Software, algorithm | UCSF ChimeraX | Pettersen et al., 2021 | https://www.rbvi.ucsf.edu/chimerax/download.html | For visualizing structural models. |
| Software, algorithm | synergy v0.4 | Wooten et al., 2021 | https://pypi.org/project/synergy/ | For observing synergy. |
| Software, algorithm | matplotlib v3.4.1 | Hunter, 2007 | https://pypi.org/project/matplotlib/ | For preparing figures. |
| Software, algorithm | seaborn v0.11.0 | Waskom, 2021 | https://seaborn.pydata.org/index.html | For preparing figures. |
| Software, algorithm | plotly v4.12.0 | 2019 Plotly, Inc | https://plotly.com/python/ | For preparing figures. |
| Software, algorithm | numpy v1.19.2 | Huo et al., 2020a | https://numpy.org | For data analysis. |
| Software, algorithm | pandas v1.1.2 | The pandas development team, 2020 | https://pandas.pydata.org | For data analysis. |
| Software, algorithm | scipy v1.5.0 | Virtanen et al., 2020 | https://www.scipy.org | For data analysis. |
| Software, algorithm | scikit-learn v0.23.2 | Pedregosa, 2011 | https://scikit-learn.org/stable/ | For data analysis. |
| Software, algorithm | python v3.8 | Van Rossum and Drake, 2009 | https://www.python.org | For data analysis and preparing figures. |
| Software, algorithm | Prism 9 | GraphPad | https://www.graphpad.com | For data analysis and preparing figures. |
| Software, algorithm | OCTET | Sartorius | For data analysis. | |
| Other | ProteOn GLC Sensor Chip | Bio-Rad | Cat# 176-5011 | For protein interaction analysis. |
| Other | Series S Sensor Chip CM5 | Cytiva | Cat# BR100530 | For protein interaction analysis. |
| Other | Hard-shell PCR plates, 96-well, thin-wall | Bio-Rad | Cat# HSP9661 | For differential scanning fluorimetry. |
| Other | Microseal ‘B’ seal | Bio-Rad | Cat# MSB1001 | For differential scanning fluorimetry. |
| Other | Precision Coverslips | Thorlabs | Cat# CG15KH1 | For mass photometry. |
| Other | CultureWell Reusable Gasket | Grace Bio-Labs | Cat# 103250 | For mass photometry. |
| Other | Protein A Sepharose 4B | Thermo Fisher | Cat# 101042 | For protein purification. |
| Other | Recombinant Protein G Sepharose 4B | Thermo Fisher | Cat# 101243 | For protein purification. |
| Other | CNBr-activated Sepharose 4 Fast Flow | Cytiva | Cat# 17098101 | For nanobody screening. |
| Other | SuperScript VILO Master Mix | Thermo Fisher | Cat# 11755250 | For viral escape analysis. |
| Other | Anti-mouse IgG Fc Capture Biosensors | Sartorius | Cat# 18-5088 | For epitope mapping. |
| Other | KOD Xtreme Hot Start DNA Polymerase | Sigma-Aldrich | Cat# 71975 | For viral escape analysis. |






