Reserpine maintains photoreceptor survival in retinal ciliopathy by resolving proteostasis imbalance and ciliogenesis defects
Figures
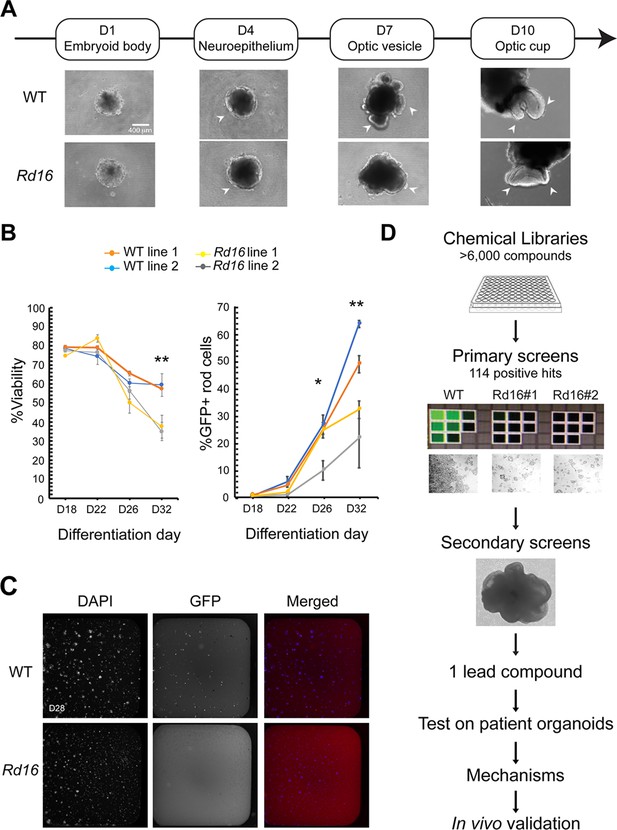
Drug discovery pipelines to identify drug candidates.
(A) Morphology of Nrl-GFP wild-type (WT) and rd16 retinal organoids differentiated from mouse-induced pluripotent stem cells (iPSC) at various differentiation time points. (B) Flow cytometry analysis of GFP+ rod photoreceptors and viability at different developmental stages. Each data point summarized at least three batches of independent experiments, each of which included at least 10 organoids. One-way ANOVA was performed to compare the %GFP+ cells and viability of organoids from four different cell lines. *p<0.05; **p<0.01. (C) Fluorescent images of dissociated day (D) 28 WT and rd16 cells stained by 4',6-diamidino-2-phenylindole (DAPI) and anti-GFP antibody. (D) Schematic outline of the drug discovery strategy.
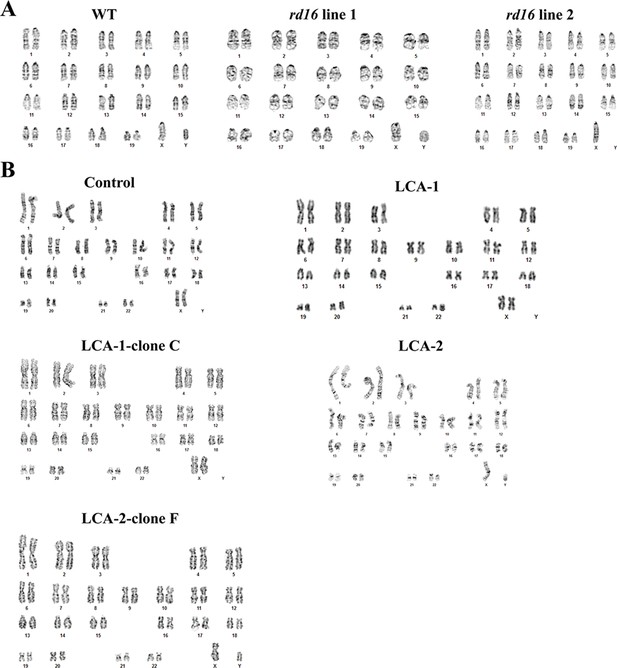
Karyotypes of induced pluripotent stem cell lines.
(A) Mouse. (B) Human. Cells were prepared for G-band stain. Fifteen unique metaphases were analyzed.
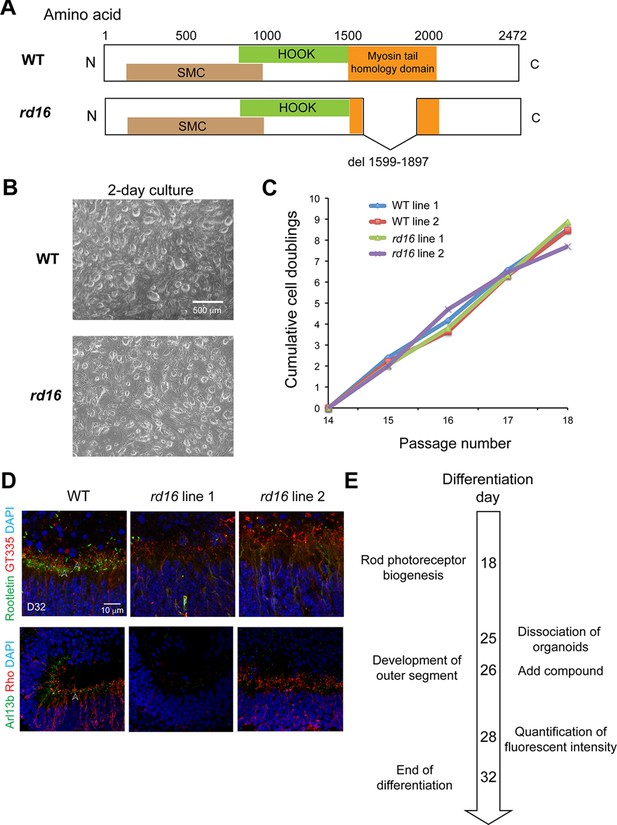
Phenotypes of retinal organoids differentiated from induced pluripotent stem cells of Nrl-GFP rd16 mice.
(A) Deletion of myosin tail homology domain of the CEP290 protein in rd16 mice compared to the wild-type (WT). (B) Morphology of induced pluripotent stem cells (iPSC) colonies, (C) proliferation rate of iPSCs, and (D) photoreceptor primary cilium of organoids were compared between WT and rd16. (E) Timeline of the key differentiation events in mouse organoids and primary screens. Images were representative of at least three independent experiments, each of which had at least three organoids.
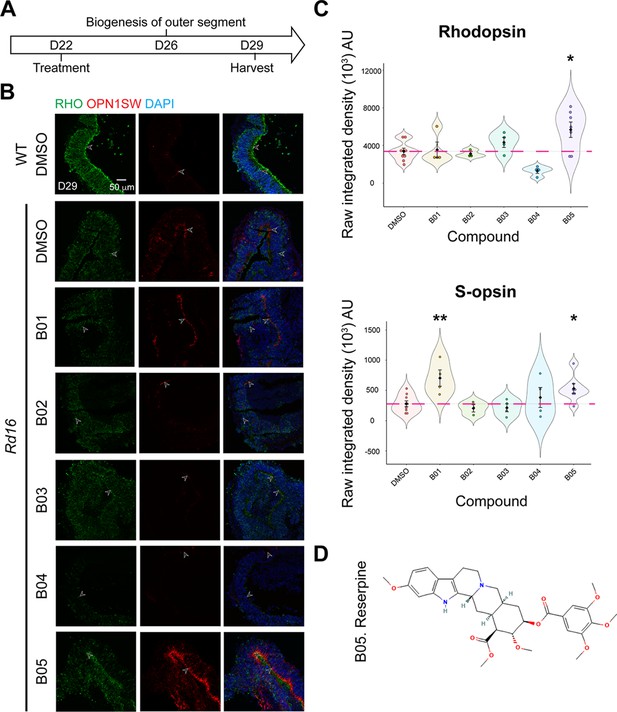
Identification of reserpine as the lead compound.
(A) Timeline for hit validation in rd16 retinal organoids. (B) Immunostaining of rod cell marker rhodopsin (RHO, green) and cone cell marker S-opsin (OPN1SW, red) in wild-type (WT) and rd16 organoids treated with non-toxic positive hits (B01–B05). Drug vehicle dimethylsulfoxide (DMSO) was used as a control. Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three independent experiments, each of which had at least three organoids. (C) Bee swarm plots show the quantification of fluorescence intensity of rhodopsin (upper) and S-opsin (lower) staining in the validation. The shape of the plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. The pink dash line shows the mean fluorescence intensity of DMSO-treated organoids. The plot summarizes at least three independent experiments with at least three organoids in each batch. One-way ANOVA followed by the Bonferroni test was performed. *p<0.05; **p<0.01. (D) Chemical structure depiction of the selected lead compound reserpine.
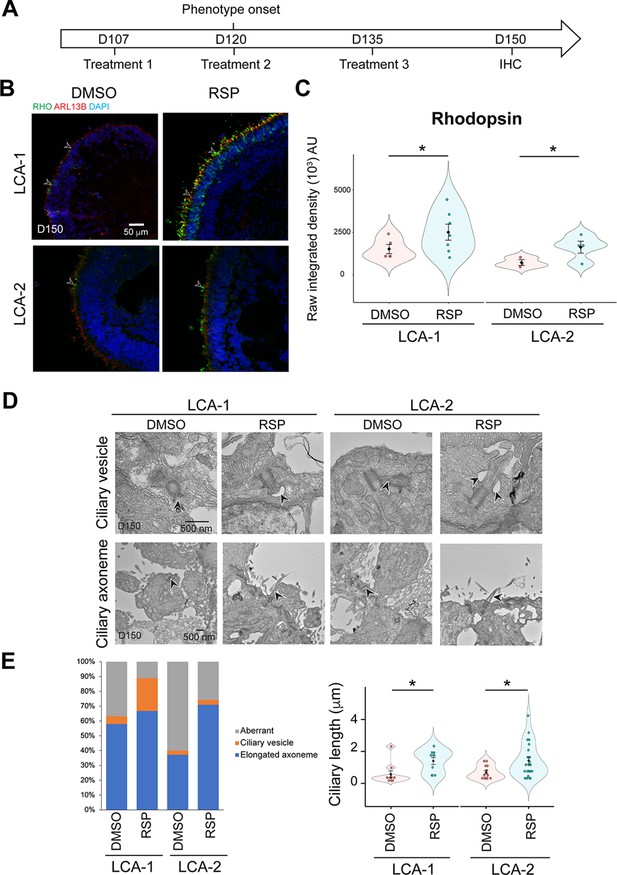
Effect of reserpine (RSP) on LCA10 patient retinal organoids.
(A) Timeline for RSP treatments and harvest of patient organoids. (B) Immunostaining of rhodopsin (RHO, green) and ARL13B (red). Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three independent experiments, each of which had at least three organoids. (C) Bee swarm plots show the quantification of fluorescence intensity of rhodopsin staining. The shape of the plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. The plot summarizes at least three independent experiments with at least one image quantified in each batch. Unpaired t-test was performed to compare untreated and treated groups. *p<0.05. (D) Transmission electron microscopy analysis of control, untreated, and treated patient organoids. Arrowheads indicate relevant staining. (E) Quantification of the number of the ciliary vesicles, elongated ciliary axoneme, and aberrant cilia (left) as well as the length of the primary cilia (right) in untreated and RSP-treated patient organoids. Aberrant cilia were defined as docked mother centrioles without ciliary vesicles or elongated ciliary axoneme. The data summarized at least four batches of independent experiments, each of which has at least two organoids and seven docked mother centrioles. The shape of the bee swamp plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. Unpaired t-test was performed to compare untreated and treated groups. *p<0.05.
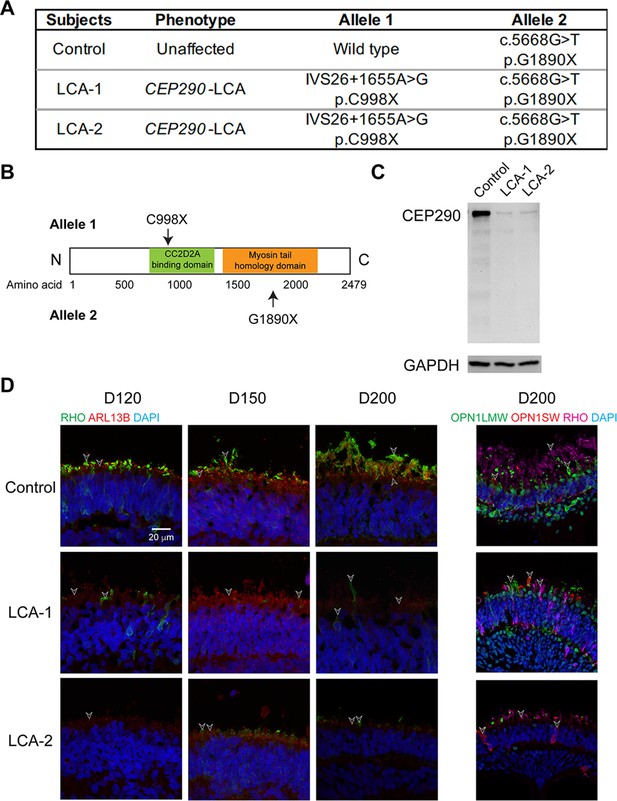
Genotype and phenotype of the CEP290-LCA family in this study.
(A) Genotypes of the subjects including a heterozygous but phenotypically normal control and two CEP290-LCA patients. (B) Location of the stop codons caused by point mutations in CEP290-LCA patients. (C) Western blot analysis of CEP290 in D200 control and patient retinal organoids. Images were representative of at least three independent experiments, each of which had at least three organoids. GAPDH was used as a loading control. (D) Immunostaining of. rhodopsin (green) and ARL13B (red) in the left panel as well as rhodopsin (magenta), S-opsin (red) and L/M-opsin (green) in the right panel. Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three independent experiments, each of which had at least three organoids.
-
Figure 3—figure supplement 1—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of CEP290.
The size of the protein ladders, CEP290, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig3-figsupp1-data1-v2.zip
-
Figure 3—figure supplement 1—source data 2
Overlay of the bright field and chemiluminescence images indicating the signal of GAPDH.
The size of the protein ladders, GAPGH, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig3-figsupp1-data2-v2.zip
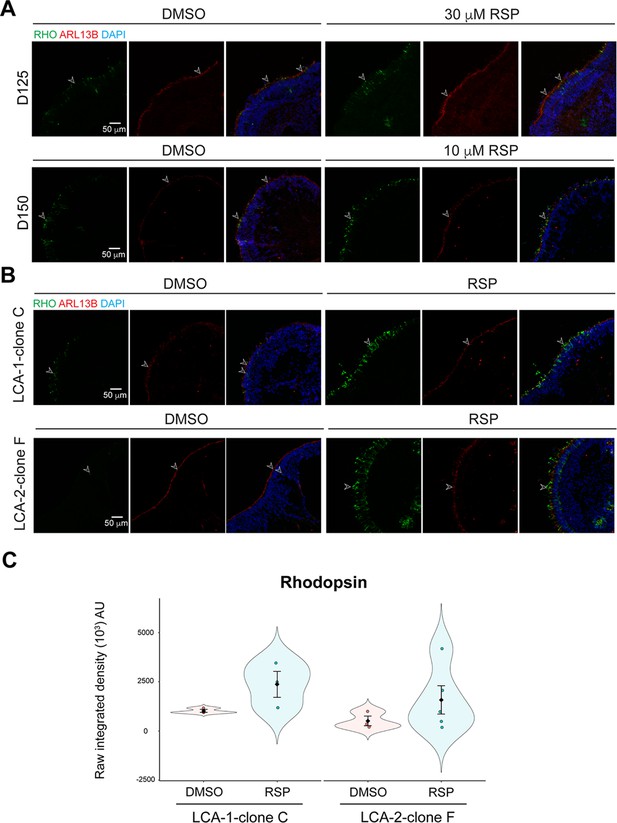
Effect of reserpine at various conditions in CEP290-LCA retinal organoids.
Rod cell marker rhodopsin (RHO, green) and ciliary marker ARL13B (red) were shown in (A) LCA1 organoids treated with 30 μM reserpine (RSP) at differentiation day (D) 125 (upper) and with 10 μM RSP at D150 (lower) and (B) organoids differentiated from clone C of LCA1 (upper) and clone F of LCA2 (lower) at D150. (C) Quantification of fluorescence intensity of rhodopsin staining. The shape of the Bee Swamp plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. The plot summarizes at least three independent experiments with at least one image quantified in each batch. Unpaired t-test was performed to compare untreated and treated groups. *p<0.05.
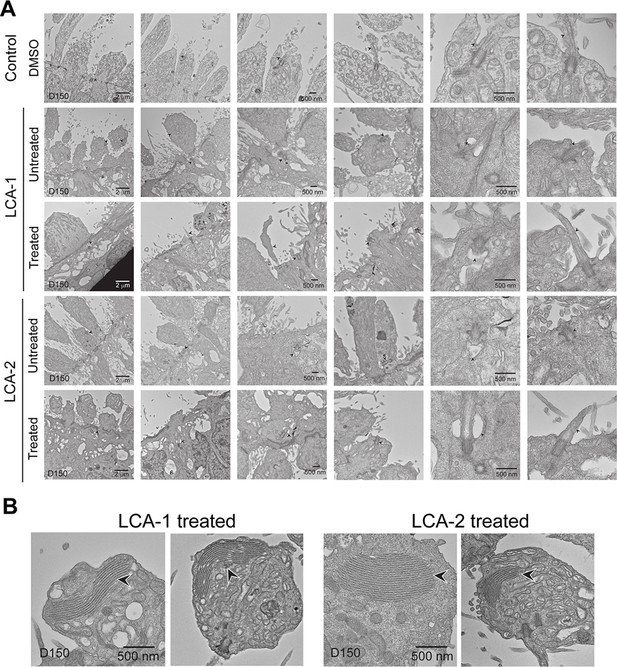
Transmission electron microscopy analyses of control, and untreated and treated CEP290-LCA retinal organoids.
(A) Morphology of photoreceptor primary cilium. (B) Stacks of disc-like structures observed in treated patient organoids. Arrowheads indicate relevant structures.
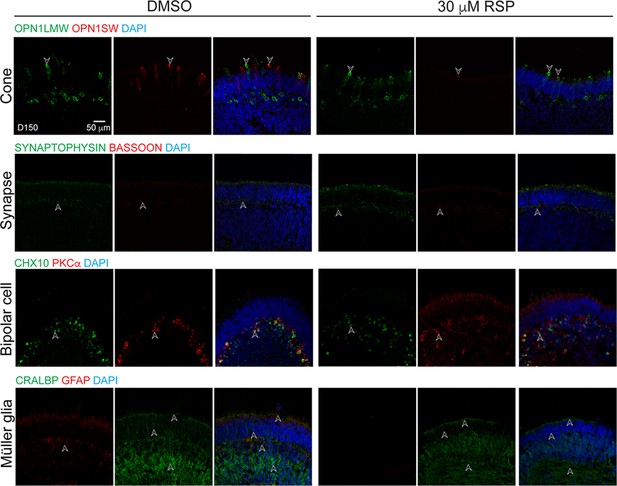
Various cell types in reserpine (RSP)-treated CEP290-LCA retinal organoids.
Immunostaining of structural or cell type-specific markers including those for L/M cones (OPN1LMW, green), S-cones (OPN1SW, red), ribbon synapses (BASSOON, red), presynaptic vesicles (SYNAPTOPHYSIN, green), bipolar cells (CHX10, green; PKCα, red), Müller glia (CRALBP, green; GFAP, red). Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three independent experiments, each of which had at least three organoids.
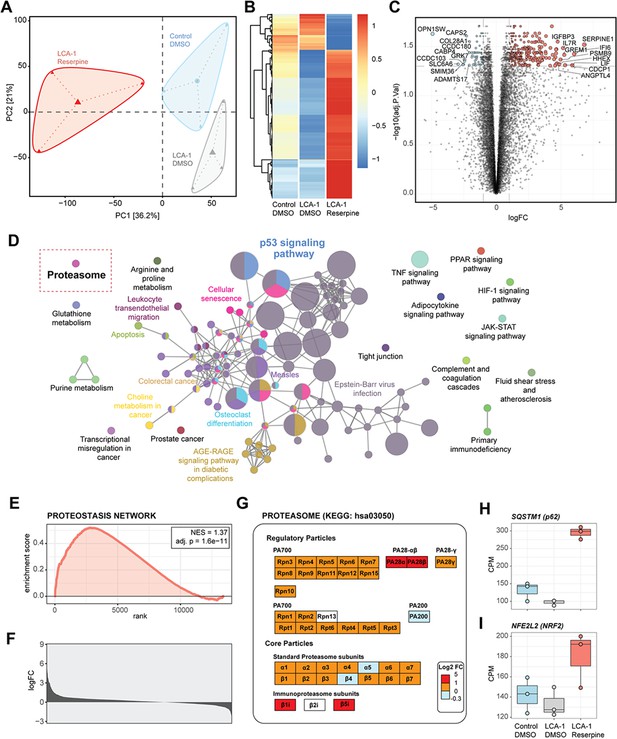
Transcriptomic upregulation of proteasomal components induced by reserpine in patient retinal organoids.
(A) Principal component analysis (PCA) diagram of patient and control organoids shows altered retinal transcriptomes after reserpine treatment. (B) Drug-induced genes in patient organoids displayed specific trends compared to control organoids. (C) Volcano plot summarizes reserpine-induced differential gene expression changes in LCA-1 organoids. (D) ClueGO analysis of KEGG pathway enrichment showed overexpressed genes mapping to protein homeostasis, metabolism, and cellular signaling processes. The red rectangle highlights the ‘Proteasome’ pathway. (E) GSEA plot showing enrichment and significance of Proteostasis Network. (F) Histogram of the log fold change of Proteostasis Network genes upon reserpine treatment. (G) Proteasomal subunits responded strongly to reserpine treatment. (H) SQSTM1 (p62) and (I) NFE2L2 (NRF2), two key regulators of protein homeostasis, showed increased expression after reserpine treatment.
-
Figure 4—source data 1
Differentially expressed genes in reserpine-treated retinal organoids.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig4-data1-v2.xlsx
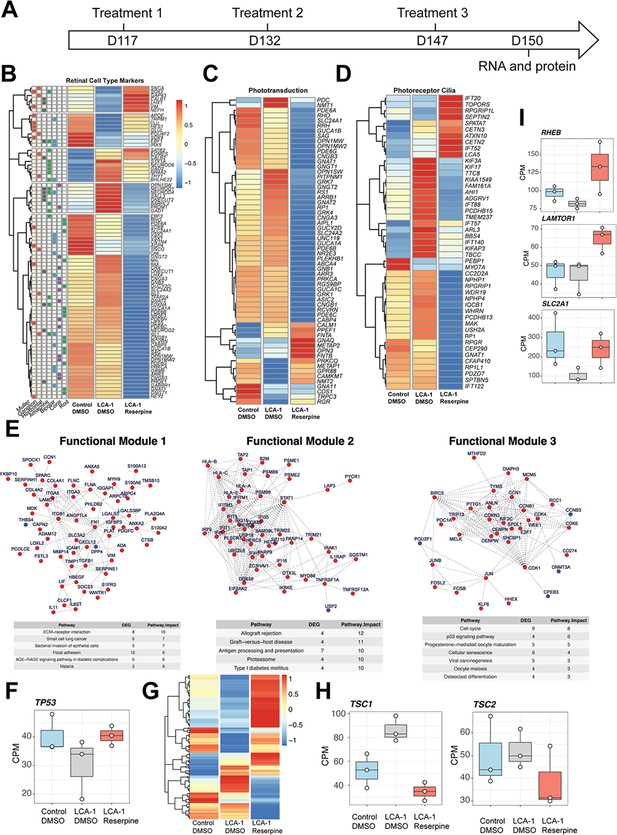
Reserpine treatment alters expression of retinal pathway genes.
(A) Timeline of treatment and sample harvest. (B) Different retinal cell types appeared to respond uniquely to reserpine. (C) Phototransduction genes were globally downregulated except for specific genes like OPN3. (D) Genes associated with photoreceptor cilia showed mixed trends. (E) Functional clustering of differentially expressed genes shows activity in cell survival pathways. (F) TP53 gene expression change with reserpine treatment. (G) Heatmap of p53 signaling pathway (combination of KEGG, Reactome, and Hallmark genes). (H) Transcriptional response of TSC1 and TSC2 genes show reduced expression after reserpine treatment. (I) Expression of mTOR pathway genes upon reserpine treatment.
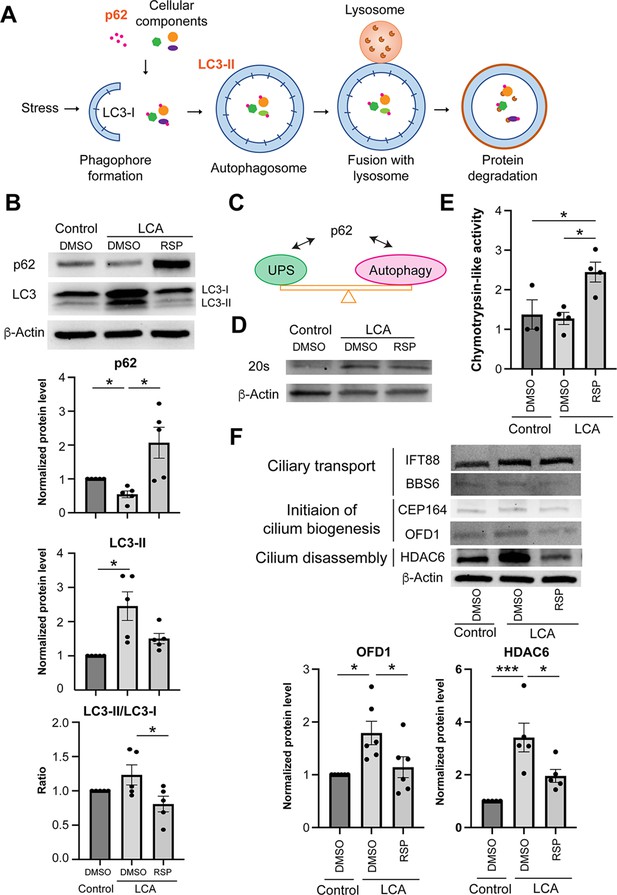
Proteostasis Network in patient organoid cultures in response to reserpine treatment.
(A) Schematic diagram of autophagy.(B) Immunoblot analyses and quantification of autophagy cargo adaptor p62 and autophagosome marker LC-II in control and patient organoids treated with reserpine (RSP). (C) Schematic diagram showing the proteome balance between ubiquitin-proteasome system (UPS) and autophagy is mediated through p62 as documented in the literature. (D) Immunoblot analysis of the 20S proteasome in control, DMSO-, and RSP-treated cultures. β-Actin was used as the loading control. (E) Proteasomal chymotrypsin-like activity in organoids. (F) Immunoblot analyses and quantification of key regulators of cilium assembly/disassembly in control, untreated, and RSP-treated patient organoids. The drug vehicle dimethylsulfoxide (DMSO) was added to the cultures in the untreated group at the same volume as the drugs. β-Actin was used as the loading control. The histograms summarize data in at least three batches of experiments, each of which had at least three retinal organoids per group. Each dot in the histogram shows data in one batch of experiment and are presented as mean ± standard deviation. One-way ANOVA followed by Tukey’s test. *p<0.05; ***p<0.005.
-
Figure 5—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of p62.
RSP stands for reserpine. The size of the protein ladders, p62, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data1-v2.zip
-
Figure 5—source data 2
Overlay of the bright field and chemiluminescence images indicating the signal of LC3.
RSP stands for reserpine. The size of the protein ladders, LC3, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data2-v2.zip
-
Figure 5—source data 3
Overlay of the bright field and chemiluminescence images indicating the signal of β-Actin.
RSP stands for reserpine. The size of the protein ladders, β-Actin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data3-v2.zip
-
Figure 5—source data 4
Overlay of the bright field and chemiluminescence images indicating the signal of 20S proteosome and β-Actin.
RSP stands for reserpine. The size of the protein ladders, β-Actin, 20S proteosome, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data4-v2.zip
-
Figure 5—source data 5
Overlay of the bright field and chemiluminescence images indicating the signal of IFT88.
RSP stands for reserpine. The size of the protein ladders, IFT88, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data5-v2.zip
-
Figure 5—source data 6
Overlay of the bright field and chemiluminescence images indicating the signal of BBS6.
RSP stands for reserpine. The size of the protein ladders, BBS6, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data6-v2.zip
-
Figure 5—source data 7
Overlay of the bright field and chemiluminescence images indicating the signal of CEP164.
RSP stands for reserpine. The size of the protein ladders, CEP164, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data7-v2.zip
-
Figure 5—source data 8
Overlay of the bright field and chemiluminescence images indicating the signal of OFD1.
RSP stands for reserpine. The size of the protein ladders, OFD1, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data8-v2.zip
-
Figure 5—source data 9
Overlay of the bright field and chemiluminescence images indicating the signal of HDAC6.
RSP stands for reserpine. The size of the protein ladders, HDAC6, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data9-v2.zip
-
Figure 5—source data 10
Overlay of the bright field and chemiluminescence images indicating the signal of β-Actin.
RSP stands for reserpine. The size of the protein ladders, β-Actin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-data10-v2.zip
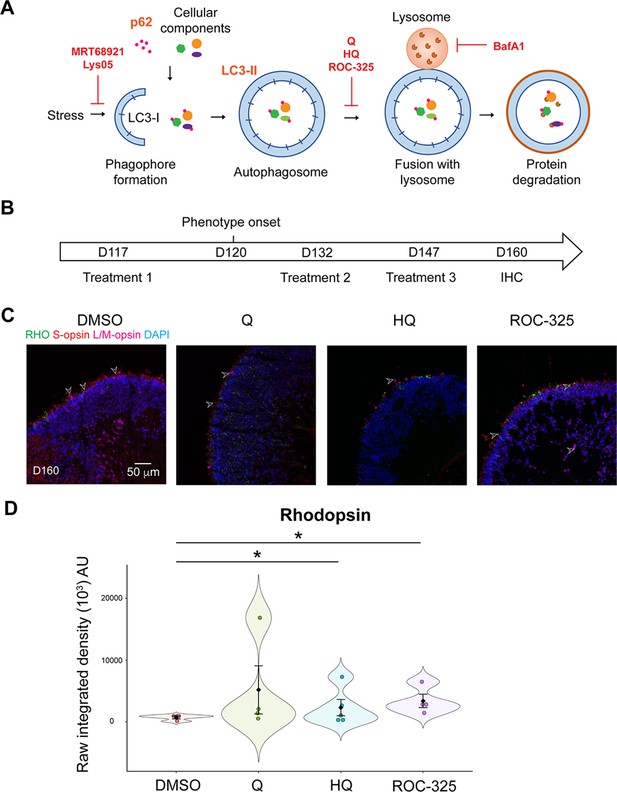
Effect of autophagy inhibitors on patient organoids.
(A) Schematic diagram of autophagy with annotation of the drug effect of autophagy inhibitors. (B) Timeline of treatment and sample harvest. (C) Immunostaining of rhodopsin (green), S-opsin (red), and L/M-opsin (magenta) were used to evaluate the effect of chloroquine (Q), hydroxychloroquine (HQ), and ROC-325. Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least two independent experiments, each of which had at least three organoids. (D) Quantification of fluorescence intensity of rhodopsin immunostaining. The shape of the Bee Swamp plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. The plot summarizes at least three independent experiments with at least one image quantified in each batch. Kruskal-Wallis test followed by Dunn’s test was used to compare the groups. *p<0.05.
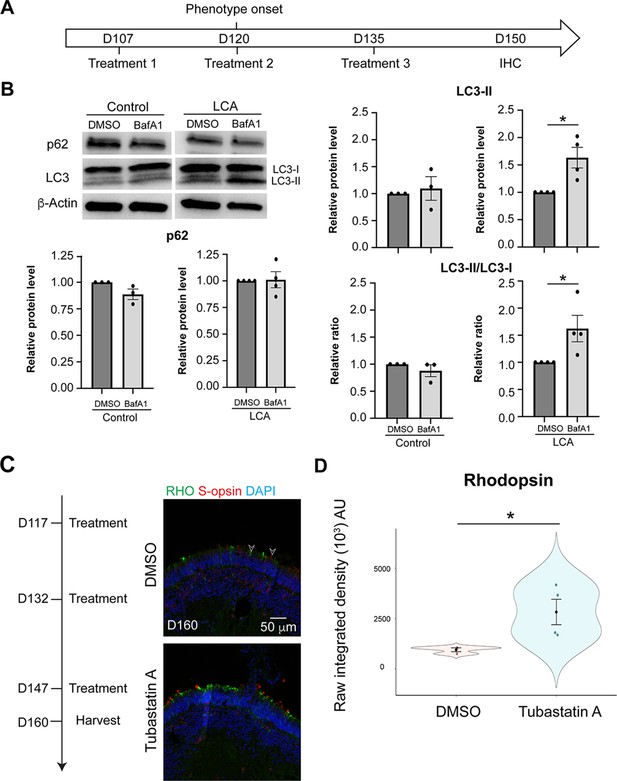
Bafilomycin A1 (BafA1) and Tubastatin A treatment on patient organoids.
(A) Timeline of BafA1 treatment. (B) Western blot and quantification of p62, LC3-II, and LC3-II/LC3-I ratio upon bafilomycin A1 (BafA1) treatment. The drug vehicle dimethylsulfoxide (DMSO) was added to the cultures in the untreated group at the same volume as the drug. β-Actin was used as the loading control. The histograms summarize data in at least three batches of experiments, each of which contained at least three retinal organoids per group. Each dot in the histogram shows data from one batch of experiment and are presented as the mean ± standard error. Unpaired t-test was performed to compare the untreated and treated groups. Unpaired t-test was used to compare the group. *p<0.05. (C) Treatment timeline (left) and outcome of Tubastatin A treatment (right) as shown by immunostaining of rhodopsin (green) and S-opsin (red). In all images, nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least two independent experiments, each of which had at least three organoids. (D) Quantification of fluorescence intensity of rhodopsin immunostaining. The shape of the Bee Swamp plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. The plot summarizes at least three independent experiments, each of which contained at least with three organoids. At least one image is quantified in each batch. Unpaired t-test was used to compare the groups. *p<0.05.
-
Figure 5—figure supplement 2—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of LC3.
BafA1 stands for Bafilomycin A1. The size of the protein ladders, LC3, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp2-data1-v2.zip
-
Figure 5—figure supplement 2—source data 2
Overlay of the bright field and chemiluminescence images indicating the signal of p62.
BafA1 stands for Bafilomycin A1. The size of the protein ladders, p62, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp2-data2-v2.zip
-
Figure 5—figure supplement 2—source data 3
Overlay of the bright field and chemiluminescence images indicating the signal of β-Actin.
BafA1 stands for Bafilomycin A1. The size of the protein ladders, b-Actin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp2-data3-v2.zip
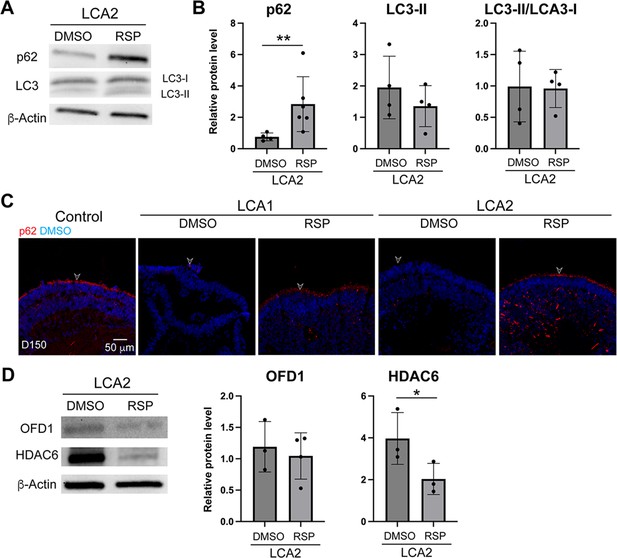
LCA2-derived retinal organoids in response to reserpine (RSP) treatment.
(A) Western blot analyses of p62 and LC3 in dimethylsulfoxide (DMSO)- and RSP-treated LCA2 organoids. β-Actin was used as the loading control. (B) Quantification of immunoblots of p62 and LC3. The histograms summarize data in at least four batches of experiments, each of which had at least three retinal organoids per group. Each dot in the histogram shows data from one batch of experiment and are presented as the mean ± standard deviation. Mann-Whitney test was used to compare the groups. *p<0.05. (C) Immunostaining of p62 in control, untreated, and treated patient retinal organoids. Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. (D) Western blot analyses and quantification of ciliary marker OFD1 and HDAC6. Data in the histogram were summarized from data from three batches of differentiation, each of which has at least three organoids. Unpaired t-test was used to compare the groups. *p<0.05.
-
Figure 5—figure supplement 3—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of p62.
The size of the protein ladders, p62, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp3-data1-v2.zip
-
Figure 5—figure supplement 3—source data 2
The size of the protein ladders, p62, and relevant sample identity are labeled.
The size of the protein ladders, b-Actin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp3-data2-v2.zip
-
Figure 5—figure supplement 3—source data 3
Overlay of the bright field and chemiluminescence images indicating the signal of OFD1.
The size of the protein ladders, OFD1, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp3-data3-v2.zip
-
Figure 5—figure supplement 3—source data 4
Overlay of the bright field and chemiluminescence images indicating the signal of HDAC6.
The size of the protein ladders, HDAC6, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig5-figsupp3-data4-v2.zip
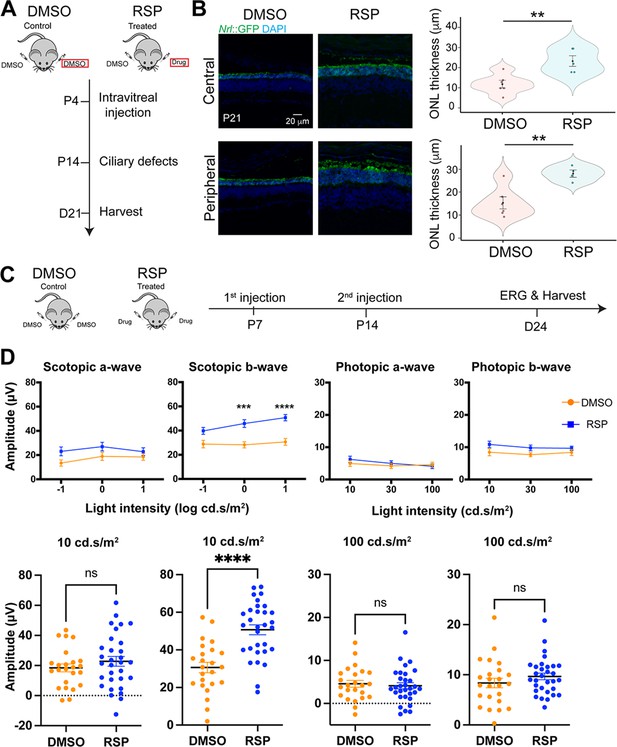
Neuroprotective effect of reserpine (RSP) in rd16 mouse retina.
(A) Timeline of in vivo intravitreal injection for quantification of outer nuclear layer (ONL), morphology, and immunohistochemistry studies. The eyes used for subsequent analyses were highlighted by red rectangles. (B) Immunostaining of dimethylsulfoxide (DMSO)- and reserpine (RSP)-treated rd16 retina (left) and quantification of the GFP+ outer nuclear layer (ONL) thickness (right). The shape of the bee swamp plot indicates the distribution of data points from four different litters, in which at least one mouse was assessed. Data points are shown in colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean. Unpaired t-test was used to compare the mean. *p<0.05. (C) Timeline of in vivo intravitreal injection for electroretinography (ERG) studies. (D) Scotopic and photopic electroretinogram responses in DMSO- and RSP-treated rd16 mice. The lower panel shows individual values of a- and b-wave amplitudes at 10 cd.s/m2 (scotopic) or at 100 cd.s/m2 (photopic). ERG was measured in both eyes of at least one mouse from five different litters in each group (12 DMSO-treated and 16 RSP-treated mice). Data were expressed as mean ± SEM, and the Mann-Whitney U test was used to compare DMSO- and RSP-treated groups. ***p<0.001; ****p<0.0001.
-
Figure 6—source data 1
Individual values of a- and b-wave amplitudes (µV) of scotopic and photopic electroretinogram responses in DMSO- and RSP-treated rd16 mice.
RSP stands for reserpine.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig6-data1-v2.zip
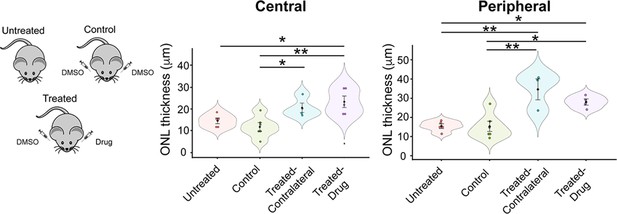
Evaluation of reserpine (RSP) injection on photoreceptor layer in rd16 mouse retina.
Evaluation of the impact of reserpine on outer nuclear layer (ONL) thickness of the central and peripheral retina. Each dot in the bee swamp plot indicates the ONL thickness of one retina from one mouse. The shape of the plot indicates the distribution of data points, which are shown by colorful circles in the center. The black diamond indicates the mean, and the error bar reveals the standard error of the mean.
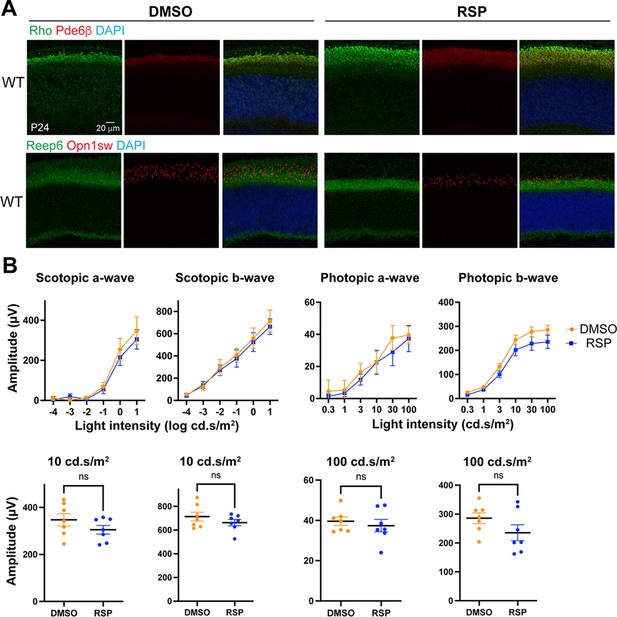
Evaluation of reserpine (RSP) injection on photoreceptor structure and function in the wild-type (WT) mouse retina.
(A) Immunostaining of rod phototransduction protein rhodopsin (Rho, green), Pde6β (red), rod cell marker Reep6 (green), and cone opsin Opn1sw (red). Nuclei were stained by 4',6-diamidino-2- phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three mice, each of which was from a different litter. (B) Scotopic and photopic electroretinogram responses in dimethylsulfoxide (DMSO)- and RSP-treated wild-type (WT) mice. The lower panel shows individual values of a- and b-wave amplitudes at 10 cd.s/m2 (scotopic) or at 100 cd.s/m2 (photopic). Electroretinography (ERG) was measured in both eyes of at least four mice from different litters in each group. Data were expressed as mean ± SEM, and the Mann-Whitney U test was used to compare DMSO- and RSP-treated groups. ***p<0.001; ****p<0.0001.
-
Figure 6—figure supplement 2—source data 1
Individual values of a- and b-wave amplitudes (µV) of scotopic and photopic electroretinogram responses in DMSO- and RSP-treated WT mice.
RSP stands for reserpine.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig6-figsupp2-data1-v2.zip
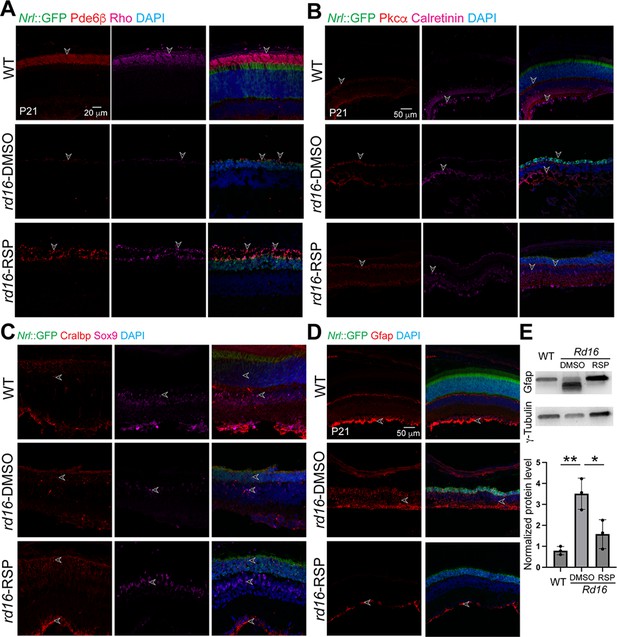
Effect of reserpine (RSP)-injection on distinct cell types in rd16 mouse retina.
Retinal structures and cell types are shown by (A) rod phototransduction protein Pde6β (red) and rhodopsin (Rho, magenta), (B) PKCα (red) for bipolar cells (red) and Calretinin (magenta) for amacrine cells, (C) Cralbp (red) and Sox9 (magenta) for Müller glia, and (D) Gfap (red) for cellular stress. Nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three mice, each of which was from a different litter. (E) Western blot analyses and quantification of Gfap. The histogram summarizes at least three retinas from three mice of different litters. One-way ANOVA followed by Tukey’s test was used to compare the groups. *p<0.05; **p<0.01.
-
Figure 6—figure supplement 3—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of Gfap.
The size of the protein ladders, Gfap, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig6-figsupp3-data1-v2.zip
-
Figure 6—figure supplement 3—source data 2
Overlay of the bright field and chemiluminescence images indicating the signal of g-Tubulin.
The size of the protein ladders, g-Tubulin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig6-figsupp3-data2-v2.zip
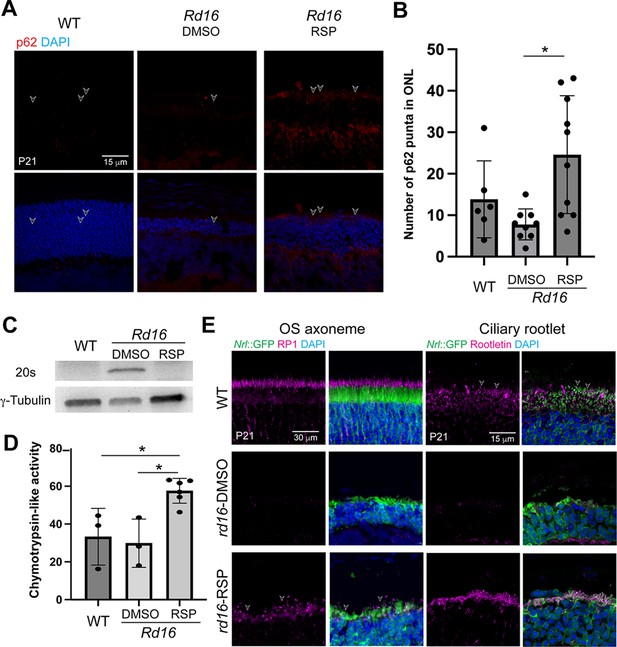
Rd16 mouse retina in response to reserpine (RSP) treatment.
(A) p62 level in dimethylsulfoxide (DMSO)- and RSP-treated photoreceptors shown by immunostaining. (B) Quantification of p62 puncta. At least three non-overlap regions were quantified in at least one mouse from at least two different litters. One-way ANOVA followed by Tukey’s test was used to compare different groups. *p<0.05. (C) Western blot analysis of 20S proteasome in wild-type (WT), DMSO-, and RSP-treated retina of rd16 mice. γ-Tubulin was used as the loading control. (D) Proteasomal chymotrypsin-like activity in rd16 retina. Data in the histogram were summarized from at least three mice from different litters and are presented as mean ± standard deviation. One-way ANOVA followed by Tukey’s test was used to compare different groups. *p<0.05. (E) Immunostaining of outer segment (OS) axoneme marker RP1 (magenta, left panel) and ciliary rootlet marker Rootletin (magenta, right panel). In (A) and (E), nuclei were stained by 4',6-diamidino-2-phenylindole (DAPI). Arrowheads indicate relevant staining. Images were representative of at least three mice from different litters.
-
Figure 7—source data 1
Overlay of the bright field and chemiluminescence images indicating the signal of 20S proteosome.
WT and RSP stand for wild-type and reserpine respectively. The size of the protein ladders, 20 S proteosome, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig7-data1-v2.zip
-
Figure 7—source data 2
Overlay of the bright field and chemiluminescence images indicating the signal of γ-Tubulin.
WT and RSP stand for wild-type and reserpine respectively. The size of the protein ladders, γ-Tubulin, and relevant sample identity are labeled.
- https://cdn.elifesciences.org/articles/83205/elife-83205-fig7-data2-v2.zip
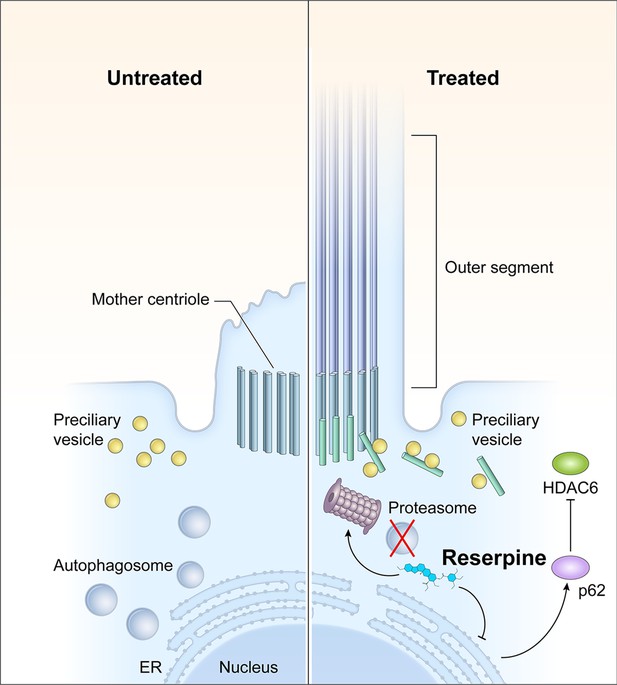
Action mechanisms of reserpine in photoreceptor survival.
CEP290 mutations lead to defects in the outer segment biogenesis and consequently, ciliary vesicles carrying building blocks of the primary cilium and ciliary proteins are accumulated in patient photoreceptors, leading to the activation of autophagy to degrade unwanted materials. As an autophagy inhibitor, reserpine downregulates autophagy and increases the p62 level in photoreceptors. As p62 is a mediator and cargo adaptor of the ubiquitin-proteasome system and autophagy, upregulation of p62 not only activates the 20 S proteasome to facilitate the clearance of the accumulated autophagosome but also facilitates the degradation of histone deacetylase 6 (HDAC6), which deacetylates microtubules. Removal of HDAC6 in photoreceptors thus should improve the stability of intracellular microtubules and facilitate the transport of pre-ciliary vesicles to the mother centriole for outer segment biogenesis.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (M. musculus) | Nrl-GFP WT | PMID: 27667917 | Reprogrammed from mouse embryonic fibroblasts by lentivirus carrying Yamanaka factors | |
| Cell line (M. musculus) | Nrl-GFP rd16 line 1 | PMID: 30332642 | Reprogrammed from mouse embryonic fibroblasts by lentivirus carrying Yamanaka factors | |
| Cell line (M. musculus) | Nrl-GFP rd16 line 2 | PMID: 30332642 | Reprogrammed from mouse embryonic fibroblasts by lentivirus carrying Yamanaka factors | |
| Cell line (Homo-sapiens) | Control | PMID: 28700940 | Reprogrammed from control fibroblasts by Sendai virus carrying Yamanaka factors | |
| Cell line (Homo-sapiens) | LCA-1 | PMID: 28700940 | Reprogrammed from control fibroblasts by Sendai virus carrying Yamanaka factors | |
| Cell line (Homo-sapiens) | LCA-2 | In this paper | Reprogrammed from control fibroblasts by Sendai virus carrying Yamanaka factors and maintained in Swaroop lab in National Eye Institute, National Institutes of Health, United States | |
| Cell line (Homo-sapiens) | LCA-1-clone C | In this paper | Reprogrammed from control fibroblasts by Sendai virus carrying Yamanaka factors and maintained in Swaroop lab in National Eye Institute, National Institutes of Health, United States | |
| Cell line (Homo-sapiens) | LCA-2-clone F | PMID: 28700940 | ||
| Antibody | anti-RHO (mouse monoclonal) | Gift of Dr. Robert Molday, University of British Columbia | IF(1:500) | |
| Antibody | anti-OPN1SW (Goat polyclonal) | Santa Cruz | Cat#: sc-14363 | IF(1:500) |
| Antibody | anti-GT335 (Mouse monoclonal) | AdipoGen | Cat#: AG-20B- 0020 | IF(1:500) |
| Antibody | anti-Rootletin (Chicken polyclonal) | PMID: 12427867 | IF (1:200) | |
| Antibody | anti-ARL13B (Rabbit polyclonal) | Proteintech | Cat#: 17711–1-AP | IF (1:500) |
| Antibody | anti-OPN1LMW (Rabbit polyclonal) | Millipore | Cat#: AB5405 | IF (1:500) |
| Antibody | anti-CEP290 (Rabbit monoclonal) | Bethyl laboratories | Cat#: A301-659A | WB (1:500) |
| Antibody | anti-GAPDH (Mouse monoclonal) | Millipore | Cat#: G8795 | WB (1:1000) |
| Antibody | anti-PDE6b (Rabbit polyclonal) | PMID: 33107904 | IF (1:500) | |
| Antibody | anti-Synaptophysin (Mouse monoclonal) | Abcam | ab8049 | IF (1:200) |
| Antibody | anti-p62 (Rabbit polyclonal) | Abcam | ab109012 | IF (1:200) WB (1:500) |
| Antibody | anti-LC3A/B (D3U4C) (Rabbit monoclonal) | Cell Signaling | 12741 | For human retinal organoids, WB (1:1000) |
| Antibody | anti-LC3B (Rabbit polyclonal) | Abcam | ab51520 | For mouse retina, WB (1:1000) |
| Antibody | anti-b-Actin (Mouse monoclonal) | Millipore | A5316 | WB (1:5000-1:10000) |
| Antibody | anti-20S (Rabbit polyclonal) | Enzo Life Sciences | BML-PW8155-0100 | WB (1:500) |
| Antibody | anti-IFT88 (Rabbit polyclonal) | Proteintech | 13967–1-AP | WB (1:500) |
| Antibody | anti-BBS6 (Rabbit polyclonal) | Michel Leroux | #17 | WB (1:500) |
| Antibody | anti-CEP164 (Rabbit polyclonal) | GeneTex | GTX85298 | WB (1:500) |
| Antibody | anti-OFD1 (Rabbit polyclonal) | Thermo | PA5-115684 | WB (1:500) |
| Antibody | anti-HDAC6 (Rabbit polyclonal) | Abgent | AP1106A | WB (1:1000) |
| Antibody | anti-RP1 (Chicken polyclonal) | PMID: 11773008 | IF (1:2000) | |
| Antibody | anti-Bassoon (Rabbit monoclonal) | Cell Signaling | 6897 S | IF (1:200) |
| Antibody | anti-CHX10 (Sheep polyclonal) | Abcam | ab16141 | IF (1:200) |
| Antibody | anti-PKCa (Rabbit monoclonal) | Thermo Fisher | MA1-157 | IF (1:1000) |
| Antibody | anti-CRALBP (Mouse monoclonal) | Abcam | ab15051 | IF (1:200) |
| Antibody | anti-GFAP (Rabbit polyclonal) | Dako | Z0334 | IF (1:500) WB (1:1000) |
| Antibody | anti-Calretinin (Rabbit monoclonal) | Millipore | MAB1568 | IF (1:200) |
| Commercial kits | Proteasome 20 S Activity Assay Kit | Sigma-Aldrich | MAK172-1KT | |
| Chemical compound, drug | Reserpine | Sigma Aldrich | 06859 | |
| Chemical compound, drug | MRT68921 | Sigma Aldrich | SML1644 | |
| Chemical compound, drug | Lys05 | Sigma Aldrich | SML2097 | |
| Chemical compound, drug | Chloroquine | Sigma Aldrich | C6628 | |
| Chemical compound, drug | Hydroxychloroquine | Sigma Aldrich | H0915 | |
| Chemical compound, drug | ROC-325 | Selleck | S8527 | |
| Chemical compound, drug | Bafilomycin A1 | Sigma Aldrich | 19–148 | |
| Chemical compound, drug | Tubastatin A | Sigma Aldrich | SML0044 | |
| Software, algorithm | Photoshop CC 2019 | Adobe | ||
| Software, algorithm | FIJI | FIJI | ||
| Software, algorithm | R | R-core team | ||
| Software, algorithm | Biowulf Linux cluster | National Institutes of Health | http://biowulf.nih.gov | |
| Other | DAPI stain | Invitrogen | D1306 | (1 µg/mL) |





