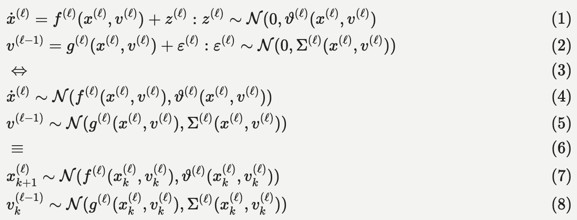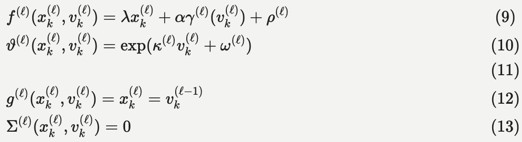Peer review process
Not revised: This Reviewed Preprint includes the authors’ original preprint (without revision), an eLife assessment, and public reviews.
Read more about eLife’s peer review process.Editors
- Reviewing EditorPeter KokUniversity College London, London, United Kingdom
- Senior EditorMichael FrankBrown University, Providence, United States of America
Reviewer #1 (Public review):
I enjoyed reading this long but compelling account of the new (generalised) version of the Hierarchical Gaussian filter (HGF). Effectively, it describes an extension of the HGF to accommodate the influence of latent states on volatility - and vice versa. This paper describes a generalisation that has been made available to the community via the TAPAS software. This contribution will be of special interest to people in computational psychiatry, where the application of the HGF has been the most prevalent.
I thought the background, motivation, description and illustration of the scheme were excellent. The paper is rather long; however, it serves as a useful technical reference.
There are two issues that I think the authors need to address.
(1) The first is the failure to properly relate the current scheme to standard implementations of Bayesian filtering under hierarchical state-space models.
(2) The second is that whilst the paper is well-written, some of the mathematical notation is cluttered. Furthermore, I think that the authors need to motivate the otherwise overengineered description of the requisite variational message passing and decomposition into update steps.
I think that the authors can address both of these issues by including a technical section in the introduction, relating the HGF to state-of-the-art in the broader field of Bayesian filtering and predictive coding. They can then explain the benefits of the particular generative model - to which the HGF is committed - by drilling down on the update scheme and its implementation in the remainder of the paper.
I was underwhelmed by the account of predictive coding and its relationship to Bayesian filtering. I think that the authors should suppress the references to predictive coding in the recent machine learning literature. Rather, the presented narrative should emphasise the fact that predictive coding and Bayesian filtering are the same thing. The authors could then explain where the hierarchical Gaussian filter fits within Bayesian filtering and why its particular form lends itself to the variational updates they subsequently derive.
The authors could add something like the following to the introduction (accompanying PDF has the equations). There is a summary of what follows in the Wikipedia entry on generalised filtering, in particular, its relationship to predictive coding (https://en.wikipedia.org/wiki/Generalized_filtering).
Relationship to Existing Work
Technically, the hierarchical Gaussian filter is a Bayesian filter under a hierarchical state-space model. The most general form of these models can be expressed as stochastic differential or difference equations as follows, c.f., Equation 9 in (Feldman and Friston, 2010):
This functional form implies a hierarchical decomposition into hierarchical levels (l) that are linked through latent causes (v), with dynamics among latent states (x) at each level. From the perspective of the HGF, the state-dependency of state (z) and observation (e) noise at each level is a key feature. The variance (i.e., inverse precision) of the random fluctuations z is known as volatility, which - in a hierarchical setting - can depend upon latent causes and states at higher levels. The variational inversion of these models - sometimes called variational or generalised filtering - finds a number of important applications: a key example here is dynamic causal modelling, typically in the analysis of imaging timeseries. In this setting, unknown or latent states, parameters and precisions are updated in variational steps by minimising variational free energy (a variational bound on negative log marginal likelihood).
In engineering, the simplest form of generalised filtering is known as a Kalman filter, in which all the equations are linear, and volatility is assumed to be constant. In neurobiology, there is an intimate relationship between generalised filtering and predictive coding: predictive coding was originally introduced for timeseries analysis and compression of sound files (Elias, 1955). Subsequently, the implicit filtering or compression scheme was considered as a description of neuronal processing in the retina (Srinivasan et al., 1982) and then cortical hierarchies (Mumford, 1992; Rao, 1999; Rao and Ballard, 1999). The formal equivalence between predictive coding and Kalman filtering was noted in (Rao, 1999). Kalman filtering itself was then recognised as a special case of generalised filtering that could be read as predictive coding in the brain (Friston and Kiebel, 2009). The estimation of precision in these predictive coding schemes has been associated with endogenous (Feldman and Friston, 2010) and exogenous (Kanai et al., 2015) attention; i.e., with and without state dependency, respectively. Subsequently, precision estimation or uncertainty quantification has become a key focus in computational psychiatry.
In machine learning, there have been recent attempts to implement predictive coding via the minimisation of variational free energy under generative models with the functional form of conventional neural networks: e.g., (Millidge et al., 2022; Salvatori et al., 2022). However, much of this work is nascent and does not deal with dynamics or volatility. There is an interesting exception in machine learning, namely, transformer architectures, where the attention heads can be read as implementing a form of Kalman gain, namely, estimating state-dependent precision, e.g., (Buckley and Singh, 2024).
Within this general setting, the HGF emphasises the importance of precision estimation or uncertainty quantification by committing to a particular functional form for the generative model that can be summarised as follows:
"We will unpack this form below and show how it leads to a remarkably compact and efficient Bayesian belief updating scheme. We will appeal implicitly to variational message passing on factor graphs (Dauwels, 2007; Friston et al., 2017; Winn and Bishop, 2005) to decompose message passing between nodes and, crucially, within-node computations. These computations furnish a scalable and flexible form of generalised Bayesian filtering. In principle, this scheme inherits all the biological plausibility of belief propagation and variational message passing in cortical hierarchies (Friston et al., 2017)."
It might be worth the authors [re-]reading the abstracts of the above papers, for a clearer sense of how those in computational neuroscience and state-space modelling (but not machine learning) think about predictive coding and its relationship to Bayesian filtering. They could then go through the manuscript, nuancing your discussion of the intimate relationship between variational Bayes, generalised filtering, predictive coding and hierarchical Gaussian filtering.
Reviewer #2 (Public review):
Summary:
The authors introduce a generalised HGF featuring (1) volatility coupling (rate of change), value coupling (phasic or autoregressive drift) [and 'noise coupling', which is a volatility parent of an outcome state] (2) parameters: volatility coupling κ, tonic volatility ω, value coupling α, tonic drift ρ, {plus minus}auto-regressive drift λ (3) inputs at irregular intervals (but still discrete time steps, unlike continuous time belief evolution in predictive coding) (4) states with multiple parents or parents with multiple child states (5) value parents by default have a volatility parent, and volatility parents have a value parent (or none) (6) linear or non-linear (including ReLU) functions (7) also beliefs can be any exponential family distribution (incl binary, categorical), hence can also model POMDPs
They describe the 3 steps involved in updating (for both value and volatility): (1) prediction (2) update posterior (entails passing both pwPE and prediction precision from lower to upper node - the latter is not found in other predictive coding schemes) (3) prediction error NB this makes the network modular, so nodes can be added/removed without recomputing all the update equations.
They give some examples of models working using simulated data: (1) sharing of parent nodes can generalise an update from one context to another (2) sharing of child nodes enables multisensory cue combination (e.g. auditory-visual, or interoceptive-exteroceptive).
The authors further discuss a potential shortcoming of the HGF - its discretisation of timesteps - which is less naturalistic but nevertheless makes it very amenable to fitting trial-wise experimental data. They propose to extend the HGF to modelling within-step dynamics in future, which could make testable continuous time neuronal predictions.
Strengths:
Overall, I think the paper is excellent - it contributes an important extension to a popular modelling tool which substantially increases the number of potential applications. It is well written, and I have almost no criticisms to make.
Weaknesses:
The authors state that this generalised HGF will "make it easy to build large networks with considerable hierarchical depth", comparable to neural network architectures. The examples they give are extremely simple; however, it would be good to see a more complex one.
Reviewer #3 (Public review):
Summary:
In this paper, Weber and colleagues develop a generalization of the HGF, a widely used modeling tool. The generalization allows coupling between latent variables that was not possible in the original HGF. The resulting inference algorithm invites a predictive coding interpretation. The modular structure allows the construction of complex models out of simpler building blocks.
Strengths:
Overall, I think this is a valuable technical contribution, which will have applications to neuroscience, behavior, and psychiatry. It is mathematically rigorous, and the exposition is, for the most part, clear. It also comes with open-source software, so it should be a valuable resource to the modeling community.
Weaknesses:
My main concern is that the way that this paper is written will only be accessible and interesting to a niche audience interested in particular kinds of approximate inference schemes. The paper doesn't draw out the implications until the very end, so it's hard for readers to understand the motivation for certain modeling choices. It also requires readers to work through many pages of math before getting to applications. The applications themselves are very abstract.