By Ian Mulvany, eLife Head of Technology (i.mulvany@elifesciences.org, @ianmulvany)
Exploring new ways to evaluate and publish great science is at the heart of what we do at eLife. We are particularly interested in how new technologies can help and have recently achieved a significant landmark that we hope will help us, and others, improve how science is published online.
Until recently, the eLife journal has been hosted by Highwire Press, who helped us to build and launch the journal website under a very tight schedule in 2012. However, to fully explore how we might bring science to life in a digital environment, we need to have as much control as possible over the tools that we use. So, last year, we made the decision to build our own publishing platform: eLife Libero. The name comes from our desire to create a platform that fully supports continuous publication, and that will continually develop, which is part of the eLife philosophy.
The decision to develop our own infrastructure also came naturally once we had put into place a number of small, purpose-built solutions necessary to accommodate our growth – in publications and type of content, and demand for new functionalities.
Committing to the development of our own entire infrastructure, however, relied on a few key factors:
- Clear communication between the developers and other stakeholders
- More than doubling the capacity of our development team
- Negotiating priorities and expectations across the entire company.
Having made the decision to build our own platform, and having put these things in place, we switched to eLife Libero in February. The transition went extremely smoothly and people outside of eLife noticed no change whatsoever; the mark of a really great transition.
Features
eLife Libero opens the door to a number of handy new features:
- We can now do true continuous publication, and introduce content at any hour, on any day. We also have full support for new article versions even after the final version of record is published.
- We have full control of the conversion of our article XML to HTML, and can control the pace at which we update our XML in order to adopt best practices for tagging, such as the recommendations coming out of JATS4R.
- We can make copies of our site easily for testing and development using Amazon Web Services, which we leant on heavily while building the platform.
- Our content processing workflow is extensible, and the reporting dashboard can be modified in the future to show information about the full life-cycle of an article, from submission through to post-publication activities, such as deposit to PubMed Central.
The most direct result of creating our own infrastructure is that our production team have support in their time zone, so any problems with publication can be dealt with immediately.
We are also in control of our own prioritisation, making problem-solving as well as experimenting with new ideas faster, and enabling us to deliver more effectively on eLife’s mission: to help scientists accelerate discovery by operating a platform for research communication that encourages and recognises the most responsible behaviours in science.
The following diagram gives a high-level overview of what we have built. eLife Libero is composed of blocks that receive data from our content processor, prepare it for publication, provide a publishing dashboard for our production team, and finally publish that content onto our website.
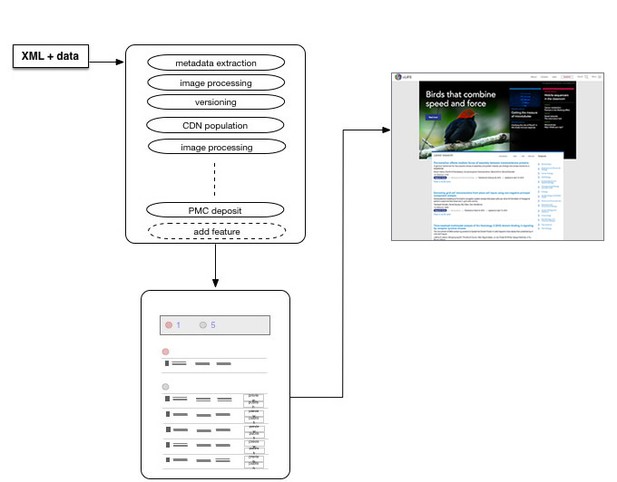
Fig1. Our new system receives article packages from our content processor, runs them through a number of steps, provides a preview via the publishing dashboard, and then, on publication, makes that content available on our site.
Goals
When we started developing the system, we set out a few goals:
- To publish articles from the publishing dashboard within one minute
From receipt of the data from our content processors, an article takes about five minutes to run through the system, and can be published instantaneously from the dashboard. - A system that is a delight to use
Our production team are happy with the system we have created, saying, “The interface is great, very unfussy and easy to use.” - To be stable and flexible
Our use of a workflow-based system means we can add features to it easily, and we have a roadmap of features that we are working on. - To have software tests in all critical parts of the system
While we have the capacity to create copies of our site for testing, the test coverage is not as extensive as we would like it to be, but we are currently increasing this. - To allow the software to be updated whenever we want
We can, but – more importantly – we hope that you may be able to help us do this, too.
Next steps
Right now, we are learning from running in a real environment, and have decided to add some new features which were not obvious to us in the design phase. We are also making the code less eLife-specific so that others may use it.
We will open-source eLife Libero over the Summer. When we release the code publicly, we will make available some of the more technical details of how the system is put together, and we will welcome suggestions for improvement, or feature requests.
Overall, it has been an exciting year for us, developing this new platform. Most of our expectations for the new system have been met and we can see how further progress may be made.
What’s really empowering is that we now control the code for our own publishing system. By having both insight and influence over the entire process, we can work towards really improving the experience of publishing science online.
We will be hosting a webinar to introduce eLife Libero to publishers, developers, and anyone else interested. We have not yet set a date. If you’d like to hear more about it, please let us know on staff@elifesciences.org.




