By Andy Collings, Executive Editor at eLife.
Norbert Perrimon approached eLife in March 2017 to see if we would be interested in introducing a Key Reagent (or Key Resources) Table in published papers, as had been done by the Cell Press STAR Methods and the GSA journals. Madeline Crosby, along with Norbert and the FlyBase Consortium, had been working with other model organism databases and had developed a prototype reagent table (presented in this F1000Research poster). The aim was to facilitate the handling of reagent sources and identifiers at multiple steps, to help researchers, journals, and biological databases.
In the view of Norbert Perrimon, “Journals have a unique opportunity to have a huge impact in promoting rigour and transparency in science by implementing Author Reagent Tables”.
Indeed, if journals can work with authors to keep better track of the resources being used, this enhances transparency (and makes it easier to locate the same resource), helps to give credit where it’s due, and it aids the curation of databases and datasets.
After consultation with our editors and some revisions to the template table, we started encouraging authors in decisions sent out in August 2017 to include a Key Resources Table when they came to resubmit. Our intention was to highlight the genetically modified organisms and strains, cell lines, reagents, and software that are essential to reproduce the results presented. However, Key Resources Tables do not need to be a comprehensive list of all the materials and resources used.
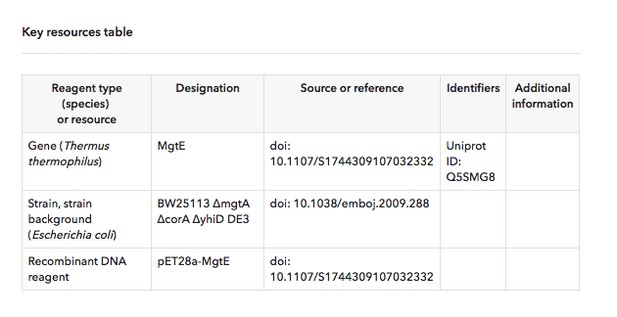
An example of a short Key Resources Table from eLife 2018;7:e31596.
While these tables may not be helpful for all forms of research, it is particularly useful for bench research. Where relevant, authors of eLife papers invited for revisions are strongly encouraged to download and complete this template for the Key Resources Table to ensure consistency. The template includes detailed notes on completion as well as an example. We indicate the details of newly generated (and previously published) datasets separately, so the resource categories in eLife’s Key Resources Tables follow:
- Gene (indicating species)
- Strain, strain background (indicating species)
- Genetic reagent (indicating species)
- Cell line (indicating species)
- Transfected construct (indicating species)
- Biological sample (indicating species)
- Antibody (including host organism common name and clonality)
- Recombinant DNA reagent (cultured clones, plasmids, cDNAs, and so on)
- Sequence-based reagent (oligonucleotides, primers, morpholinos, and so on)
- Peptide, recombinant protein
- Chemical compound, drug
- Commercial assay, kit
- Software, algorithm
- Other (this could include histological stains, among other resources)
The completed Key Resources Table should be incorporated at the beginning of the Materials and Methods section. We have been encouraged by the response from authors. In the three months covering December 2017 to February 2018, we accepted 298 research papers for publication, with 121 (41%) including a Key Resources Table. The Key Resources Table in the paper “Gq activity- and β-arrestin-1 scaffolding-mediated ADGRG2/CFTR coupling are required for male fertility” by Dao-Lai Zhang, Yu-Jing Sun, Ming-Liang Ma, Yi-jing Wang, and colleagues is a particularly good example, using catalogue numbers in some cases and RRIDs where possible.
We hope more and more authors will include a Key Resources Table going forward, and we welcome feedback or suggestions for how we can better present them, or make the process of completing and submitting them easier. You can contact us at editorial [at] elifesciences [dot] org.
For the latest in published research plus papers available in PDF shortly after acceptance sign up for our weekly email alerts. You can also follow @eLife on Twitter.




