By Adrià Mercader and Jo Barratt, Open Knowledge International, and Naomi Penfold, eLife
eLife supports the sharing of research data at the time of publication if not before. To enable the community to derive greater value from this shared data, we are interested in helping researchers improve its quality. Open Knowledge International (OKI) develops technical solutions to make it easier to share reusable data, and the organisation helped us understand the reusability of data published on eLife through the application of its goodtables analysis tool. We found that researchers tend to present data with a view to a human visually inspecting it. The reusability of datasets shared with eLife could be improved by helping researchers to prepare data in machine-friendly ways, where this is appropriate.
Open data and reusability: the next frontier
Data sharing is an important cornerstone in the movement towards more reproducible science: it provides a means to validate assertions made, which is why many journals and funders require that research data is shared publicly and appropriately within a reasonable timeframe following a research project. At eLife, authors are encouraged to deposit their data in an appropriate external repository and to cite the datasets in their article or, where this is not possible or suitable, publish the source data as supplements to the article itself. The data files are then stored in the eLife data store and made available through download links available within the article.
Source data shared with eLife is listed under the Figures and data tab. Source: Screenshot from eLife 2017;6:e29820.
Open research data is an important asset in the record of the original research, and its reuse in different contexts helps make the research enterprise more efficient. Sharing and reuse of research data is fairly common, and researchers may reuse others’ data more readily than they might share their own.
The exact nature of data reuse, however, is less clear: 40% of Wellcome Trust-funded researchers make their data available as open access, and three-quarters report reusing existing data for validation, contextualisation, methodological development and novel analyses, for example (Van den Eynden et al, 2016). Interestingly, a third of researchers who never publish their own data report reusing other researchers’ open data (Treadway et al, 2016) and dataset citation by researchers other than the original authors appears to be growing at least in line with greater availability of data (for gene expression microarray analysis; Piwowar & Vision, 2013). However, only a minority of citations (6% of 138) pertained to actual data reuse when citation context was manually classified in this study.
Making reuse of open data frictionless
OKI is a global non-profit organisation focused on realising open data’s value to society by helping civil society groups access and use data to take action on social problems.
Informed by work building and deploying the CKAN open-source data portal platform, and learning about various data publication workflows, the Frictionless Data team at OKI is investigating how to remove the barriers in obtaining, sharing and validating data – barriers that stop people from truly benefiting from the wealth of data being opened up every day. This work takes the form of a growing series of tools, standards and best practices for publishing data. One such tool is goodtables – a library and web service developed to support the validation of tabular datasets both in terms of structure and also with respect to a published schema.
Understanding data reusability at eLife using OKI’s goodtables
At csv,conf,v3 in Portland in June, which featured presentations by both Naomi Penfold of eLife and Adrià Mercader from the Frictionless Data team, it became clear that eLife, with its strong emphasis on research data, would be a great candidate for a pilot exploring the application of goodtables’ automated data validation.
In order to assess the potential for a goodtables integration at eLife, we first needed to measure the quality of source data shared directly through eLife. Using goodtables, we performed a validation on all files made available through the eLife API, and generated a report on their data quality. This allowed us to understand the current state of eLife-published data, and opened the possibility of doing more exciting things with it, such as more comprehensive tests and visualisations.
We went through the following process:
- We downloaded a large subset of the articles’ metadata made available via the eLife public API.
- We parsed all metadata files in order to extract the data files linked to each article, regardless of whether it is was an additional file or a figure source. This gave us a direct link to each data file linked to the parent article.
- We then ran the validation process on each file, storing the resulting report for future analysis.
All scripts used in the process, as well as the outputs, can be found on Github: https://github.com/frictionlessdata/pilot-elife.
We analyzed 3,910 articles, 1,085 of which had data files. The most common format was Microsoft Excel Open XML Format Spreadsheet (xlsx), with 89% of all 4,318 files being published in this format. Older versions of Excel and CSV files made up the rest.
In terms of validation, more than three quarters of the articles analyzed (844/1,085) contained at least one ‘invalid’ file. In this context, ‘invalid’ and ‘valid’ are arbitrary terms based on the tests that are set within goodtables, and the results therefore need to be carefully reviewed to understand the pertinence of these terms as representative of actual addressable issues.
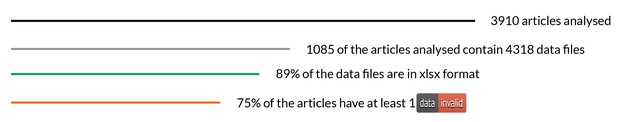
A summary of the eLife research articles analysed as part of the Frictionless Data pilot work. Open Knowledge International.
Here’s a summary of the raw number of errors encountered (for a more complete description of each, see the Data Quality Specifications):
| Blank rows | 45,748 |
| Duplicate rows | 9,384 |
| Duplicate headers | 6,672 |
| Blank headers | 2,479 |
| Missing values | 1,032 |
| Extra values | 39 |
| Source errors | 11 |
| Format errors | 4 |
Following analysis of a sample of the results, the vast majority of the errors appear to be due to the data being presented in nice-looking tables, using formatting to make particular elements more visually clear, as opposed to a machine-readable format:
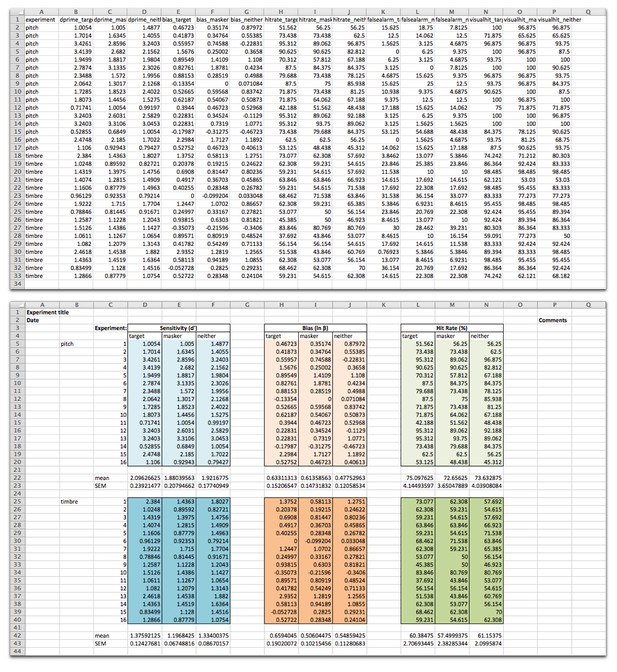
Data from Maddox et al. was shared in a machine-readable format (top), and adapted here to demonstrate how such data are often shared in a format that looks nice to the human reader (bottom). The data file is presented as is (top) and adapted (bottom, source data) from Maddox et al., eLife 2015;4:e04995, under the Creative Commons Attribution License (CC BY 4.0).
Because these types of errors are so common, particularly with Excel files, we have introduced default ‘ignore blank rows’ and ‘ignore duplicate rows’ options in our standalone validator goodtables.io to help bring more complex issues to the surface and focus attention on those which may be less trivial to resolve. Excluding duplicate and blank rows as well as duplicate headers (the most common but also relatively simple errors), 6.4% (277/4,318) of data files had errors remaining, affecting 10% of research articles (112/1,085).
Much less frequent errors were related to difficulties retrieving and opening data files. It was certainly helpful to flag articles with files that were not actually possible to open (source-error), and the eLife production team are resolving these issues. While only representing a small number of datasets, this is one key use case for goodtables: enabling publishers to regularly check continued data availability after publication.
Overall, the findings from this pilot demonstrate that there are different ways of producing data for sharing: datasets are predominantly presented in an Excel file with human aesthetics in mind, rather than structured for use by a statistical program. We found few issues with the data itself beyond presentation preferences. This is encouraging and is a great starting point for venturing forward with helping researchers to make greater use of open data.
Areas for future work
Work to improve the reusability of research data pushes towards an ideal situation where most data is both machine-readable and human-comprehensible. In general, the eLife datasets had better quality than, for instance, those created by government organisations, where structural issues such as missing headers and extra cells are much more common. So, although the results here have been good, the community may derive greater benefit from researchers going that extra mile to make files more machine-friendly and embrace more robust data description techniques like Data Packages.
Tools such as goodtables help to flag the gap between the current situation and the ideal of machine-readability, and help researchers and publishers identify how datasets could be reshaped to make them more reusable for humans and machines alike.
We would like to help researchers to share their best work. Help us to understand how you would like datasets to be shared for your reuse, whether to validate, contextualise, or form the basis for subsequent analyses:
- Do you download the datasets shared with eLife articles?
- Are the data presented in a way that is easy for you to use for your purposes?
- Are there ways in which you would prefer data to be formatted or presented?
- As an author, would you appreciate being able to present your data both in a visually clear way and in a machine-readable format? If so, would you be interested in tools that help to improve datasets in both kinds of format?
Moving forward, we are interested in tools and workflows that help to improve data quality earlier in the research lifecycle or make it easy to reshape at the point of sharing or reuse. Tools and initiatives that reduce the burden of preparing data for reuse include those designed for handling ‘Big Data’, such as OpenRefine and the Data Civilizer System.
Most of the issues identified by goodtables in the datasets shared with eLife relate to structuring the data for human visual consumption: adding space around the table, merging header cells and such like. We encourage researchers to make data as easy to consume as possible, and recognise that datasets built primarily to look good to humans may be only be sufficient for low-level reuse. For researchers interested in preparing data for sharing in a machine-readable format, these resources explain key principles to bear in mind:
- Data organisation in spreadsheets, Broman & Woo (2017)
- How to share data for collaboration, Ellis & Leek (2017)
- Ten Simple Rules for Digital Data Storage, Hart et al (2016)
- Tidy data, Wickham (2014)
- Learning resources from Data Carpentry
Get in touch
Researchers, we welcome your feedback on the potential utility of reusable data in the life sciences and suggestions for how to make it easier for you to reuse research data shared with us.
Technologists, we welcome your input regarding tools that help researchers to share reusable data with minimal effort.
Contact eLife by email (innovation [at] elifesciences.org) or Twitter (@eLifeInnovation). Contact OKI via the Discuss forum or Gitter chat.
This content is adapted and cross-posted on the OKI blog and as a goodtables case study.
For the latest in innovation, eLife Labs and new open-source tools, sign up for our technology and innovation newsletter. You can also follow @eLifeInnovation on Twitter.
Do you have an idea or innovation to share on eLife Labs? Please send a short outline to innovation [at] elifesciences.org.




