The Natural History of Model Organisms: C. elegans outside the Petri dish
Figures
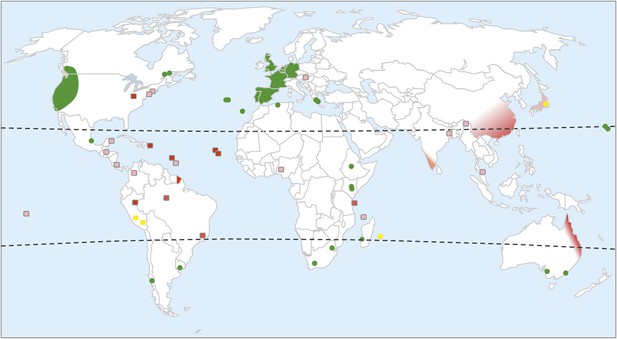
Worldwide distribution of C. elegans.
Green shading highlights areas where C. elegans has been repeatedly collected. Green dots mark islands or locations where C. elegans has been collected at least once. Yellow squares represent areas where many Caenorhabditis species have been sampled and where C. elegans is present but rare (often found at altitude). Red shading highlights where C. elegans has never been collected despite the intensive sampling of many other Caenorhabditis species. Pink shading highlights where C. elegans has not been collected, despite the sampling of several other Caenorhabditis species. White represents areas that have never been sampled for C. elegans or very rarely. The distribution is inferred from published data (Abdul Kader and Côté, 1996; Barrière and Félix, 2005a, 2005b, 2007; Dolgin et al., 2008; Wang et al., 2010; Kiontke et al., 2011; Andersen et al., 2012; Félix and Duveau, 2012; Dey et al., 2013; Félix et al., 2013), WormBase, and our lab collection (http://www.justbio.com/worms/index.php).
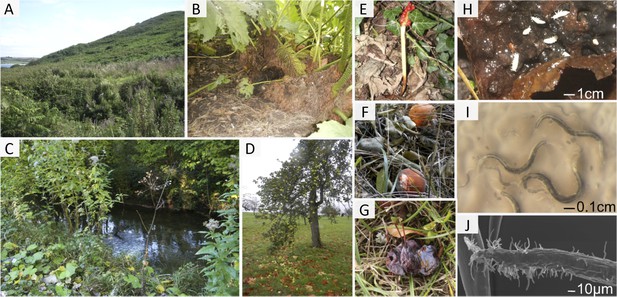

The habitat of C. elegans at different scales.
(A–D) Landscapes that correspond to the macroscale C. elegans habitat; all are relatively humid areas where C. elegans has been found: (A) wet shrubland; (B) urban garden; (C) riverbank; and (D) fruit trees. (E–G) Bacteria-rich decomposing vegetal substrates, corresponding to the microscale C. elegans habitat: (E) Arum stem; (F) oranges and (G) plums. (H) Detail of a rotting apple at the stage where C. elegans proliferates. Springtails (white) and a mite are examples of animals that share the bacteria-rich habitat of C. elegans and that are potential carriers and/or predators (see also Table 1). (I) C. elegans nematodes on an E. coli lawn, just coming out of a rotten fruit. (J) Scanning electron micrograph of C. elegans infected with the fungus Drechmeria coniospora. Image credits: Marie-Anne Félix.
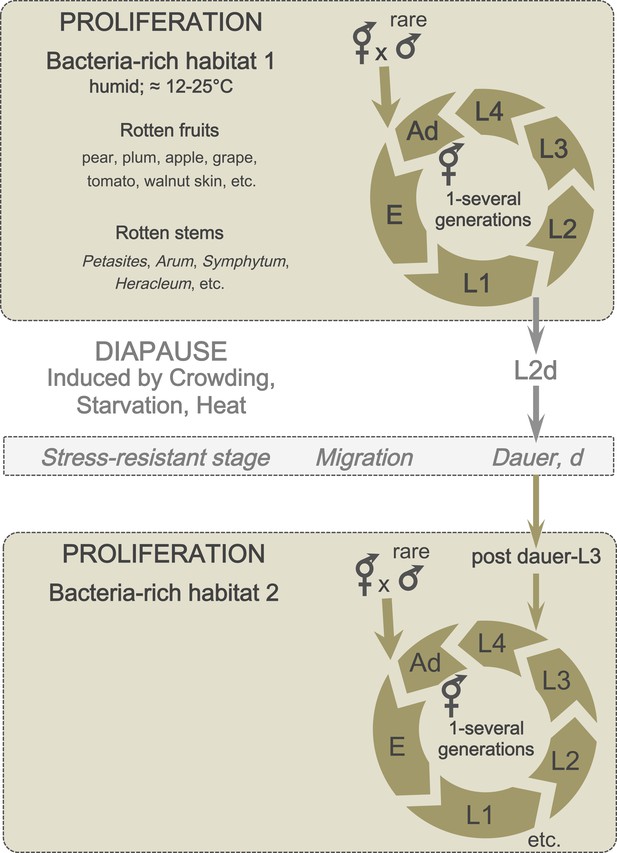
A schematic of C. elegans lifecycle in the wild.
In bacteria-rich habitats (beige), the C. elegans life cycle begins with an embryonic (E) stage, followed by four larval (L1-L4) stages, and ends with an adult stage (Ad). Most animals are self-fertilizing hermaphrodites; males are rare and breeding with males therefore uncommon. Under suboptimal conditions (such as crowding and starvation), L1 larvae can enter a predauer stage (L2d) followed by the diapause stage (dauer). When better conditions arise, dauers develop into postdauer L3 larvae and re-enter the lifecycle at the L4 stage.
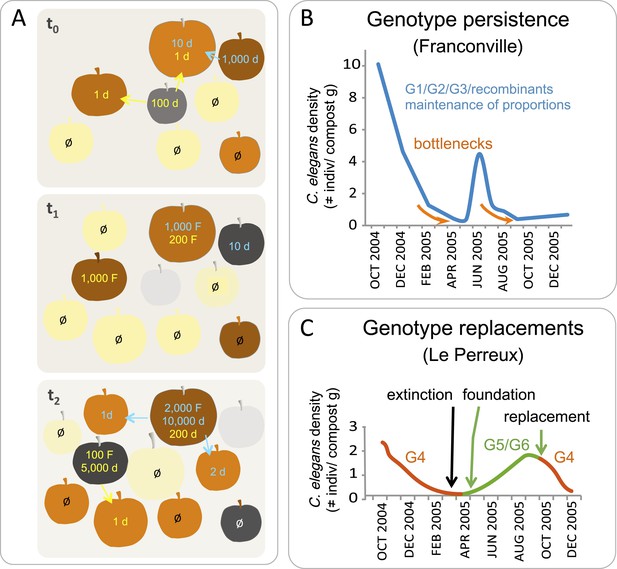
C. elegans population dynamics in a natural habitat.
(A) A schematic of C. elegans population dynamics in an orchard. Population growth on a given apple has not been monitored to date and so is inferred here from data in (Félix and Duveau, 2012), based on many time points in an orchard and single time points on a given fruit. The fruits are shown at three time points (t) in identical positions on each panel, with t0 the first and t2 the last timepoint. Fruit colors indicate the degree of fruit decomposition, from early stages (yellow) to brown and dark grey (later stages), until disappearance (light grey). The number of feeding (F) and non-feeding dauer (d) individuals are indicated in color, with different colors representing different genotypes. Ø represents no colonization of a fruit by C. elegans. Arrows indicate dauer migration. How often different fruits or stems are colonized by several genotypes remains to be tested (Barrière and Félix, 2007; Andersen et al., 2012). (B–C) Actual population dynamics at the scale of a compost heap, from Barrière and Félix (2007). (B) In the first (Franconville) example, three main genotypes, G1, G2 and G3, persist in the heap at similar frequencies over the time period shown. In the second example (Le Perreux-sur-Marne), a single genotype, G4, was present, became extinct, then two new genotypes, G5 and G6, founded a new population. G4 reappeared in September, while G8 and G9 (not shown) disappeared. Genotypes were characterized using microsatellite markers.
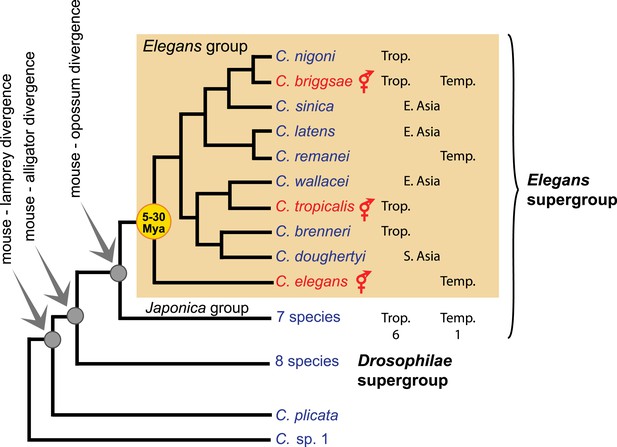
Phylogeny of the Caenorhabditis genus, with emphasis on the Elegans group, based on (Kiontke et al., 2011). Trop.: tropical distribution. Temp.: temperate. See http://worldwideworm.banshy.fr/ for geographical distributions.
Tables
The biotic environment of C. elegans
-
Some organisms may act as both food and pathogen, or as both vector and predator. Other nematodes may be competitors for food but also predators. The predatory relationships are inferred from the co-occurrence of the two species in the wild, from laboratory predation assays, and, in the case of mites and fungi, by observation on the laboratory isolation plate; specific studies would be needed to observe them directly in a natural setting.
-
*
and our unpublished observations.





