Genetic diversity affects ecosystem functions across trophic levels as much as species diversity, but in an opposite direction
Figures
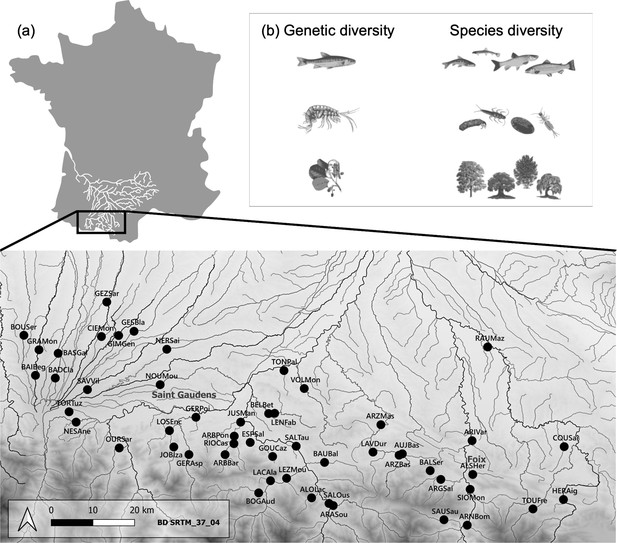
Location of sampling sites and illustration of the trophic chain.
(a) Distribution of the 52 sampling sites (black dots) spanning an east-west gradient at the foothills of the Pyrenees Mountains (France). Each site is denoted by a six-letter code, with the first three letters indicating the river name and the last three letters indicating the closest city or village. (b) Our study focused on a tri-trophic food chain commonly found in mountain rivers, consisting of riparian trees, macroinvertebrate shredders, and fishes (from bottom to top). Within each trophic level, we measured two facets of biodiversity: genetic diversity in a single target species within each trophic level (specifically P. dragarum, Gammarus sp., and A. glutinosa), and species diversity from communities.
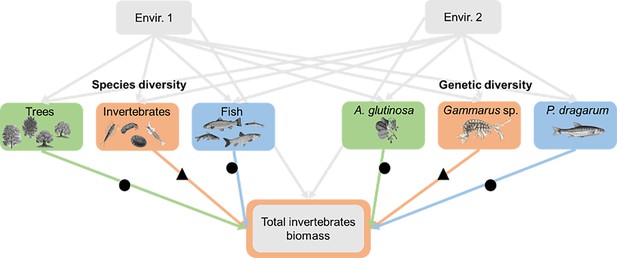
Example of one of the seven causal models used to quantify the relationships between (species and genetic) diversity and ecosystem functions.
We focused on seven ecosystem functions associated with genetic and species diversity at three trophic levels (green for primary producer, orange for primary consumer, and blue for secondary consumer). Each relationship between biodiversity and ecosystem functions (n=6 values per function, but for some functions for which irrelevant links were not considered, see the text, n=32 values in total) was measured at the same trophic level (triangles) or at another trophic level (dots).
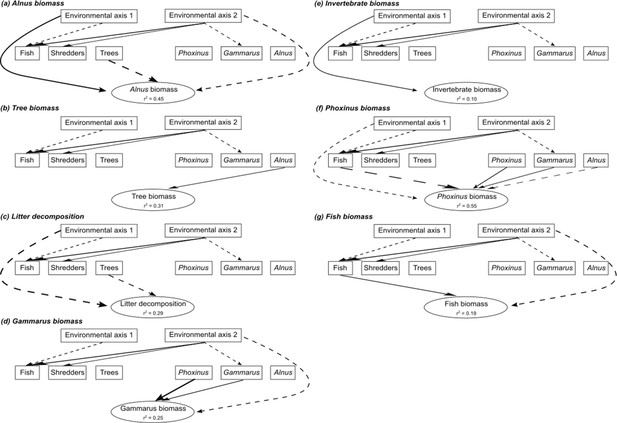
Details of the seven causal models linking abiotic parameters, species and genetic diversity, and ecosystem functions.
Each causal graph (a–g) represents a simplified illustration of the relationships between the two principal component analysis (PCA) axes synthesizing the environmental parameters of each sampling site (Environmental axis 1 and 2), the species diversity estimated at each trophic level (boxes ‘Fish’, ‘Shredders’, and ‘Trees’), the genomic diversity estimated from each focal species at each trophic level (boxes ‘Phoxinus’, ‘Gammarus’, and ‘Alnus’), and each ecosystem function (one model per function). Only the relationships for which the p-value was inferior to 0.20 are indicated for visual simplification. Full arrows indicated positive effects, whereas dotted arrows indicated negative effects. The width of the arrows is proportional to the size of their effects. The percentage of variance explained by environmental and biodiversity effects on ecosystem functions (r2) is indicated for each function.
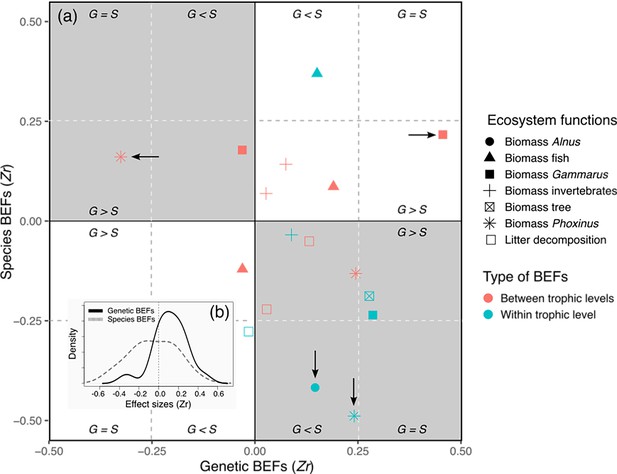
General description of individual effect sizes measured between biodiversity estimates and ecosystem functions (BEFs) in a riverine trophic chain.
(a) The magnitude and direction of individual effect sizes (Zr) of biodiversity is shown for each ecosystem functions as a biplot between Zr associated with genetic diversity (y-axis, genetic BEFs) measured for one of three target species (A. glutinosa, Gammarus sp., and P. dragarum) and Zr associated with species diversity (x-axis, species BEFs) measured for one of three trophic levels (trees, invertebrates, and fish). For each ecosystem function (but the biomass of trees and of A. glutinosa), a total of six Zr are depicted in the biplot; four of them are associated with biodiversity measured at another trophic level than the one of the target functions (red symbols, e.g. effect of fish diversity on invertebrate biomass) and two of them are associated with biodiversity measured at the same trophic level than the one of the target functions (blue symbols, e.g. effect of fish diversity on fish biomass). The arrows indicate significant Zr (95% confidence intervals excluded 0, see Supplementary file 2); vertical arrows are for significant genetic BEFs, horizontal arrows are for species BEFs. White quadrats stand for situation in which genetic and species BEFs are in the same direction, whereas gray quadrats indicate situation in which genetic and species BEFs are in the opposite direction. Within each quadrat, sub-quadrats indicate the relative magnitude of BEFs, i.e., whether genetic BEFs are stronger, weaker, or equal in magnitude than species BEFs. (b) Density plots displaying the distribution of individual Zr for species and genetic BEFs (dotted and full lines respectively).
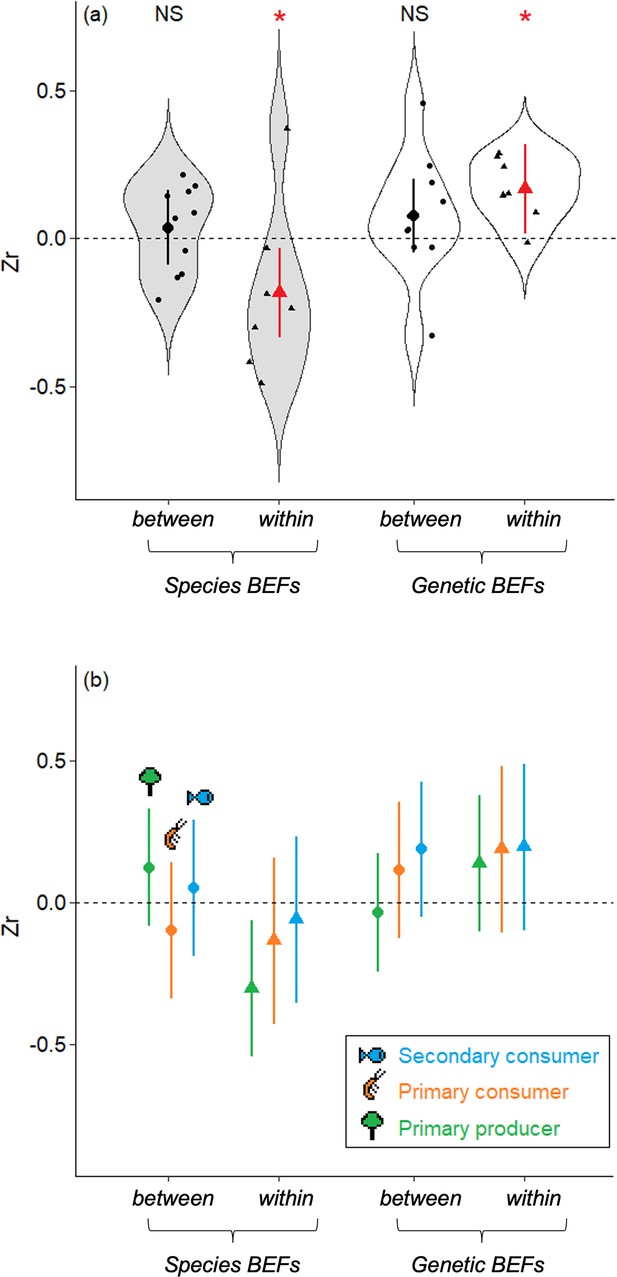
Magnitude and direction of the mean effects sizes estimated from the relationships between biodiversity and ecosystem functions (BEFs) measured in a riverine trophic chain.
(a) The magnitude and direction of BEFs are expressed as effect sizes (Zr) and are displayed according to the facet used to measure biodiversity (genetic or species diversity, light gray and white boxplots respectively) and to the type of BEFs (within-trophic level BEFs or between-trophic level BEFs, triangles and dots respectively). Red color and stars indicate global effect sizes that are significantly different from zero (p-value<0.05). Large symbols are mean ± 95%IC estimated as marginal effects from the meta-regressions. Small symbols are raw estimates. (b) Same representation as (a) but with details at each trophic level (mean ± 95% IC estimated as marginal effects from the meta-regressions, green for primary producer, orange for primary consumer, and blue for secondary consumer). The trophic level at which BEFs are measured is coherent across all trophic levels.
Tables
Characteristics of the first two principal components identified by the principal component analysis (PCA) ran on the 13 environmental variables.
The part of the total environmental variance (%) and the contribution of each variable on each component are shown. The variables that contributed significantly to the axis are highlighted in bold.
| Component 1 | Component 2 | |
|---|---|---|
| Part of total variance (%) | 21.66 | 16.37 |
| River width | 0.596 | 0.320 |
| Connectivity | –0.155 | 0.646 |
| Altitude | 0.648 | –0.385 |
| Distance from outlet | 0.528 | –0.331 |
| East-west gradient | 0.105 | –0.795 |
| Oxygen concentration | 0.738 | 0.343 |
| Oxygen saturation | 0.266 | –0.509 |
| Water temperature | –0.594 | –0.012 |
| Specific conductivity | –0.463 | –0.045 |
| pH | 0.573 | 0.398 |
| Concentration in NO3+NO2 | –0.369 | –0.286 |
| Concentration in NH4+ | –0.019 | –0.207 |
| Concentration in PO43- | –0.279 | 0.236 |
| Global characteristic | Low altitude, poorly oxygenated site – high altitude, highly oxygenated sites | Poorly connected east site – highly connected west site |
ANOVA table for the linear mixed model testing whether the relationships between biodiversity and ecosystem functions (BEFs) measured in a riverine trophic chain differ between the biodiversity facets (species or genetic diversity) and the types of BEF (within- or between-trophic levels).
A Wald chi-square test is used to test the significance of each fixed effect.
| Degree of freedom | Chisq-value | p-Value | |
|---|---|---|---|
| (Intercept) | 1 | 0.287 | 0.595 |
| Biodiversity facet | 1 | 0.232 | 0.630 |
| Type of BEF | 1 | 5.393 | 0.020 |
| Biodiversity facet*Type of BEF | 1 | 5.567 | 0.018 |
Additional files
-
Supplementary file 1
ANOVA table for the linear mixed model testing whether the relationships between biodiversity and ecosystem functions (BEFs) measured in a riverine trophic chain differ between the biodiversity facets (species or genetic diversity), the types of BEF (within- or between-trophic levels), and the trophic levels at which BEFs are estimated (primary producers, primary consumers, or secondary consumers).
A Wald chi-square test is used to test the significance of each fixed effect.
- https://cdn.elifesciences.org/articles/100041/elife-100041-supp1-v1.docx
-
Supplementary file 2
Estimates of individual effect sizes of biodiversity and ecosystem functions (BEFs) (Zr, n=34) for each ecosystem function, each biodiversity facet (genetic or species diversity), and each type of BEF (within- or between-trophic levels).
95% confidence intervals are provided together with the estimate of each BEF. BEFs are considered as significant when the 95% CI does not overlap 0. p-Values estimated from t-test are also provided.
- https://cdn.elifesciences.org/articles/100041/elife-100041-supp2-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/100041/elife-100041-mdarchecklist1-v1.docx





