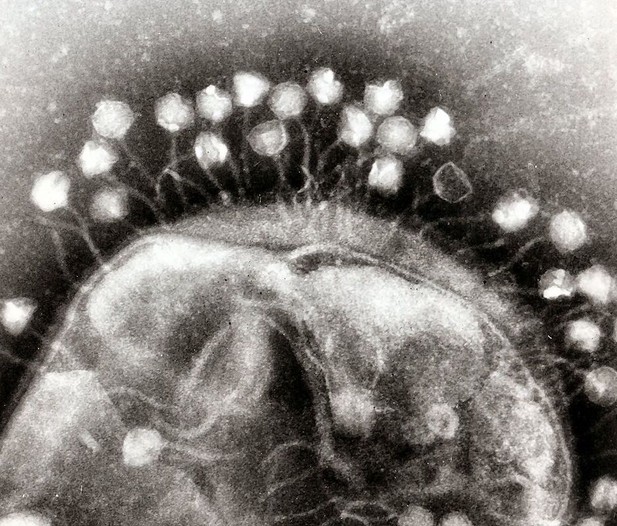
Multiple bacteriophages attached to a bacterial cell wall. Image credit: Modified from Dr Graham Beards (CC BY-SA 3.0)
Viruses that attack bacteria are known as bacteriophages, or phages for short. Bacteria have developed an antiviral immune system called CRISPR-Cas that works by targeting particular genetic sequences, such as those of an invading phage, for destruction. To counteract this immune system, phages have evolved proteins that can block CRISPR-Cas known as anti-CRISPRs.
Researchers have studied the CRISPR-Cas bacterial defense systems intensively over the past decade but much less is known about anti-CRISPRs. For example, the natural diversity and prevalence of anti-CRISPRs is still unknown, and identifying these proteins has proven difficult.
To address this gap, Forsberg et al. developed a technique to identify new anti-CRISPRs based on their ability to inhibit CRISPR-Cas activity. The method relies on three elements. First, a piece of DNA that lets bacteria resist a specific antibiotic. Second, a test piece of DNA that might code for an anti-CRISPR. Third, a CRISPR-Cas system designed to target and destroy the antibiotic resistance DNA. The three elements are put into bacteria, and two things can happen. If the ‘test DNA’ does not code for an anti-CRISPR, then the CRISPR-Cas system destroys the antibiotic resistance DNA and the bacteria die when exposed to the antibiotic. On the other hand, if the test DNA does code for an anti-CRISPR, it will inhibit the CRISPR-Cas system and the antibiotic resistance DNA will remain intact. This means that the bacteria will survive when grown in the antibiotic, and new anti-CRISPRs can be found by examining the test DNA in those bacteria.
Forsberg et al. employed this strategy to screen a huge library of DNA pieces, uncovering several new anti-CRISPRs. They then focused on an anti-CRISPR that was very common in the human gut called AcrIIA11. Biochemical characterization showed that AcrIIA11 inhibited CRISPR-Cas via a different mechanism from other known anti-CRISPRs. Moreover, it could inhibit CRISPR-Cas systems from many different bacteria.
The potential to systematically identify anti-CRISPRs able to resist any bacterium’s CRISPR-Cas defense system could lead to the design of phages that can infect bacteria which are otherwise difficult to destroy. In the future, these phages could be used to clear antibiotic-resistant infections. Beyond its role as an antiviral system in bacteria, CRISPR-Cas is a widely used tool for genetic modification in biomedical research. Using anti-CRISPRs to regulate where, how, and when CRISPR-Cas systems act could make their many emerging applications safer and more effective.




