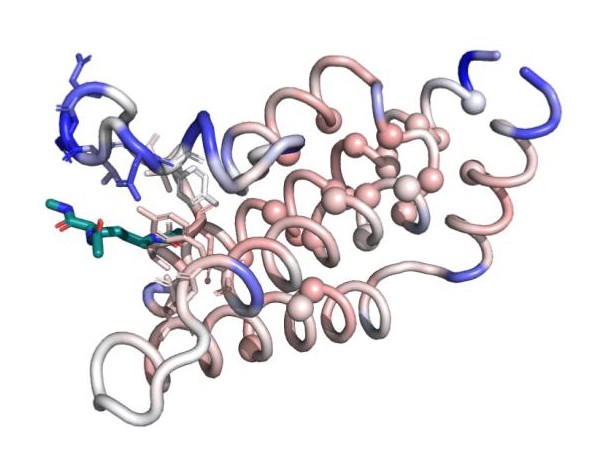
Structure of the bromodomain of the human protein ATAD2, colored according to the flexibility of each residue: blue residues are less dynamic than the average binding site residue and red residues are more dynamic than the average binding site residue. Binding site residues are represented by sticks, and the ligand is colored in teal. Image credit: Stephanie Wankowicz (CC BY 4.0)
Proteins are the workhorses of our cells. They are large molecules that ‘fold’ into specific, often highly complex, three-dimensional configurations. These structures are not static, but rather dynamic and flexible. In other words, proteins can shift between different three-dimensional shapes to perform their tasks within the cell.
To perform their roles, many proteins have to bind to small molecule ligands. Many ligands are drugs, which means that their effectiveness depends on their ability to bind to and impact the proteins involved in the disease they are treating.
When a ligand binds to a protein, it can reshape the protein. For example, certain conformations of the protein, which were difficult for the protein to be in on its own, may become more stable when the ligand binds. Additionally, upon ligand binding, some parts of the protein may move relative to each other. Previous studies have shown that these movements can affect the interaction between ligand and protein. However, these studies only examined a small number of proteins. Therefore, Wankowicz et al. set out to determine, in greater detail, what happens to protein flexibility upon ligand binding.
First, a pipeline was created to model alternative configurations of the protein both with and without ligands attached. These models measured flexibility within protein structures. The models revealed that when ligands bind to proteins, the flexibility of different regions of the protein changes – and does so in a consistent way. Proteins that become more rigid in the region interacting with their ligands become less rigid in other, distant regions, and vice versa. In other words, the rest of the protein is able to compensate for any changes in flexibility caused by ligand binding, which may contribute to how well a ligand binds to a protein.
This study demonstrates the ability of ligands to affect the entire structure of the proteins they bind to, and therefore sheds new light on the role of proteins’ innate conformational flexibility during this process. These results will contribute to our understanding of how the ligands and proteins involved in different cellular processes interact with each other – and, potentially, how these interactions can be manipulated.




