Probing the functional segments, or ‘motifs’, of proteins has helped scientists identify the minimal ingredients needed for them to form biological patterns.
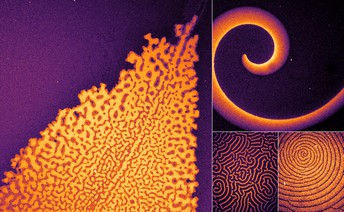
Different patterns formed by the team's minimal biochemical interaction networks. The modular replacements for MinE create this diverse set of patterns when co-reconstituted with MinD on membranes. Credit: Glock et al. (CC BY 4.0)
Writing in the journal eLife, the researchers describe how they dissected the biological phenomenon of protein pattern formation into its main functional modules, and then rebuilt the process from the ground up in a completely new way.
Proteins self-organise to form patterns in living cells, which are essential for key functions such as cell division, communication and movement. A striking example is the MinDE system of the bacterium Escherichia coli (E. coli). This system produces oscillations of two protein types, MinD and MinE, between two poles of the rod-shaped bacteria, positioning the machinery for cell division to midcell. It can be reconstituted in the laboratory, allowing scientists to control and manipulate the functional elements needed for pattern formation via protein mutations.
“Because of its simplicity, the MinDE system has been invaluable in understanding the mechanisms of protein-based pattern formation,” says Philipp Glock, a PhD student at the Max Planck Institute of Biochemistry in Munich, Germany, and co-lead author alongside Fridtjof Brauns and Jacob Halatek, both from the Ludwig Maximilians University of Munich. “A key question that remains is whether this structural and functional complexity can be reduced further to reveal a set of minimal ingredients for pattern formation.”
To answer this, Glock and his colleagues created a minimalistic version of MinE, which plays an antagonistic role in the two-protein MinDE system, by dissecting the protein in a set of core functional motifs, guided by theoretical modelling. One motif, the short helical sequence of amino acids which MinE uses to interact with MinD, is not enough to produce patterns on its own. But adding other functional motifs of MinE one at a time enabled the scientists to fully design new minimal pattern-forming protein mutants.
The team found that at least one other functional motif is required to form patterns. This can either be a motif for membrane binding or a dimerizing motif, which binds to other molecules of the same kind. Neither of these motifs needs to be from native MinE, but can be replaced and potentially simplified further.
Mathematical modelling then allowed the authors to explain why these features are required and how they enable patterns to form. Moreover, they predicted how these patterns adapt to the cell shape in E. coli. The team says that testing these predictions is an exciting goal for future experiments.
“Our work provides a starting point for a modular and tunable experimental platform to design protein-based pattern formation from the bottom-up,” says Petra Schwille, PhD, Director of the Department of Cellular and Molecular Biophysics at the Max Planck Institute of Biochemistry, and co-senior author alongside theoretical physicist Erwin Frey, from the Ludwig Maximilians University of Munich. She adds that while the patterns created by the new system are less regular than those formed by the native MinDE system, they are still sufficient for reproducing and studying basic biological processes.
The model can now be used to study which functional features, regardless of a particular protein system, need to be combined to allow for self-organisation and pattern formation in biology. “Our modular approach may also provide the necessary data for computer modelling of pattern formation in other types of bacteria, as well as more complex organisms,” Schwille concludes.
The above image and videos accompanying this study are available to view and download here.
Media contacts
Emily Packer
eLife
e.packer@elifesciences.org
+441223855373
About
eLife is a non-profit organisation inspired by research funders and led by scientists. Our mission is to help scientists accelerate discovery by operating a platform for research communication that encourages and recognises the most responsible behaviours in science. We publish important research in all areas of the life and biomedical sciences, including the Physics of Living Systems, which is selected and evaluated by working scientists and made freely available online without delay. eLife also invests in innovation through open-source tool development to accelerate research communication and discovery. Our work is guided by the communities we serve. eLife is supported by the Howard Hughes Medical Institute, the Max Planck Society, the Wellcome Trust and the Knut and Alice Wallenberg Foundation. Learn more at https://elifesciences.org/about.
To read the latest Physics of Living Systems research published in eLife, visit https://elifesciences.org/subjects/physics-living-systems.




