Quick question for open science enthusiasts, live from our #eLifeSprint project to increase the impact of scientific Tweets:
Could you connect us with relevant people at research institutes with a strong #openscience agenda and social media presence to get them involved?Michiel van der Vaart (@Michiel_vdVaart)
The eLife Innovation Sprint 2020 took place on September 2 and 3, online. This is a global gathering of researchers, developers, designers, technologists, publishers and others to develop open-source prototypes to redefine the ways we share and do research.
A total of 80 participants worked on 14 projects, from a database collecting information on Article Processing Charges and waivers, to software to track and cite other research artefacts.
You can find out more about these projects and related resources below – we invite your feedback and contributions.
If you are interested in joining the eLife Innovation Sprint in 2021, you can sign up for email updates.
A Badge for FAIR software
The team wants to incentivise research software engineers and researchers who develop software to implement five simple recommendations for making research software more FAIR (Findable, Accessible, Interoperable, Reusable). Their idea was to make use of Shields badges, a common, easy way to indicate the status of a GitHub repository. The badge will visualise the software’s compliance with the five recommendations, thus making the research software developers’ effort and compliance more visible to the rest of the world.
During the Sprint, the team designed the badges and developed a GitHub Action that automatically checks for compliance with the recommendations and suggests a badge according to the check results. The GitHub Action is now available for all to use with their projects on the Marketplace, and you can also visit the project’s GitHub repository.
The team members were:
- Mateusz Kuzak (Project lead)
- Carlos Martinez (Project lead)
- Jurriaan Spaaks (Project lead)
- Edidiong Etuk
- Andrew Holding
- Shyam Saladi
- Chris Shaw
A Clean Slate
Bullying in academia is prevalent but often a very isolating experience. The project leads’ vision is to create a safe space for early-career researchers (ECRs) who experience bullying in academia to share their stories anonymously and support each other. At the Sprint, they started with creating two empathy canvases to collectively gain a better understanding of both the victims of bullying and the bullies. They have also started working on the infrastructure of the platform (a Discourse forum and a website), and put together a quick guide and questionnaire for users.
The team invites everyone interested in contributing to the project to join their Slack community space.
The Discourse Forum prototype.
The team members were:
- Renuka Kudva (Project lead)
- Nafisa Jadavji (Project lead)
- Gabriela Gaona (Project lead)
- Zhang-He Goh
- Bruno Paranhos
- Fatima Pardo Avila
- Helena Perez Valle
- Christine Blume
- Gyan Prakash Mishra
- Parvathy V R
- Michiel van der Vaart
Affordabee: Low-Cost Open Access Publishing
To publish their research open access, researchers are often asked to pay an Article Processing Charge (APC). This is a burden especially for ECRs and students in low and middle-income countries. The Afffordabee team hopes to build a platform that can help students and early-career researchers identify journals that can offer waivers or subsidies, and how to access them, and ultimately to reduce the open-access publishing cost barrier for students from resource-poor settings.
During the Sprint, the team successfully expanded the database of publishers offering APC waivers and subsidies. They also added information on possible APC funding options. Contributors also helped create a website, design a logo and built a process to crowdsource APC waiver and subsidy information. The team hopes to continue to collaborate, and at the same time to recruit more contributors to continue to add useful information to this important resource.

The Affordabee platform. Source.
The team members were:
- Caleb Kibet (Project lead)
- Jennifer Mutisya (Project lead)
- Kennedy Mwangi (Project lead)
- Ryan Davies
- Ana Dorrego-Rivas
- Delwen Franzen
- Arup Ghosh
- Gigi Kenneth
- Elizabeth Oliver
- Subhashish Panigrahi
- Gyan Prakash
ANN: A platform to annotate text with Wikidata IDs
With the ever-growing amount of scientific literature, there’s a pressing need to make more of this literature machine readable. Annotation, the linking of words and concepts with unambiguous external identifiers, is often time-consuming and requires domain-specific knowledge.
At the Sprint, the team brainstormed both technical and practical aspects of developing a tool to crowdsource annotations linking scientific concepts to Wikidata entities. They designed a gamified interface with the aim to incentivise researchers to participate in annotating. Additionally, they studied Natural Language Processing approaches for extracting scientific entities, and assembled a series of perspectives on how to implement such an annotation tool in the current research environment.
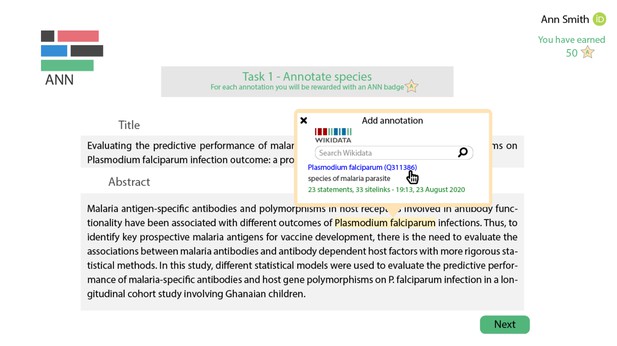

A mock-up of the ANN gamified annotation tool. Source.
The team hopes to continue developing ANN after the Sprint – see the project’s GitHub issues page to explore how you can help.
The team members were:
- Tiago Lubiana (Project lead)
- Dayane Araújo
- Ana Dorrego-Rivas
- Delwen Franzen
- Arup Ghosh
- Inés Hojas
- Bruno Paranhos
- Sabine Weber
- Gabriela Nogueira Viçosa
Code is Science
Code is an important part of modern research, and journals and publishers have established guidelines to help authors make their research code open and reusable by others. Yet, these guidelines are not always followed, negatively impacting the reproducibility and reusability of the published research. The Code is Science team aims to build a central resource to list journal policies regarding code artifacts, but also to explore and evaluate journal code policy compliance, and ultimately, to incentivise publishers towards more open practices.
At the Sprint, the team created user personas for editors, reviewers, code authors and coders to better understand their preferences and needs when interacting with code. They also made crucial front- and back-end infrastructure decisions, and started implementing the platform. Finally, with various contributors at the Sprint, the team brainstormed ways and challenges towards assessing journal code policy compliance. They invite everyone interested in the project and issues to watch the Code is Science project GitHub repository. Please also read project lead Yo Yehudi’s summary of their Sprint work.

Some of the personas built by the Code is Science team. Source.
The team members were:
- João Paulo Taylor Ienczak Zanette (Project lead)
- Isaac Miti (Project lead)
- Yo Yehudi (Project lead)
- Billy Broderick
- Delwen Franzen
- Ryan Davies
- Edidiong Etuk
- Iain Hrynaszkiewicz
- Stuart King
- Dipanshu Nagar
- Festus Nyasimi
- Helena Pérez Valle
- Raniere Silva
- Emmy Tsang
- Abigail Wood
Covidpreprints.com
Preprints and the commentaries around them are important assets, especially in the current pandemic. Covidpreprints.com was started with the aim to enable greater accessibility to and understanding of new research outputs by placing them in the context of the pandemic progression, thus helping readers understand the role of open scientific discourse in solving global problems.
The covidpreprints.com project aimed to find solutions for two questions at the Sprint: (1) how to keep up with the ever-growing literature, and (2) how to best present the information to all users. The team devised a scalable workflow, designed a logo and built a new user interface. In the longer term, they hope to explore the clustering and curation of preprints around themes, and also to make the platform more portable, such that it could be used not only for COVID-19, but other areas of research too.

Covidpreprints.com’s new timeline interface. Source.
Please visit covidpreprints.com to explore this resource, and their GitHub repository to find out how you can contribute.
The team members were:
- Jonathon Coates (Project lead)
- Gautam Dey (Project lead)
- Zhang-He Goh (Project lead)
- Hugo Gruson
- Chris Huggins
- Allan Ochola
- Bruno Paranhos
- Michael Parkin
Crawling data portals
There are many open data portals around the globe that could help address a variety of research questions and issues, but to make the most out of the open data, researchers need a way to find and select the most appropriate portals for their research needs. The crawling data portals team’s goal is to produce open, accessible and easily reusable data for the research community.
At the Sprint, the team first mapped out the front-end, back-end and design tasks that they needed to complete towards a prototype of their vision. They designed a user interface to show interactive benchmarking data of the different portals and created some APIs to allow the community to reuse the data.
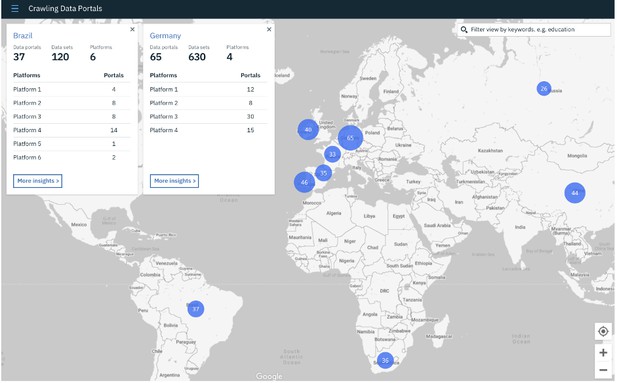
The crawling data portals’ user interface. Source.
They would like to complete the missing APIs and front-end development – please visit their GitHub repository if you would like to contribute to the project.
The team members were:
- Andreiwid Correa (Project lead)
- Kellyton Brito (Project lead)
- Nick Duffield
- Raoni Lourenço
Expanding Open Grants
To have a successful career in academia is not only dependent on one’s research output, but also implicit knowledge around grant proposals, fellowship applications, job statements and so on – the “hidden curriculum” of academic research. Open Grants aims to demystify and democratise research and this hidden curriculum with an open repository of successful and unsuccessful applications and proposals.
At the Sprint, the team set out to improve the usability and functionality of the current platform. From designing and running a user survey, they identified that most users would like to see a keyword search on the platform. In order to facilitate this, the team worked on front and back-end infrastructure and their integration to be able to search both the grant PDF files and metadata.

The Open Grants platform with a keyword search. Source.
Looking forward, the team would like to plan a roadmap towards sustainability and growth. The team invites you to join and help drive forward the project by visiting their GitHub repository.
The team members were:
- Hao Ye (Project lead)
- Esha Datta
- Daniel Nüst
Frictionless DataScriptor
Research papers often contain tables: they exist in different sections of the paper and are used for a variety of purposes. The lack of consistency in how these tables are structured, even when they are used for similar purposes, makes it difficult for machines to understand and extract information from large corpuses of tables.
At the Sprint, the team first extracted and analysed the corpus of tables from eLife papers, to look at what they are used for. They then created a library of article table templates for the main categories of tables – for example, Key Resource Tables, results reporting for differential analyses and so on. In the future, the team hopes to integrate this work with the DataScriptor web application to help researchers automatically generate these tables within their research manuscripts.
The team members were:
- Lilly Winfree (Project lead)
- Philippe Rocca-Serra (Project lead)
- Bradly Alicea
- Billy Broderick
- Jez Cope
- Alf Eaton
- Delwen Franzen
- Danny Garside
- Iain Hrynaszkiewicz
- Chris Shaw
- Gabriela Nogueira Vicosa
Research References Tracking Tool (R2T2)
Research software is not built in isolation – it often codes for an existing theory or uses published datasets. However, there are no existing tools that allow software to cite other research artefacts. The R2T2 team envisioned building a tool to improve citations from software, to give credit to and increase the visibility of all research work that the software is built upon.
During the Sprint, the team worked on a few workflows to allow research software engineers and other researchers to more easily add references to their software, namely workflows that parse Bibtex files, DOIs and docstrings. They also added documentation and examples to help guide users to use their Python package.
Next, they would like to explore outputting the citation information in different formats, such as Citation File Format (CFF) or CodeMeta. They invite everyone who is interested to try the tool.
The team members were:
- Diego Alonso Álvarez (Project lead)
- Jez Cope (Project lead)
- Tino Sulzer (Project lead)
- Billy Broderick
- Daniel Ecer
- Chas Nelson
- Raniere Silva
SciGen.Report
Researchers often try to reproduce or replicate others’ work, and while these results are potentially extremely useful to others in the community, they are rarely published or communicated. SciGen.Report’s vision is to develop and provide a platform for anyone to easily share information on the reproducibility of a publication.
At the Sprint, project lead Cassio worked with designers, editors and others to build a new wireframe for the platform. In addition to working on improving the design of the workflow and user interface, the contributors also engaged in some deeper discussions around the problem that SciGen.Report is trying to solve, and what a “reproducibility score” could look like.
The goal of the tool is ultimately to drive a “third branch” of research communication beyond preprints and publications, to bring visibility and recognition to post-publication reviews. You can find out more about Cassio’s vision in the ORION Open Science Podcast’s coverage of the Sprint.
The team members were:
- Cassio Amorim (Project lead)
- Ryan Davies
- Giuliano Maciocci
- Veethika Mishra
- Helena Perez Valle
- Emmy Tsang
Tweet your research suggestions
An increasing number of researchers are using social media to share their work and increase their visibility, one of the most popular platforms being Twitter. This creates an opportunity to explore whether Twitter can be used to spark discussions, create collaborations and motivate reuse of published research, if tweets can be better formulated to highlight research insights for specific researcher audiences.
The team identified four components that should be included in a tweet: who it is for, why they should care, what this means in practice and a call to action. They tested the prototype by formulating tweets according to these steps to ask for help during the Sprint, and successfully received replies to their question. They have also devised a workflow to help researchers easily implement these rules in their research article tweets day-to-day.
Moving forward, the team has engaged with eLife to design and carry out a pilot study to critically assess the impact of tweeting research with these guidelines.
The team members were:
- Jan Paul Grollé (Project lead)
- Michiel van der Vaart (Project lead)
- Ryan Davies
- Inés Hojas
- Dasapta Erwin Irawan
- Danny Garside
- Stuart King
- Giuliano Maciocci
- Gavin McStay
- Veethika Mishra
- Daniel Nüst
- Helena Pérez Valle
- Micah Vandegrift
The Software Citation Project
Software is an integral part for most modern-day research, yet software citation is still an issue: software is often not cited correctly. The Software Citation Project builds on work in this area carried out by many, including the team at the eLife Innovation Sprint 2019, with the aim to enable research software engineers (RSEs) to add citability to open-source software without hassle. One of the ways to achieve this is to build a tool to allow RSEs to easily add a Citation File Format (CFF) file to their software release.
Through running a survey, the team realised that while the awareness and adoption of CFF files are quite low, there is a demand for a tool that is interoperable with the existing software publishing and archiving workflow and which can help create a CFF file at the same time. During the Sprint, the team also improved both the front and back ends of the existing prototype.
Moving forward, they hope to continue to work on supporting conversion of CFF files to other citation formats, as well as providing more information on the website on software citation.
The team members were:
- Sarthak Sehgal (Project Lead)
- Naomi Aro
- Nick Duffield
- Rohit Goswami
- Chas Nelson
#
We welcome comments, questions and feedback. Please annotate publicly on the article or contact us at innovation [at] elifesciences [dot] org.
Do you have an idea or innovation to share? Send a short outline for a Labs blogpost to innovation [at] elifesciences [dot] org.
For the latest in innovation, eLife Labs and new open-source tools, sign up for our technology and innovation newsletter. You can also follow @eLifeInnovation on Twitter.




