Blogpost by Naomi Penfold, Innovation Officer
Today, we announced a new project collaboration between Substance, Stencila and eLife, to support the development of an open technology stack that will enable researchers to publish reproducible manuscripts through online journals. In this blogpost, we outline the background and remit of this project. We welcome feedback and contributions: please comment publicly on the article using Hypothes.is or email innovation@elifesciences.org.
Building on the traditional research article
The research article: a concise narrative describing the background, methods, results and interpretation of a body of research work. And while the established format of research articles generally allows researchers to describe their work in sufficient detail to allow others to repeat, understand, critique and validate it, that same format falls short when it comes to communicating the complexity of modern research.
For example, an early finding from the Reproducibility Project: Cancer Biology (RPCB) is that research methods are not generally described in sufficient detail to allow other researchers to faithfully repeat key experiments without engaging the original authors. And although engaging the author is an important part of the scientific conversation, the findings that form the very building blocks of scientific progress should ideally be communicated in a way that outlasts the original author’s memory.
To this end, researchers today have access to a multitude of dedicated tools and services aimed at facilitating the sharing of specific methods and resources: Bio-protocol (a journal), Protocols.io (an online repository and forum), and resource-specific initiatives such as AddGene and Chemical Probes). Additionally, data and code repositories, and dedicated journals such as Scientific Data, JOSS and JORS provide platforms to more completely describe research data and software. Yet, while the abovementioned initiatives help researchers to improve the documentation of all the assets that lie behind a research finding, they also share one key obstacle to reproducibility: they are not integral to the research article itself.
These days, data trails are often a morass of separate data and results and code files in which no one knows which results were derived from which raw data using which code files.
— Professor Charles Randy Gallistel, Rutgers University
Supporting a reproducible document format
A reproducible document is one where any code used by the original researcher to compute an output (a graph, equation, table or statistical result) is embedded within the narrative describing the work.
Reproducible document formats like R Markdown and Jupyter Notebooks help researchers organise both data and code into an understandable workflow that not only improves documentation for the authors themselves, making it faster to build on previous work when new data becomes available, but which also aids collaboration, training and reproducibility.
Tools like Jupyter notebook enable a level of interaction that can transform scientific communication. Exchange between authors and reviewers and, ultimately, between authors and readers could be very focused on technical aspects. This would counteract a trend by which papers are seen as tools to advertise research work, rather than to enable other investigators to proceed with research.
— Professor Vincenzo Carnevale, Temple University
But when submitting to a journal, authors today have to export their manuscripts in a flattened format (Word doc, PDF) or submit in LaTeX. This process strips code from being embedded into the narrative of the research into a separate asset. What started as a self-contained, fully reproducible document is transformed into a collection of linked assets.
We are already composing our research manuscripts in a format that is intimately paired with the methodology and data underlying the findings. It's frustrating to have to convert these rich and reproducible documents into flattened formats, such as PDF and Word documents, in order to satisfy the needs of the traditional publishing workflow. I would like to see publishers supporting me to share my work and data in the reproducible formats I prepare them in.
— Dr Laurent Gatto, University of Cambridge
Earlier this year, researchers familiar with reproducible document tools told us they were interested in being able to share and read research articles with features that support better code and data sharing as well as greater interactivity and executability:
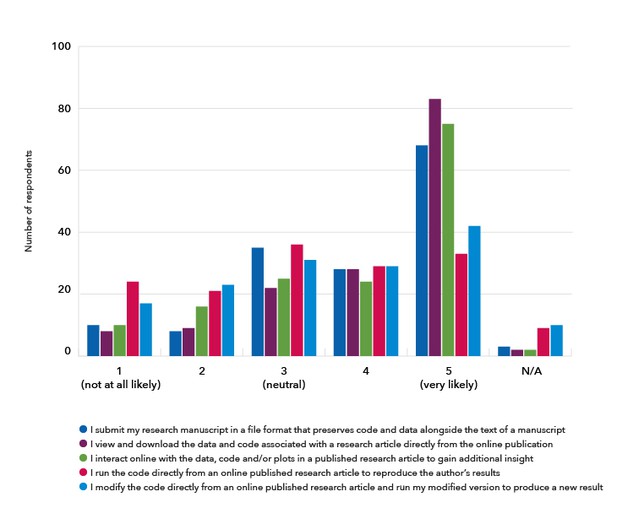
Source: https://elifesciences.org/inside-elife/e832444e/innovation-understanding-the-demand-for-reproducible-research-articles.
As Dr Ithai Rabinowitch at the Hebrew University of Jerusalem, notes, embedding data and code in an organised manner would not only aid reproducibility efforts, it could also “make science much more efficient as a community endeavour”. Professor Gary Bader, University of Toronto, adds:
Having the code available for analysis workflows would really speed up research. For instance, one could copy and improve a workflow for use with another data set. We've seen this in my lab with the use of R/Jupyter notebook.
Finally, providing a means for authors to enrich their research article in this way would recognise the researchers who invest resources in preparing these assets. This is a key point given the imbalance between effort and reward has been identified as a key barrier to researchers adopting more open research practises.
The Reproducible Document Stack Project
We wish to support reproducible document formats from authoring through publication. A valuable step forward would be to enable researchers to publish the code and data behind their analyses in a form that can be easily examined, and thereby create figures and whole documents where users can faithfully reproduce key aspects of the work.
As such, we are embarking on a project with Substance and Stencila to develop the technology required to support the Reproducible Document through authoring, sharing and publication.
We will have succeeded once any scientist can easily create a standardised reproducible publication and submit it to a journal. Together with our supporters we are building an open source toolset, to be used by authors, journals and readers.
— Michael Aufreiter, Substance
The creation of an open standard for the exchange, submission and publication of reproducible documents is critical for widespread adoption by academic publishers, and will be beneficial for the discovery and persistence of research reported in this form. Therefore, a key output of this project will be a Reproducible Document Exchange Format, which will allow the data, code and computed outputs (graphs, statistical results, tables) embedded in a reproducible document to be recognised and presented online as an enhanced version of the published research article. In order to do this, Substance is investigating how to represent these assets in JATS XML, the publishing standard through which research manuscripts are processed through the publishing workflow:
<fig id="f1">
<caption>
<title>Figure 1</title>
<p>Biodiversity on Mars</p>
</caption>
<alternatives>
<code executable="yes" specific-use="input" language="mini">
bars(counts_by_species)
</code>
<code specific-use="output" language="json">
{
"execution_time": 322,
"value_type": "plot-ly",
"value": {...}
}
</code>
<!-- static version for existing JATS toolchains -->
<graphic xlink:href="89f8b53e361f.svg"/>
</alternatives>
</fig>This complements Substance's work to develop Texture, an XML-based text editor for authoring, and contributions towards eLife Lens, the side-by-side article reader.
We recognise that these computational research tools are not the mainstay of all life sciences research. Efforts to encourage greater adoption of reproducible research methods by researchers who are less familiar with programming are welcomed to aid interdisciplinary communications, and facilitate those who wish to learn code-based research practises. Substance and eLife are already supporting the development of Stencila, an authoring platform and execution engine for reproducible documents that is targeted at researchers who are less comfortable with programming and more familiar with Microsoft Word and Excel.
The calls for research to be reproducible are growing louder. But many of the tools for reproducible research are code focussed and can be intimidating to non-coders. We’re creating tools with the same intuitive, visual interfaces that researchers are familiar with but built from the ground up with reproducibility in mind. We want to create a platform that is accessible to a greater range of scientists - without dumbing down their research or restricting their ability to learn new computational methods.
— Nokome Bentley, Stencila
With intuitive text and spreadsheet authoring interfaces capable of producing XML documents, Stencila offers a platform to feed directly into the reproducible document publishing workflow.
In addition, we are interested in developing features that will enable readers to make the most of this enhanced article. Above all, the task of replicating an author’s original research – at least from a computational and analysis angle – should be trivial from the reader’s standpoint.
To meet the needs of researchers, the project will need to address several key technological considerations, including:
- The diversity of programming environments used in research must be supported – a variety of languages and environments are used to perform computations in research, from Python and R, to Bash and Matlab, and sometimes multiple languages are combined in a single workflow. The Reproducible Document Exchange Format will be designed with applications like Jupyter and R Studio (a popular R Markdown authoring tool) in mind, but ultimately be platform-agnostic to enable researchers freedom of choice in their own workflow.
- Researchers should be able to follow their current workflows – the rich ecosystem of platforms and tools for conducting, storing, documenting and sharing computational research is a resource that we intend to build on. Further, some scripts and data are too complex to sit within a reproducible document. Where possible, we aim to support as many use cases and workflows as possible, and we welcome input from the community to help us to understand these use cases (email innovation@elifesciences.org).
- Composing reproducible manuscripts should be easy – current authoring tools such as Microsoft Word are popular for a reason: What You See Is What You Get (WYSIWYG). We intend to support tools that make authoring and consumption of computationally reproducible documents as intuitive as possible for the user. It was with this principle in mind that we recently redesigned our online journal.
Get involved
We are at the beginning of the process to develop a complete publishing solution to support the Reproducible Document envisioned above. We expect this process to take at least 12 months.
We are open to your input and feedback throughout, whether about the concept or regarding specific elements. Please consider commenting publicly on this blogpost using Hypothes.is, or emailing us directly at innovation@elifesciences.org. For more news, please sign up for our technology and innovation newsletter.
For specific elements contributing towards this project, please see:
- Texture is an open format manuscript editor: the code is open source at https://github.com/substance/texture
- Stencila is a office suite for reproducible research: it is being developed openly at https://github.com/stencila/stencila and you can participate in discussions at https://community.stenci.la/
Future considerations
The ultimate goal of this project is to enable researchers to share their research manuscripts in a reproducible format. This will complement existing practises to share research data and code, and readers will remain free to use the assets as they wish. However, this project will also open up new avenues for innovation, including the executability of articles online, and the potential to capitalise on reproducible resources during peer review.
The idea of supporting executable notebooks is not new. In 2014, Nature piloted a feature that supported an online interactive iPython notebook (now Jupyter notebook). This was a popular feature, likely due to the novelty, with more than 20,000 notebook runs computed. The team behind Authorea, the online collaborative authoring platform, continue to add features that enable authors to include dynamic and interactive plots in their manuscripts and to highlight the source data underlying a figure. How these features could be supported by the ultimate publisher of the peer-reviewed work remains to be addressed, but F1000 Research has piloted similar features and continues to experiment along these lines. Further, journals in other domains are responding to the needs of their own computational research communities: Distill, an online journal for machine learning research, includes a ‘reactive diagram’ feature and, in some articles, readers can even input data to trial an algorithm.
Any third-party review of data and code assets is challenging. Currently, reviewers have to in many cases take the authors at their word that the code and algorithms they employed in their work are robust. At Nature Neuroscience, the efficacy of code sharing is currently under investigation via a pilot to review the code used to compute key figures in new articles. We hope that offering a means to share reproducible documents will help to incentivise the sharing of demonstrably reusable data and code underlying a research article. Further, opening up the means to support the submission of research in a reproducible format could assist interested reviewers to reproduce computations with as minimal extra effort as is feasible.
For the latest in innovation, eLife Labs and new open-source tools, sign up for our technology and innovation newsletter. You can also follow @eLifeInnovation on Twitter. We welcome comments and questions, so please comment using Hypothes.is.




