By Emily Schlafly1, Anthea Cheung2, Samantha W Michalka3, Paul A. Lipton4, Caroline Moore-Kochlacs1, Jason Bohland5, Uri T. Eden2,6, Mark A. Kramer2,6
1. Graduate Program in Neuroscience, Boston University, United States
2. Department of Mathematics and Statistics, Boston University, United States
3. Olin College of Engineering, United States
4. Princeton School of Public and International Affairs, Princeton University, United States
5. Department of Communication Science and Disorders, University of Pittsburgh, United States
6. Center for Systems Neuroscience, Boston University, United States
Sophisticated data collection requires sophisticated data analysis: skills in computer programming, statistics, and mathematics are now required of neuroscientists [1], [2]. However, most experimental and clinical neuroscientists lack sufficient training in these skills, and the idea of tackling a standard technical course or textbook on data analysis without sufficient, relevant tutorials, examples and hands-on practice is daunting. Therefore, a fundamental gap has emerged in neuroscience labs between highly sophisticated and rich data collection procedures, and limited training to assess and understand these data with state-of-the-art quantitative techniques. While important for training the next generation of neuroscientists, computational neuroscience education remains a challenge [1], compounded by recent requirements for remote learning and virtual classrooms [3].
The data, analysis methods and code are not toy examples, but instead correspond directly with modern techniques in use today.
To help address these challenges, we present here an online, freely available resource to enhance training in neural data analysis. To reach the practicing neuroscientist, we assume only a basic knowledge of calculus and statistics, common to those trained in biological sciences [1]. We also invert the standard presentation of dta analysis techniques. Typical statistics textbooks in data analysis begin with the development of theory and then describe applications. We instead begin each topic with an example case study of neural data (e.g., action potentials from rat hippocampus, or scalp electroencephalogram data recorded from a human subject). These data then motivate the development and application of modern analysis techniques (e.g., visualization approaches, spectral analysis, bootstrapping, and generalized linear models). We emphasize a hands-on approach, example data sets are provided, and computer (Python) code interspersed within the material encourages direct interaction with the concepts. The data, analysis methods and code are not toy examples, but instead correspond directly with modern techniques in use today. Upon completing each case study, the learner will be able to immediately deploy these data analysis tools in his or her own research.
Features
All notebooks have consistent structure. This structure consists of:
- A brief synopsis of the notebook data and analysis methods.
- An on-ramp to immediately introduce and evaluate in Python a core data analysis concept.
- A description of the data, method of collection, and scientific question motivating analysis.
- A detailed statement of the module goals and the data analysis tools to be studied.
- Data analysis begins with visualization approaches.
- Data analysis continues with implementation of the techniques specific to the notebook, including development of appropriate theoretical material and practical implementations.
- Data analysis concludes with rendering of results.
Content focuses on practical data analysis applications. The outline from one notebook is shown in Figure 1. In this case study, the research question asks whether a stimulus evokes a response in the EEG. To answer this question, we show how to compute the event-related potential (ERP), generate confidence intervals using a bootstrapping technique, and establish the statistical significance of the evoked effect.
Other examples of case studies include:
- Identifying rhythms in EEG data using common spectral analysis tools like the Fourier transform, power spectral density, and spectrograms.
- Quantifying coupling between two brain regions in invasive EEG recordings using cross-covariance and coherence.
- Characterizing the properties of a neuron recorded in rodent hippocampus using point process theory and generalized linear models.
A full list of goals, available datasets, and analytical tools is given in Table 1.
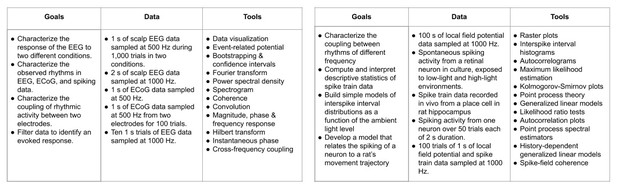
Table 1. Research goals, data, and tools discussed in Case Studies.

Figure 1. Example content outline from a notebook.
Notebooks are modular and can be completed in any order. Although we organize the notebooks sequentially, each notebook is independent of any other. In this way, learners and educators may develop their own individual learning paths through the material. For example, a learner with limited programming experience who is interested in the fundamentals of EEG analysis could begin with the extensive Python tutorial (Notebook #1), and then proceed to notebooks introducing the evoked response potential (Notebook #2) and spectral analysis (Notebooks #3 and #4). Or, an educator teaching a graduate course in computational neuroscience may introduce the topic of cross-frequency coupling (CFC, an active research area [4]) and provide students with a hands-on example of implementing and applying a measure of CFC (Notebook #7). A motivated individual could also complete the entire sequence of notebooks.
GitHub, Python, and Jupyter notebooks for an interactive online learning environment. To facilitate an open, freely available distribution of the material, we use the programming language Python, host all material on GitHub, and organize the material into Jupyter notebooks [5], each focused on analysis of a specific data set. To interact with the material, a learner may download (or clone) the entire repository (including all data, Python code, and written material) for use on his or her own device. Alternatively, a learner may interact directly with the material through his or her web browser; no downloads or software installations required. Using Binder or ThebeLab, the Jupyter notebooks become fully interactive environments; the data analysis (Python code) becomes executable and modifiable directly in most modern web browsers.
Use cases for learners and educators
The material was developed for investigators of any ability and at all career levels – from the beginning undergraduate researcher to the established PI – to analyze and understand neural data. For those new to Python, Notebook #1 is dedicated to a thorough but rapid and practical introduction to the use of Python 3. We anticipate many scenarios for participation, for example:
- A clinical research lab requires new data analysis skills in scalp EEG. Lab members complete notebooks describing analysis of brain rhythms and rhythmic coupling of field data (Notebooks #3-5).
- A postdoctoral researcher requires modern spike-field coherence analysis techniques as part of her ongoing research. An expert in Python, she completes a single notebook (Notebook #11).
- An instructor plans to update his undergraduate laboratory course in basic neuroscience. In the lab, students record spike train data. The instructor augments this lab by requiring students complete a notebook introducing spike train analysis (Notebook #8) and apply basic visualizations to the spike train data recorded in the lab.
- An instructor must suddenly teach her neuroscience course remotely. She selectively chooses content from a subset of notebooks to aggregate into a coherent curriculum. Students complete notebooks outside of class, and discuss material during (virtual) classroom time [3].
We expect interested researchers, students, and educators in the neuroscience community will develop many other uses beyond the examples listed here.
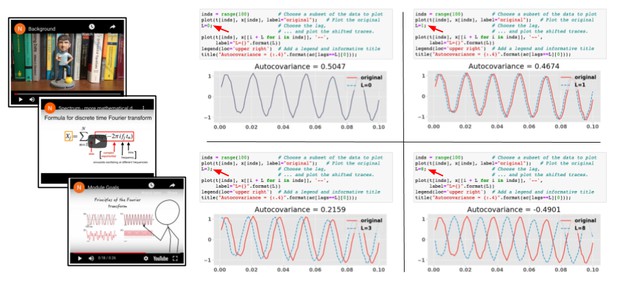
Figure 2. Notebooks introduce concepts with short videos and interactively explore mathematical ideas. In this example, the learner explores how the autocovariance is related to the lag between signals. Changing the parameter L (red arrows) within the executable notebook supports interactive learning, visualization, and exploration.
Notebooks are dynamic and available for further development by the community. The collaborative nature of GitHub allows for dynamic evolution of the material. A modification (e.g., an error correction) suggested by any learner can be flagged as an issue and incorporated into the material; versioning allows a complete record of attributable changes. Moreover, the repository is expandable. For example, a tutorial that implements and applies a new data analysis method to an example data set may be contributed as a standalone notebook. In this way, the community’s expertise allows the material to grow and include important topics not yet presented (e.g., analysis of imaging data).
An accessible data analysis resource for practicing neuroscientists. We present here an online platform for training in neural data analysis. The material is modular, motivated by real-world examples in neural data analysis, and executable directly in most modern web browsers. We invite the neuroscience community to explore the resource and contribute to improving, expanding and updating its material, methods and datasets, to make it accessible and useful to more in the community.
Acknowledgements
The authors acknowledge support from NIH NIGMS R25GM114827 and NSF DMS #1451384 (MAK). The authors thank MIT Press for publishing a complementary version of this material.
References
[1] M. S. Goldman and M. S. Fee, “Computational training for the next generation of neuroscientists.,” Curr Opin Neurobiol, vol. 46, pp. 25–30, Jul. 2017, doi: 10.1016/j.conb.2017.06.007.
[2] M. A. Kramer and U. T. Eden, Case Studies in Neural Data Analysis. Cambridge, MA: MIT Press, 2016.
[3] S. Sandrone and L. D. Schneider, “Active and Distance Learning in Neuroscience Education,” Neuron, vol. 106, no. 6, pp. 895–898, Jun. 2020, doi: 10.1016/j.neuron.2020.06.001.
[4] A. Hyafil, A.-L. Giraud, L. Fontolan, and B. Gutkin, “Neural Cross-Frequency Coupling: Connecting Architectures, Mechanisms, and Functions.,” vol. 38, no. 11, pp. 725–740, Nov. 2015, doi: 10.1016/j.tins.2015.09.001.
[5] “Introducing Binder 2.0 — share your interactive research environment,” eLife, Nov. 30, 2017. https://elifesciences.org/labs/8653a61d/introducing-binder-2-0-share-your-interactive-research-environment (accessed Jun. 19, 2020).




