Rigid firing sequences undermine spatial memory codes in a neurodegenerative mouse model
Figures
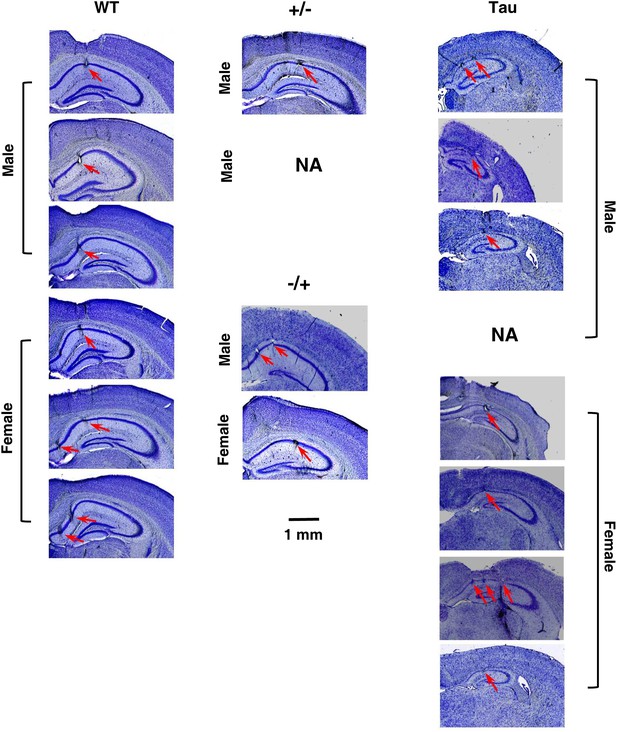
Histological confirmation of recording sites and neurodegeneration in Tau mice.
The images are cresyl violet stained coronal brain sections, showing (1) the recording sites at the CA1 pyramidal cell layer of HP and (2) the prominent cell loss in Tau mice, compared with WT mice and mice with other two control genotypes (+/− and −/+). One section per recorded mouse is shown. One +/− mouse was not available for sectioning (NA). The recorded side of the brain in one Tau mouse was damaged during sectioning (NA). Recording sites at CA1 in these two mice were identified by electrode depths and the presence of sharp-wave ripple complexes (Csicsvari et al., 2000). Scale bar applies to all sections. Arrows: recording sites. Note the smaller hippocampi and thinner CA1 pyramidal cell layers in Tau mice.
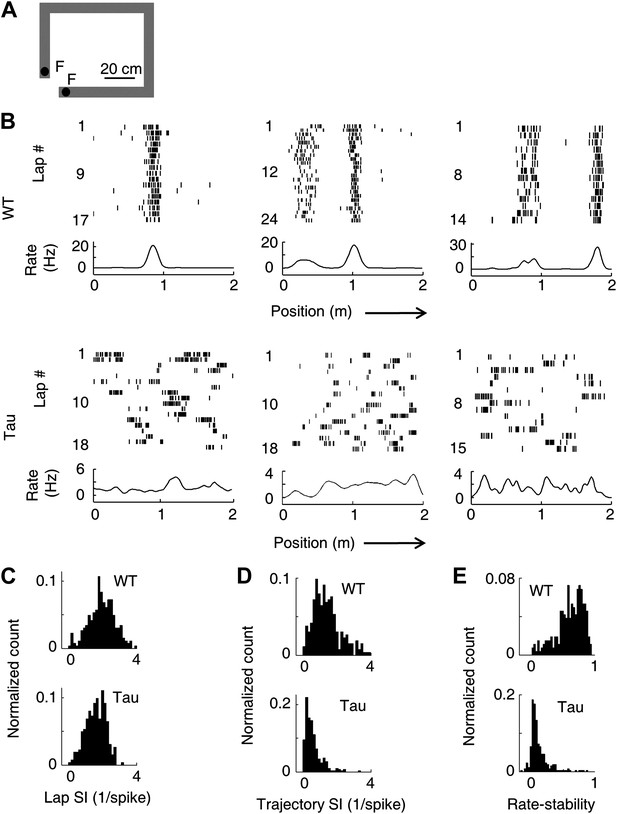
Unstable firing locations of Tau neurons led to the loss of overall location-specificity on familiar trajectories.
(A) The rectangular familiar track. F: food wells. (B) Lap-by-lap spike raster of three WT and three Tau neurons, each from a different animal, on one trajectory of the track. The trajectories were linearized and plotted as the x-axis (arrows: running directions). Each tick represents a spike. Bottom curves: firing rates on the trajectories averaged across all laps. Note the unstable firing locations of the Tau neurons. (C)–(E) Distribution of lap SI (C), trajectory SI (D), and rate-stability (E) of WT and Tau neurons (see text for definitions). Plots are histograms normalized by total numbers of samples, each computed for one neuron on one trajectory.
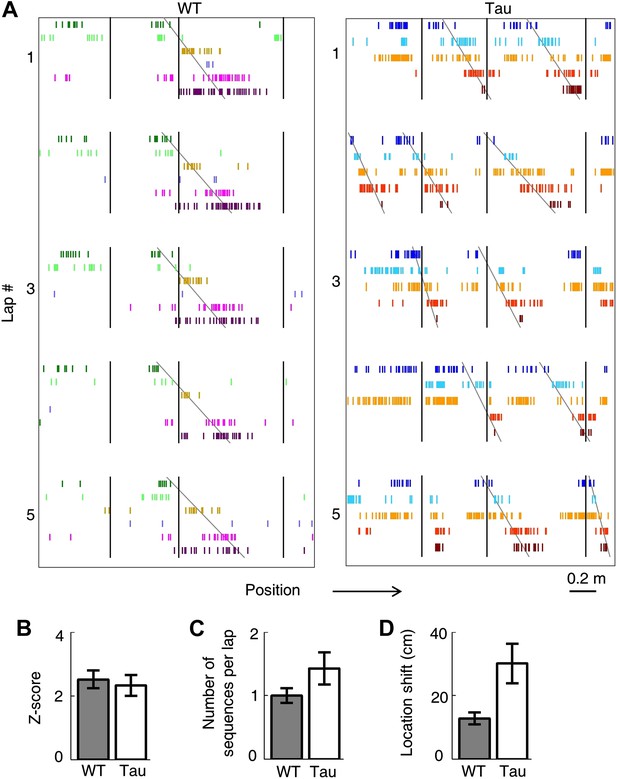
Tau neurons maintained stable firing sequences on familiar trajectories.
(A) Lap-by-lap spike raster of six WT neurons and that of five Tau neurons on a trajectory (arrow: running direction). Each color-coded row within a lap represents a neuron. Each tick represents a spike. Only five laps are shown. Angled lines: detected firing sequences that matched with the templates derived on the trajectories (see Figure 3—figure supplements 1 and 2 for details of template generation and sequence detection). Vertical lines: landmark positions (three corners of the track in Figure 2A) along the trajectory. Note that the Tau neurons shifted their firing locations, but maintained the same firing sequences across the laps. See more data from more animals with more laps in Figure 3—figure supplement 3. (B) Mean number of detected sequences (in Z-score) on the track trajectories in WT and Tau mice. The robustness of this sequence detection result is examined in Figure 3—figure supplement 4. (C and D) Mean number of sequences per lap (C) and mean location shift (D) for the detected sequences in WT and Tau mice.
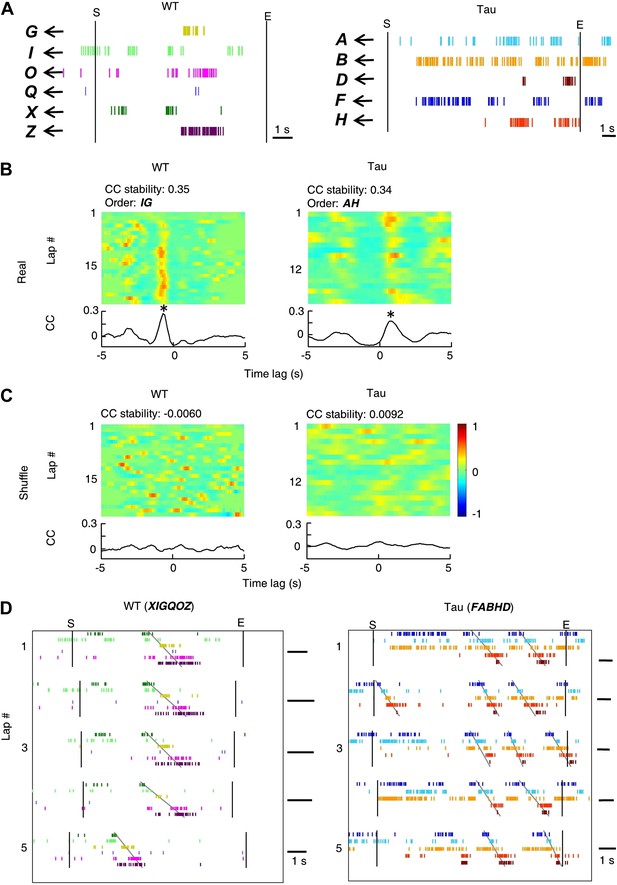
Deriving template sequences based on pair-wise cross-correlations.
(A) First, we assigned letter identities (bold letters) to cells active on a trajectory. Spike raster of six WT and five Tau cells (same as those in Figure 3A) during a single running lap are plotted in time. Vertical lines: start and end times of the laps. (B and C) Second, we determined whether pairs of cells had stable firing relationships based on the stability of their cross-correlations (CCs). For examples, the CCs of a pair of WT (cell G and cell I in A) and a pair of Tau cells (cell A and cell H in A) are shown in (B). Each panel shows the color-coded CC for every lap and the bottom curve shows the lap-averaged CC. The color bar is shared by all panels in (B) and (C). For every lap, a CC was generated by cross-correlating the two cells' binned firing rates (bin size: 100 ms) in the lap. Note the stable peak locations of the lap CCs and the prominent peak of the lap-averaged CC for both the WT and Tau pairs in (B). For each cell pair, we computed a CC-stability, defined as the average correlation-coefficient between any two laps' CCs. To evaluate the significance of the CC-stability, we also computed a shuffled version of CCs (C) after the two cells' firing rates were circularly slid with independent, random time intervals (slide-shuffling). Note the inconsistent peaks in the lap CCs of both the WT and Tau pairs and consequently small CC-stability values in (C). For each cell pair, we generated 1000 shuffled CCs by slide-shuffling and therefore obtained 1000 chance level CC-stability values. A cell pair was considered to have a stable CC if (1) CC-stability is greater than the upper 1% of the chance level (p<0.01) and (2) there was a peak in the lap-averaged CC (the maximum value of the Z-transformed CC ≥ 1). Third, we ordered the cells in those pairs with a stable CC. For the pairs in (B), because the peak of the lap-averaged CC between cell G and I occurred at a negative time lag (*), we ordered the pair as IG. Because the peak between cell A and H occurred at a positive time lag (*), we ordered the pair as AH. If the peak of a pair occurred at exactly time lag 0, we randomly ordered the cells in the pair. (D) Finally, we constructed template sequences that agreed with all the ordered pairs (see ‘Materials and methods’). The derived template sequences for the WT (XIGQOZ) and Tau (FABHD) cells are shown. Spike raster of the cells, ordered according to the templates, are displayed in the time domain during the same five laps as in Figure 3A. Note the repeated occurrence of the template sequences across laps. This also shows that firing sequences of WT and Tau cells in the time domain were very similar to that in the space domain (Figure 3A).
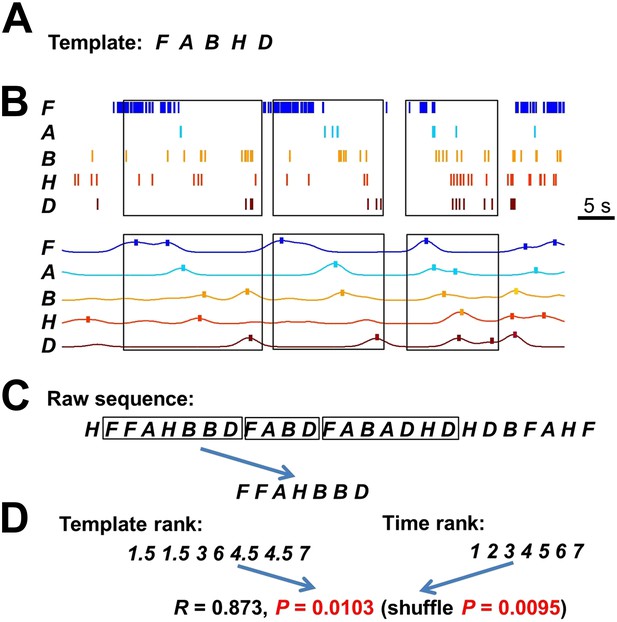
Detecting matches with a template sequence.
(A) An example template sequence (FABHD) made of five cells (A, B, D, F, H). (B) The spike pattern of the five cells in a time period. Here we show how the spike pattern was searched through to find matches with the template sequence. First, the spike trains of the five cells (top) were transformed into instantaneous firing rate curves (bottom) using a Gaussian kernel (σ = 1 s) (Ji and Wilson, 2007). The local maxima of the temporal rate curves greater than a threshold (here was the mean rate of a cell in the entire running session) were detected (ticks). Boxes: detected matches. (C) Second, the detected local maxima of all the cells were ordered into a raw sequence according to their times. Now the spike pattern was translated into a long raw sequence. Boxes: segments of the raw sequence that matched with the template sequence, as determined below. (D) Third, we searched through the raw sequence to find the segments that match with the template sequence. A match was defined by a significant rank order correlation. For example, given a segment FFAHBBD, we determined a rank for each letter according to its order in the template (template rank) and a rank according to its time (time rank). The correlation coefficient of the two ranks was computed and its significance was assessed by using 1000 random shuffles of the letters in the segment (shuffle p). In pilot computations, we found that the p value determined from Pearson's R (p=0.0103 for this example) was a close approximation of shuffle p (0.0095 for this example). To save computing time, we later used a threshold (p<0.05) on the Pearson's p value to define matches.

More examples of WT and Tau firing sequences on familiar trajectories.
The first 15 laps are shown. Each panel (A, B, or C) was from a separate WT or Tau mouse, which was different from the one in Figure 3A.
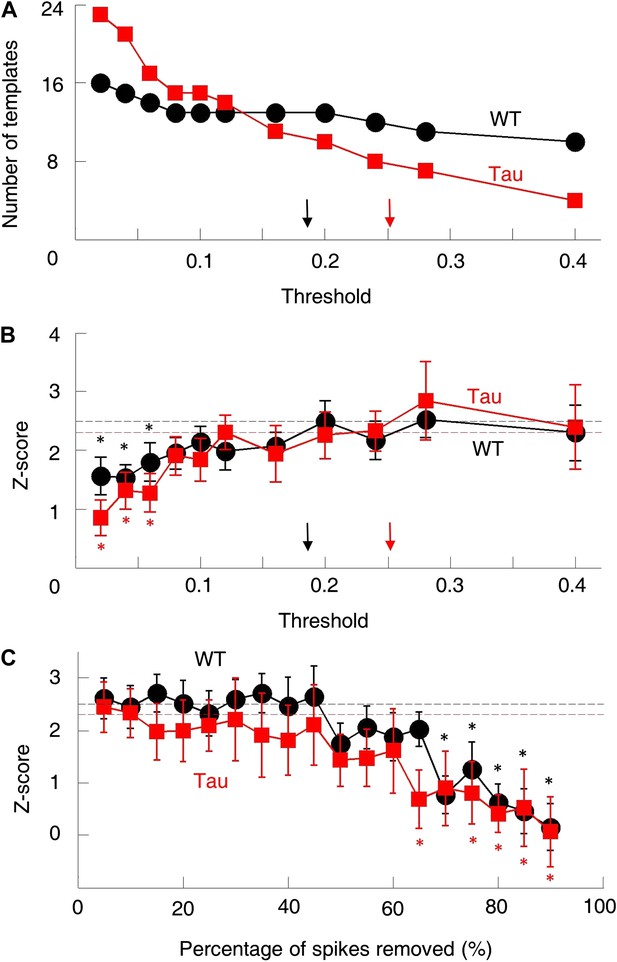
The robustness of the sequence detection result.
We examined the robustness of our sequence detection result to answer two questions. (A and B) First, the template sequences were generated by using a threshold to select pairs of cells with consistently high correlations. How did the sequence detection result depend on the particular threshold chosen? The number of templates (A) and mean number of detected sequences (in Z-score) (B) on familiar trajectories in WT (black) and Tau (red) mice were computed using a wide range of cell selection threshold (0.05–0.4). Arrows: actual thresholds (black: WT, red: Tau). Different actual thresholds were used for WT and Tau mice because the thresholds were computed from chance-level correlations (Figure 3—figure supplement 1), which differed between WT and Tau cell pairs. Dashed horizontal lines: actual Z-scores at the actual thresholds. *p<0.05 from the actual Z-scores. The Z-scores changed very slowly as the threshold was decreased. Only when the threshold became extremely low (<0.08), did the Z-scores drop significantly. (C) Second, Tau cells fired more broadly on the track, which might have increased temporal overlap in their firing patterns and yield more opportunities for sequence detection. Were the Z-scores sensitive to the possible overlapped firing of Tau cells? To address this, we systematically down-sampled the spike trains during trajectory running and re-computed the Z-scores. We binned spikes of each cell into 1-s long bins and removed spikes within randomly chosen bins. This manipulation reduced the temporal overlap among cells at a time scale of our sequence detection. The result is plotted similarly as in (B). It is clear that the actual Z-scores were insensitive to the down-sampling. Only when more than 60% spikes removed, did Z-scores start to drop significantly below the actual values. Therefore, although the absolute number of detected sequences on the track inevitably depended on the exact parameters chosen, the normalization (Z-scoring relative to the chance level) in our sequence detection method resulted in a robust quantification of the sequential structure in the multicell spiking patterns.
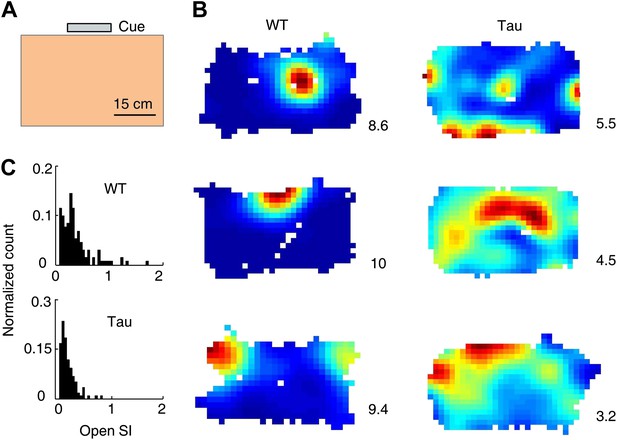
Tau neurons fired with low location-specificity in a familiar open box.
(A) The open box with its interior color and cue card (Cue) shown. (B) Color-coded firing rate maps of three WT and three Tau neurons, each from a different animal, in the open box. Numbers: peak (red/black) rates in Hertz. Note the broader firing areas of Tau neurons than those of WT neurons. (C) Distribution of open SI of WT and Tau neurons. Plots are histograms normalized by total numbers of samples, each computed for one neuron in one open box session.
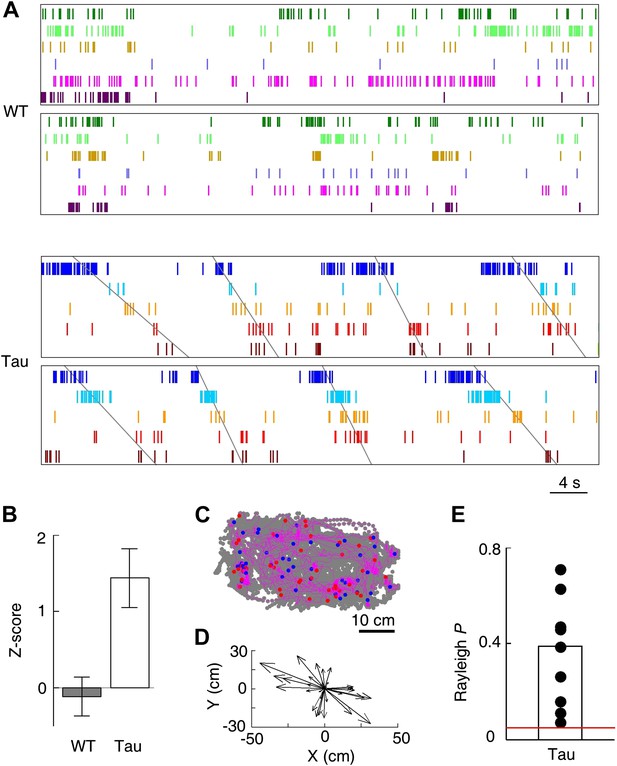
Tau, but not WT, sequences seen on the familiar trajectories also appeared in the familiar open box.
(A) Spike patterns of the same WT and Tau neurons as in Figure 3A within two time periods of an open box session. Neurons are color-coded. Each tick represents a spike. Angled lines: sequences detected as matches with the Tau sequences shown in Figure 3A. See more data from more animals in Figure 5—figure supplement 1. (B) Mean number (in Z-score) of detected WT and Tau sequences in open box sessions. (C) Running paths of a Tau mouse for all the detected sequences in one open box session. For every sequence, the animal’s position at its start/end time was marked by a blue/red dot and its running path in between by a purple line. Gray dots: positions of the animal during the entire session. Note that the sequences occurred everywhere in the box. (D) Vector representation of the paths in panel (C). For each path, a vector was drawn from its start to end location and was plotted with the start location translated to the origin. Note that there was no preferred direction for the vectors. (E) Rayleigh p values for individual searches in open box sessions of Tau mice. Each dot represents a search for one trajectory template sequence in one open box session. Bar: mean values. Red line: p=0.05.
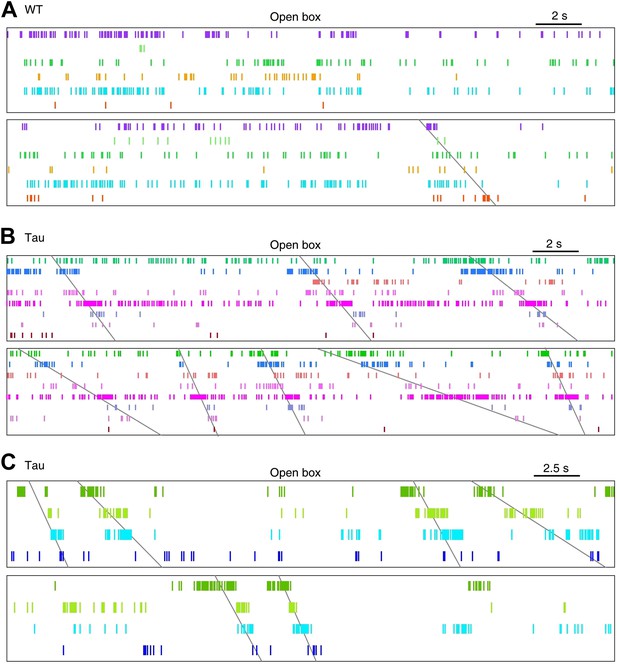

More examples of WT and Tau firing patterns in the familiar open box.
Neurons in each panel (A, B, or C) were the same ones plotted in the corresponding panel (A, B, or C) of Figure 3—figure supplement 3.
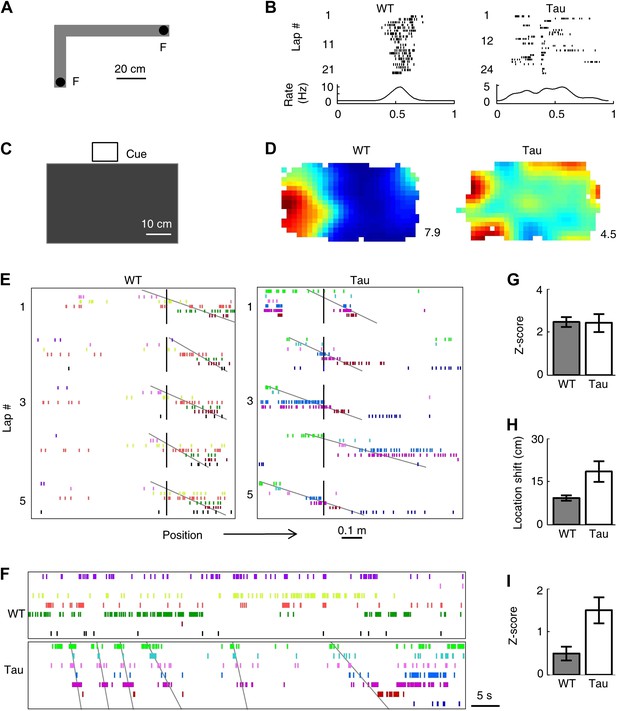
Tau neurons formed robust firing sequences on a novel track and in a novel open box, despite their low location-specificity.
(A) The L-shaped novel track. F: food wells. (B) Lap-by-lap spike raster of a WT and a Tau neuron on one trajectory of the track (see Figure 2 for details). (C) The novel open box with its interior color and cue card (Cue) shown. (D) Firing rate maps of a WT and a Tau neuron on the open box. Numbers: peak (red/black) rates in Hertz. (E) Examples of WT and Tau sequences (see Figure 3 for details) on one trajectory of the L-track. (F) Spike patterns of the same WT and Tau neurons within a time period of an open box session. (G and H) Mean number of sequences (in Z-score) (G) and mean location shift (H) for the sequences detected on the novel trajectories in WT and Tau mice. (I) Mean number of sequences (in Z-score) detected during the novel open box sessions in WT and Tau mice.
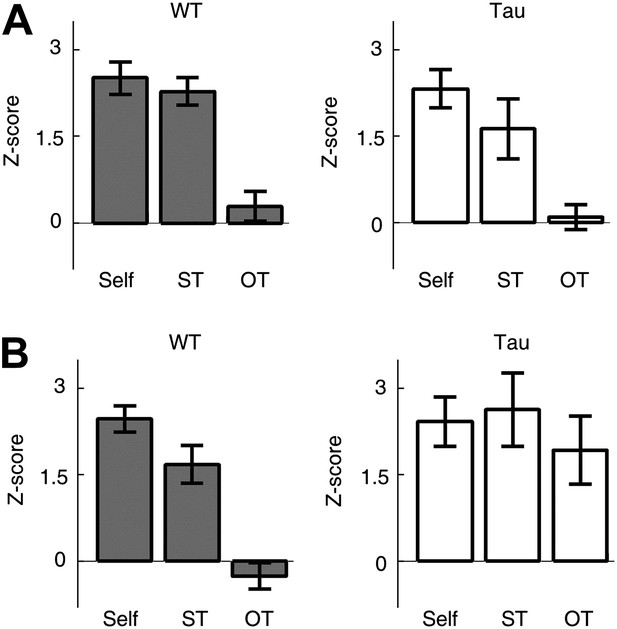
Tau sequences were trajectory-selective on the familiar track, but not on the novel track.
(A and B) Mean number of sequences on three types of trajectories (Self, ST, OT) of the familiar (A) and novel (B) track. For each template sequence derived on a trajectory of a session, we computed the number of sequences (in Z-score) that matched with the template on the trajectory itself (Self), on the same trajectory (ST) but in the other session, and on the other trajectory (OT) in the same session.
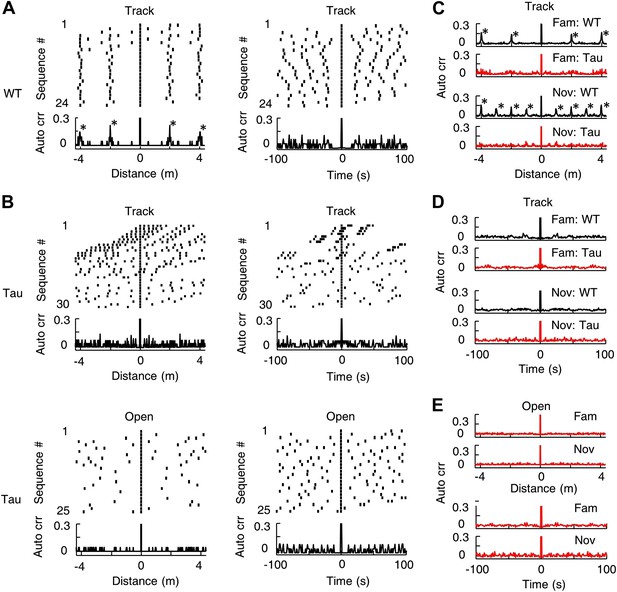
Tau sequences did not display obvious periodicity in space or time.
(A) The raster and auto-correlogram (Auto crr) of all sequences matched with a template during a track session of a WT mouse, plotted in the spatial (left) and temporal (right) domain. Each tick represents a sequence. For each sequence, we aligned its location/time at 0 and plotted the sequences occurring before and after at corresponding location/time gaps (top). The histogram of this raster plot is equivalent to the auto-correlogram of the sequence locations/times (bottom). The auto correlation value at 0 was truncated. Note the regular intervals in the spatial raster plot and periodic peaks in the spatial auto-correlogram. (B) Same as (A), but for the sequences matched with a template during a track session (Track) and an open box session (Open) of a Tau mouse. Note the inconsistent spatial intervals and the disappearance of the peaks in the track spatial auto-correlogram. (C) The average sequence spatial auto-correlograms during familiar (Fam; WT: N = 9 templates; Tau: N = 6) and novel (Nov; WT: N = 12 templates; Tau: N = 7) track sessions of WT and Tau mice. (D) Same as (C), but in the temporal domain. (E) The average sequence auto-correlograms of Tau mice in familiar (Fam; N = 10 searches) and novel (Nov; N = 4) open box sessions in the spatial (top) and temporal domain (bottom).
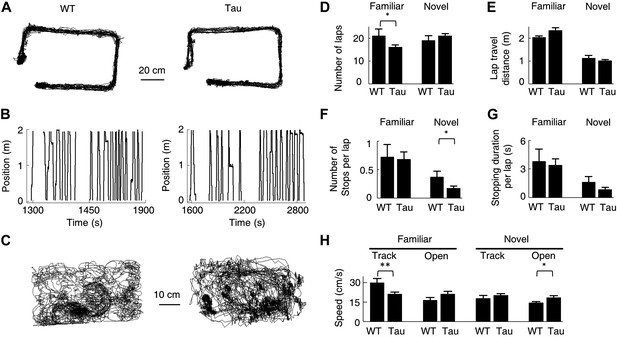
Behavioral quantifications of WT and Tau mice.
(A) The accumulative running paths of a WT and Tau mouse on the familiar rectangular track. (B) The paths in (A) were linearized lap-by-lap on the two track trajectories (ascending and descending lines) and plotted against time. The positions at the food wells were truncated. (C) The accumulative running paths of the WT and Tau mouse during an open box session. (D–G) the mean number of laps per session (D), mean travel distance per lap (E), mean number of stops per lap (F), and mean stopping duration per lap (G) for WT and Tau mice during familiar and novel track sessions. (H) Mean running speed of WT and Tau mice during familiar and novel track (Track) and open box (Open) sessions. *p<0.05; **p<0.01.
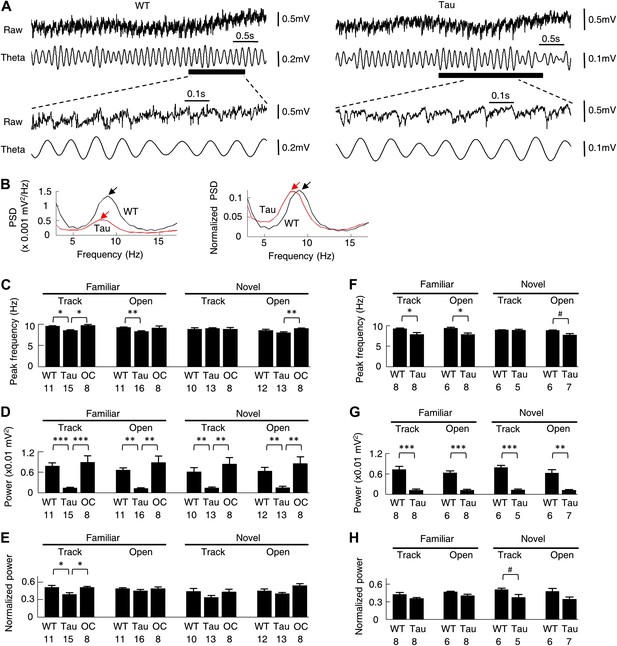
Theta oscillations were present in Tau mice.
(A) Raw and theta-filtered (6–12 Hz) LFP traces recorded from a WT and Tau mouse. Black bars: time windows during which a WT and a Tau firing sequence were detected. The raw and theta-filtered LFP traces within the time windows are expanded on the bottom. Note the similar theta oscillations between the WT and Tau mouse, but with different scale bars. (B) The absolute (left) and normalized (right) PSD curves of the raw LFP traces shown in (A). Black: WT; Red: Tau. Arrows: peaks in the theta frequency band. (C–E) mean peak theta frequency (C), mean absolute theta power (D), and mean normalized theta power (E) of raw LFPs recorded from WT and Tau mice and from mice with two other control genotypes (OC; +/− and −/+ combined), during familiar and novel track running (Track) and open box sessions (Open). Numbers: number of samples. (F–H) same as in (C)–(E), but for raw LFPs within the start and end times of detected WT and Tau firing sequences. #p<0.05; *p<0.01; **p<0.001; ***p<0.0001. For (C)–(E), the significance threshold was lowered to 0.01 because of the multiple (3) comparisons within a group and therefore only those comparisons with p<0.01 were marked.
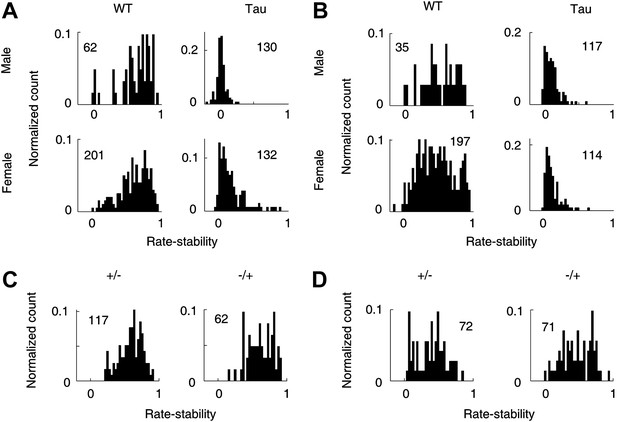
Gender and genotype differences.
(A and B) Distributions of rate-stability for WT and Tau neurons recorded from male and female mice on familiar (A) and novel (B) trajectories. (C and D) Distributions of rate-stability for neurons recorded from mice with other two control genotype (+/−, −/+) on familiar (C) and novel (D) trajectories. Plots are histograms normalized by total numbers of samples (numbers shown), each computed for one neuron on one trajectory.
Tables
Details of the recorded mice
| Animal name | Genotype | Sex | Age (m) | Brain weight (g) | CA1 thickness (µm) | Cortex thickness (µm) | Nf | Nn |
|---|---|---|---|---|---|---|---|---|
| AN1 | WT | M | 8–9 | NA | 56 | 800 | 7 | 3 |
| AN4 | WT | M | 8–9 | 0.350 | 51 | 790 | 16 | 0 |
| AN18 | WT | M | 7–8 | 0.334 | 55 | 760 | 8 | 10 |
| AN19 | WT | F | 7–8 | 0.324 | 51 | 690 | 41 | 30 |
| AN20 | WT | F | 7–8 | 0.392 | 51 | 690 | 16 | 37 |
| AN21 | WT | F | 8–9 | 0.328 | 51 | 690 | 31 | 34 |
| AN10 | +/− | M | 8–9 | 0.364 | NA | NA | 42 | 33 |
| AN11 | +/− | M | 7–8 | 0.356 | 53 | 760 | 10 | 15 |
| AN12 | −/+ | F | 7–8 | 0.310 | 50 | 800 | 12 | 18 |
| AN16 | −/+ | M | 8–9 | 0.332 | 60 | 800 | 15 | 18 |
| AN5 | Tau | F | 8–9 | 0.192 | 26 | 580 | 17 | 15 |
| AN13 | Tau | F | 8–9 | 0.226 | 22 | 580 | 14 | 10 |
| AN14 | Tau | F | 8–9 | 0.236 | 27 | 580 | 24 | 25 |
| AN15 | Tau | M | 8–9 | 0.214 | 26 | 580 | 10 | 18 |
| AN22 | Tau | M | 7–8 | NA | 28 | 550 | 20 | 10 |
| AN23 | Tau | M | 7–8 | 0.198 | 24 | 550 | 7 | 10 |
| AN24 | Tau | F | 7–8 | 0.232 | 27 | 550 | 14 | 14 |
| AN25 | Tau | M | 7–8 | 0.226 | 26 | NA | 29 | 27 |
| Total | 333 | 327 |
-
Genotype: see ‘Materials and methods’. Age: age of the mouse in months when data were recorded. Brain weight: weight of the brain with the cerebellum and olfactory bulb removed. CA1 thickness: thickness of the pyramidal cell layer in the dorsal CA1 of HP; cortex thickness: thickness of the parietal cortex above the dorsal CA1; measurements were made in the coronal sections that contained the most dorsal point of CA1, about the section at Bregma -2.06 (Figure 48) of the Franklin and Paxinos mouse atlas (Franklin and Paxinos, 2008). Nf/Nn: number of CA1 neurons recorded in the familiar/novel room. NA: number not available. Note the lower brain weight, the thinner CA1 pyramidal layer, and the thinner cortex in Tau mice.
High-order sequence analysis results for individual template sequences derived on the familiar trajectories
| Template sequence | Animal name | Genotype | Self traj. | Z (S1T1) | Z (S1T2) | Z (S2T1) | Z (S2T2) | Z (O1) | Z (O2) |
|---|---|---|---|---|---|---|---|---|---|
| AGFHJ | AN4 | WT | S2T2 | 0.01 | 2.8 | −0.4 | 3.2 | NA | NA |
| ZSBPNA | AN19 | WT | S1T1 | 2.6 | −0.7 | 1.4 | 0.3 | 0.8 | −1.7 |
| JHLKZE | AN19 | WT | S1T2 | 1.6 | 2.7 | −0.4 | 1.8 | −0.1 | 0.6 |
| IYOBZENA | AN19 | WT | S2T1 | 1.9 | 2.3 | 3.5 | 1.3 | 0.7 | 2.7 |
| ZSaFbB | AN19 | WT | S2T2 | −0.3 | 1.7 | 0.9 | 1.9 | 1.9 | 0.3 |
| GHKJBDF | AN20 | WT | S1T1 | 1.6 | 1.2 | 2.4 | 1.0 | −1.8 | 0.9 |
| JHAKC | AN20 | WT | S1T2 | −0.6 | 1.3 | −0.3 | 2.1 | 0.02 | −0.7 |
| GHCKJBDF | AN20 | WT | S2T1 | 1.8 | 0.3 | 1.4 | 1.3 | −0.6 | 1.4 |
| GJAHLC | AN20 | WT | S2T2 | 0.5 | 1.1 | 0.4 | 0.8 | −1.8 | −0.3 |
| XIGQOZ | AN21 | WT | S1T1 | 4.0 | −1.2 | 3.6 | −1.0 | −2.0 | −1.0 |
| IWQVHP | AN21 | WT | S1T2 | −0.8 | 3.0 | −1.2 | 3.3 | 0.2 | 0.0 |
| GOZTHR | AN21 | WT | S2T1 | 1.9 | 0.5 | 3.6 | 0.3 | 1.0 | −1.5 |
| OFINKDV | AN21 | WT | S2T2 | −0.9 | 3.8 | 0.5 | 3.3 | −0.5 | −1.4 |
| ACHFDG | AN5 | Tau | S2T1 | 3.0 | −0.4 | 2.5 | −0.06 | −1.2 | −1.6 |
| FABHD | AN13 | Tau | S1T1 | 4.0 | −0.1 | 2.9 | 2.0 | 2.1 | 2.9 |
| BFAHD | AN13 | Tau | S2T1 | 1.7 | 1.1 | 2.1 | −0.9 | 0.04 | 0.5 |
| FEBA | AN15 | Tau | S2T1 | 1.0 | 0.9 | 3.2 | −0.4 | 2.6 | 1.9 |
| ILFBK | AN22 | Tau | S1T1 | 1.4 | 0.5 | −0.8 | −0.8 | 1.2 | −0.5 |
| FLBDIC | AN22 | Tau | S2T1 | 0.1 | 1.4 | 2.2 | 0.7 | 2.2 | 2.9 |
| PKLEDO | AN25 | Tau | S1T2 | 1.1 | 1.9 | 0.4 | 3.3 | 2.4 | 2.4 |
| PKTLEOMA | AN25 | Tau | S2T2 | 0.7 | 1.8 | 0.0 | 1.3 | 3.4 | 1.9 |
-
Template sequence: each letter (case-sensitive) represents a cell; same letters across different templates but within the same animal represent the same cells. Self traj.: The trajectory where the template was derived. Z: Number of detected matches (in Z-score) with a template sequence on a trajectory or in an open box session. S1T1: Trajectory one of session one, S1T2: Trajectory two of session one, so forth; O1: First open box session, so forth; numbers on self-trajectories are in bold; numbers that are statistically significant (p<0.05) are in red. NA: not available (not all experiments included two trajectory running sessions and two open box sessions).
High-order sequence analysis results for individual template sequences derived on the novel trajectories
| Template sequence | Animal name | Genotype | Self traj. | Z (S1T1) | Z (S1T2) | Z (S2T1) | Z (S2T2) | Z (O1) | Z (O2) |
|---|---|---|---|---|---|---|---|---|---|
| IHFGE | AN18 | WT | S1T1 | 3.4 | 0.1 | 1.4 | 0.2 | 1.0 | 1.1 |
| HEIFG | AN18 | WT | S1T2 | 0.4 | 4.3 | −1.4 | 2.7 | 1.6 | −0.8 |
| HICFBG | AN18 | WT | S2T2 | 1.9 | 3.5 | −0.3 | 3.6 | 2.5 | 0.8 |
| NAOJBD | AN19 | WT | S1T1 | 2.5 | −1.5 | 1.4 | 0.1 | 0.1 | −0.1 |
| OKDHFBAJN | AN19 | WT | S1T2 | −0.1 | 1.6 | −1.0 | 0.7 | 1.1 | 0.1 |
| LANOB | AN19 | WT | S2T1 | 3.1 | −0.6 | 3.0 | −1.1 | −0.5 | −0.4 |
| OKFCNA | AN19 | WT | S2T2 | −0.4 | 1.3 | −0.4 | 1.7 | 1.6 | 0.1 |
| ISRPBGJXKQAY | AN20 | WT | S1T1 | 1.5 | −1.6 | 2.0 | 1.2 | 0.9 | −1.1 |
| BIPRS | AN20 | WT | S1T2 | −0.1 | 1.1 | 0.2 | −0.3 | 0.3 | 0.2 |
| CTDOISREBXG | AN20 | WT | S2T1 | 1.8 | −0.5 | 1.8 | −0.5 | 1.1 | 0.6 |
| SRUPNL | AN20 | WT | S2T2 | −0.5 | 1.8 | 1.0 | 2.8 | 0.2 | 0.9 |
| IXPHNSC | AN21 | WT | S1T1 | 2.5 | −1.5 | 3.2 | 0.0 | −0.7 | −1.1 |
| PXHYT | AN21 | WT | S1T2 | 0.3 | 1.8 | −0.4 | −0.1 | 1.5 | −0.1 |
| KIJXSC | AN21 | WT | S2T1 | 2.8 | −0.5 | 2.7 | 0.4 | 0.4 | 1.1 |
| UMNSEO | AN21 | WT | S2T2 | −1.6 | −0.4 | 1.1 | 2.5 | 1.7 | 0.7 |
| HGPOL | AN14 | Tau | S1T1 | 1.7 | −0.1 | 1.1 | −1.4 | 1.4 | −0.3 |
| GEDMHCB | AN15 | Tau | S1T1 | 3.3 | 1.7 | 2.8 | 1.6 | 0.5 | 3.1 |
| DMHCB | AN15 | Tau | S2T1 | 4.2 | 1.9 | 3.8 | −0.4 | 0.8 | 0.7 |
| GEDM | AN15 | Tau | S2T2 | 3.3 | 2.5 | 2.9 | 2.1 | 1.4 | 3.1 |
| ABDE | AN24 | Tau | S1T1 | 2.6 | 3.0 | NA | NA | 1.3 | NA |
| DEABI | AN24 | Tau | S1T2 | 4.6 | 3.5 | NA | NA | 2.6 | NA |
| SGHER | AN25 | Tau | S1T1 | 2.3 | 1.4 | NA | NA | 2.0 | NA |
| KMFGHE | AN25 | Tau | S1T2 | 1.0 | 2.3 | NA | NA | 1.4 | NA |
-
See Table 2 for details.





