Her2 activation mechanism reflects evolutionary preservation of asymmetric ectodomain dimers in the human EGFR family
Figures
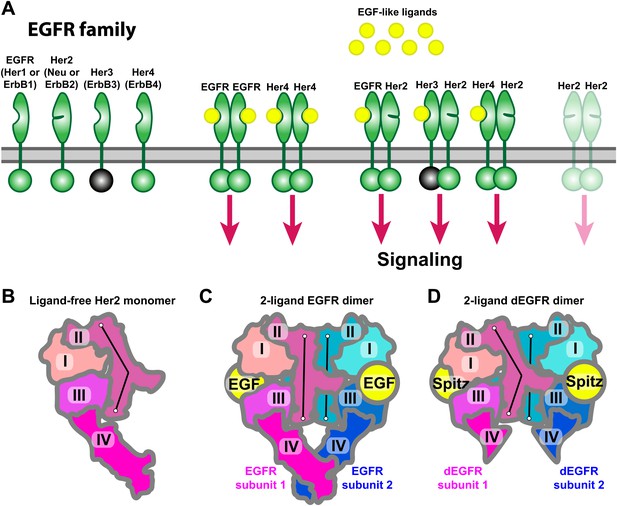
Receptors of the human EGFR family and conformations of their ectodomains.
(A) Left: the four members of the human EGFR family, each consisting of an ectodomain, a single-pass transmembrane helix, and an intracellular module that includes a kinase domain. As shown, Her2 bears a closed ligand binding site and does not bind EGF-like ligands. Her3 is kinase-dead, and its intracellular module is thus colored gray. Right: common homo- and heterodimers of the EGFR family. The Her2 homodimer is rendered semitransparent to indicate its instability in normal cell conditions. (B) Schematic of the ligand-free Her2 ectodomain monomer, as observed crystallographically (PDB entries 1N8Z, 2A91, 1S78, 3N85, and 3MZW). Domain II is bent. (C) Schematic of the crystal structure of the 2-ligand EGFR ectodomain dimer (PDB entry 1NJP). The dimer is symmetric, and domain II is straight in both subunits. (D) Schematic of the crystal structure of the dEGFR ectodomain dimer (PDB entry 3LTF). Although Spitz ligands are bound to both subunits, the structure is asymmetric; domain II is straight in one subunit but bent in the other. Domain V and part of domain IV were not resolved in this crystal structure and are not shown in the schematic. The conformations of the bent and straight domain IIs in (B), (C), and (D) are indicated by the black lines.
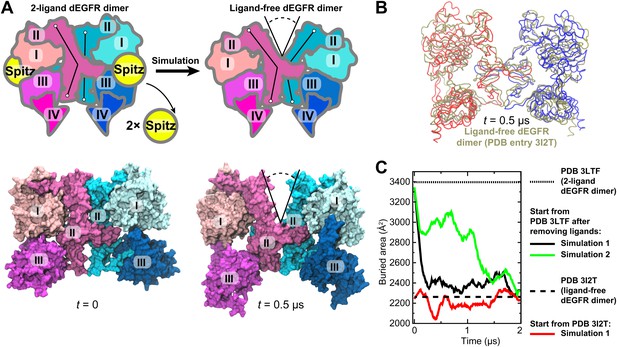
Simulations reproduce the gap in the interface of the dEGFR ectodomain dimer.
(A) The schematic shows the 2-ligand dEGFR ectodomain dimer on the left. In simulations initiated from this structure with both ligands removed, gap opened in the dimer interface, as indicated by a V-shaped outline in the right diagram. In the ligand-free dimer, domain II is bent in both subunits. The simulation snapshots are shown below the schematic diagrams. (B) The simulated dEGFR dimer from (A), at t = 0.5 µs, is compared with the crystal structure of the ligand-free dEGFR dimer (tan). Molecular renderings (A, B) omit domains IV and V for clarity (although the crystallographically resolved portion of domain IV was present in simulations). (C) The surface area buried within the dimer interface (counting the contributions from domains I, II, and III). The results of three independent simulations are shown, two starting from the 2-ligand crystal structures (after removal of the ligands), and one from the ligand-free crystal structure.
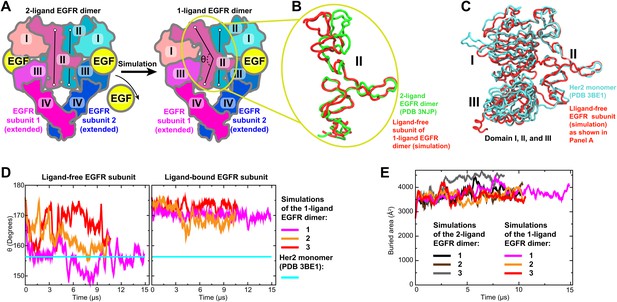
Bending of domain II.
(A) A schematic showing how the model of the 1-ligand EGFR dimer (right) is generated by simulation after the removal of one EGF ligand from the crystal structure of the 2-ligand dimer (left). The resulting structure is asymmetric: domain II of the ligand-free subunit is bent and the binding site is closed, whereas domain II of the ligand-bound subunit is straight and the binding site is open (as is the case in both subunits of the 2-ligand dimer). The angle θ, which characterizes the bending of domain II, is measured between the Cα atoms of EGFR residues 194, 239, and 296. (B) Bending of domain II. A simulation snapshot of the ligand-free subunit (red) from the 1-ligand EGFR dimer is overlaid with the crystal structures of a subunit from the 2-ligand EGFR dimer (green). EGFR residues 240–309 are used for reference. (C) An overlay of the ligand-free subunit of the 1-ligand EGFR dimer, generated by simulation (red), with the crystal structure of Her2 monomer (cyan). In the interest of clarity, domain IV is not shown. (D) The angle θ (illustrated in [A]) is shown as function of time in the three independent simulations of the 1-ligand EGFR homodimer, for the ligand-free subunit (middle) and for the ligand-bound subunit (right). The value of θ in the crystal structure of Her2 monomer is indicated by a straight line. (E) The buried surface area at the interfaces of 1-ligand and 2-ligand EGFR dimers in simulation.
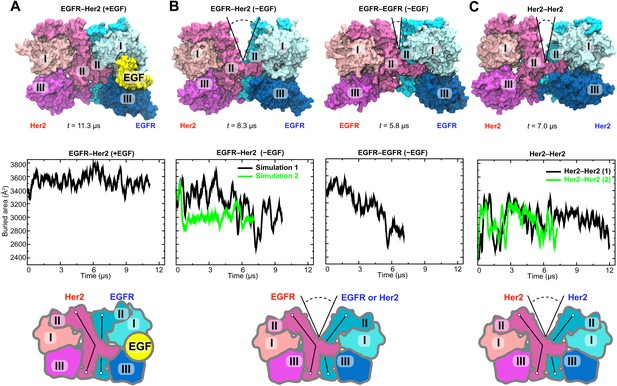
Simulations of the EGFR–Her2 heterodimer and the Her2 homodimer.
(A) The EGFR–Her2 heterodimer with the ligand bound to EGFR (‘+EGF’). (B) The ligand-free EGFR–Her2 heterodimer or EGFR–EGFR homodimer (‘−EGF’). (C) The Her2 homodimer. Top: snapshots from the simulations. Middle: plots of the surface area buried within the dimer interface (counting the contributions from domains I, II, and III) as a function of simulation time (for EGFR–Her2 [−EGF] and Her2–Her2, results of two independent simulations are shown). Bottom: schematics illustrating the conformation of domain II and the dimer interface. Conformations of the bent and straight domain II are highlighted by the black lines, and the gap in the dimer interface is indicated by a V-shaped outline when present. For clarity, domain IV is not shown.
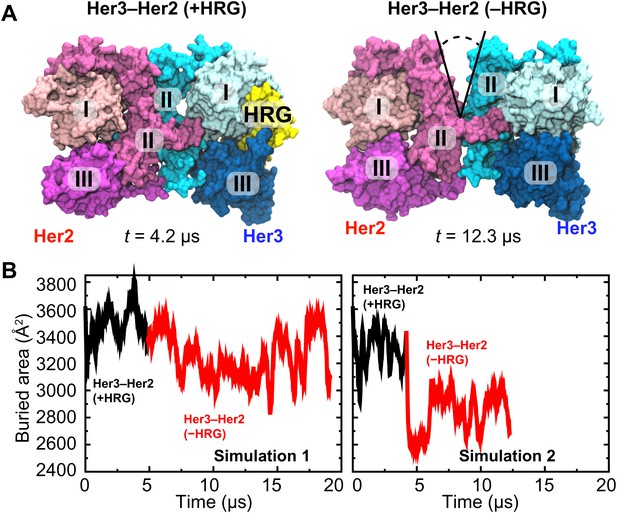
Her3–Her2 heterodimer.
(A) Snapshots from the simulations of the Her3–Her2 heterodimer with (left) and without (right) HRG bound to Her3. At the end of the simulation with HRG bound to Her3, HRG was removed, and the resulting system was resolvated and further simulated without the ligand. A gap opened in the dimer interface, as illustrated by the snapshot on the right. For clarity, these images omit domain IV. (B) The surface area buried within the dimer interface, counting the contributions only from domains I, II, and III, plotted as a function of time. Two independent sets of two simulations each (with and without HRG) are shown.
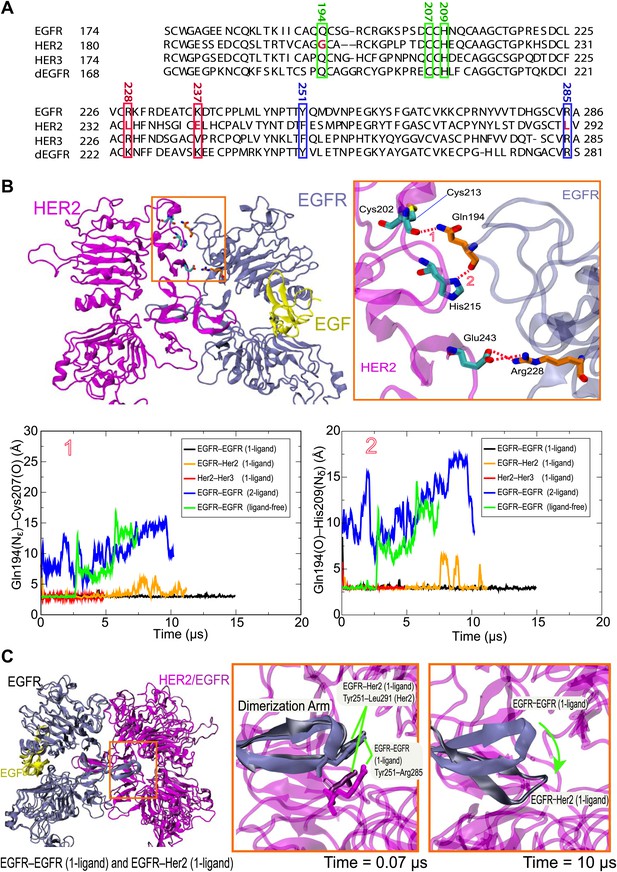
Details of the dimer interfaces in Her2 heterodimers.
(A) The sequence alignments of EGFR, Her2, Her3, and dEGFR for the part of domain II corresponding to EGFR residues 167–286. Three sets of residues (green, red, and blue, respectively), which are involved in three sets of important interactions at the dimer interface, are highlighted. (B) Top: the role of EGFR residue Gln194 in the EGFR–Her2 dimer interface. Hydrogen bonds and salt bridges are indicated by red dashed lines. Bottom: the distances between the key atoms involved in these hydrogen bonds are shown as functions of time. (C) Left: superimposition of 1-ligand EGFR homodimer and 1-ligand EGFR–Her2 heterodimer, with dimerization arms highlighted by the red box. Middle and right: Her2’s dimerization arm in the EGFR–Her2 heterodimer is conformationally less stable than the dimerization arm of the ligand-free subunit of the 1-ligand EGFR homodimer, likely because the Tyr251–Arg285 cation–π interaction is only present in the latter dimer. Dark blue is used for the EGFR dimerization arm in the EGFR homodimer, and dark gray for the EGFR dimerization arm in the EGFR–Her2 heterodimer.
Videos
The simulation of the ligand-free EGFR–Her2 ectodomain heterodimer with Her2 colored red and EGFR blue. A gap between the N-terminal portion of each subunit’s domain II develops during the simulation.





