Regulated aggregative multicellularity in a close unicellular relative of metazoa
Figures
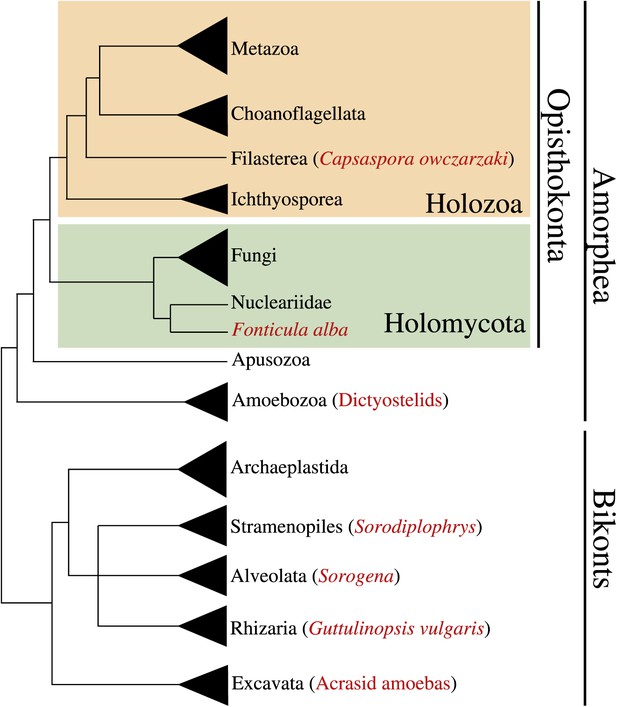
Phylogenetic position of Capsaspora owzarzaki within the eukaryotes.
The Holozoa clade (in yellow) includes Metazoa and their closest unicellular relatives: Choanoflagellata, Filasterea, and Ichthyosporea. C. owczarzaki represents one of the two known filastereans taxa that form the sister-group of choanoflagellates and metazoans. Other major eukaryotic groups are shown. Within each group, species or clades with aggregative multicellularity (see text) are highlighted in red.
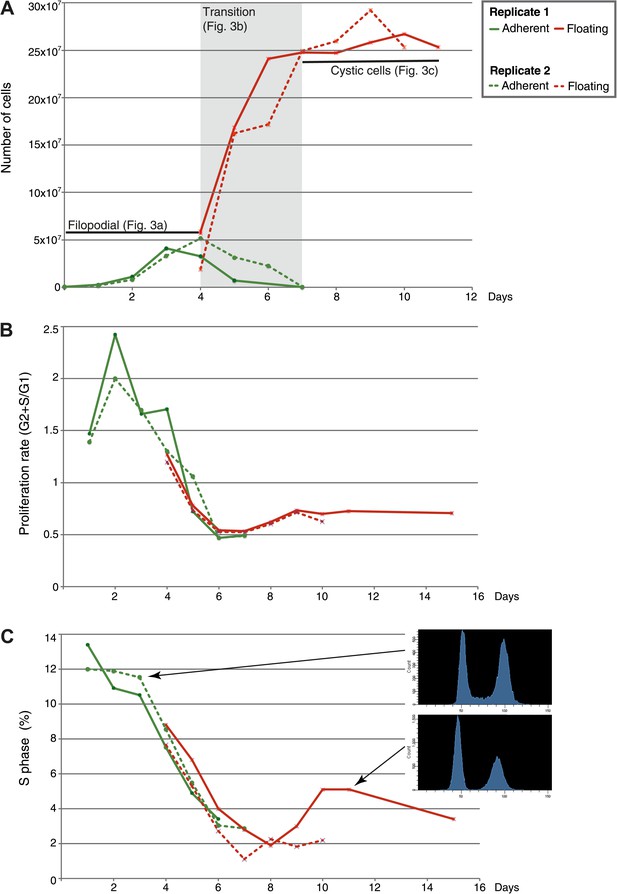
Flow-cytometry analysis of C. owzarzaki cell cycle.
(A) Total number of cells per day in each fraction (adherent and floating, see ‘Materials and methods’). (B) Proliferation rate per day. (C) Percentage of cells in S-phase per day and two examples of DNA-content profiles obtained from days 3 and 11. Note the reduction in the number of G2/M cells (second peak) and the drastic reduction in S-phase cells (the area between the two peaks). Data from adherent cells (‘Materials and methods’) is shown in green and data from floating cells in red. Experimental replicate 1 results are shown with solid lines and replicate 2 results with dashed lines.
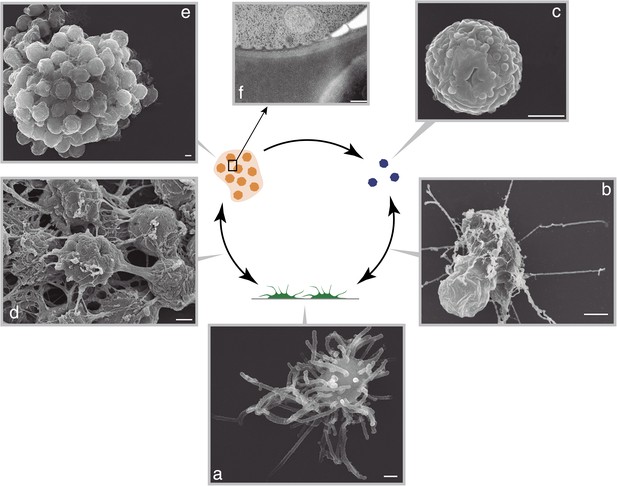
C. owzarzaki life cycle.
(A) Filopodial stage cells, amoebas with long filopodia. (B) Transition from filopodial to cystic stage: cells retract filopodia. (C) Cystic stage cells are rounded cysts without filopodia and slightly smaller than filopodial cells. (D) Transition from filopodial to aggregative stage: cells attach to each other and an extracellular matrix appears. (E) Mature aggregate. (F) Transmission EM showing adjacent cells in the aggregate separated by extracellular matrix. Arrows indicate the observed stage inter-conversions. Scale bars = 1 μm, except in panel D = 200 nm.
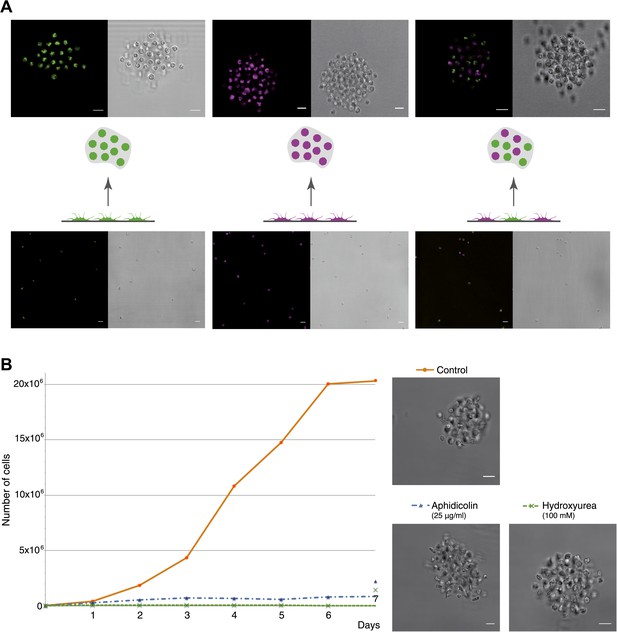
C. owczarzaki cell clusters form by active aggregation, not clonal cell division.
Aggregation was induced in different stained cell populations (‘Materials and methods’). (A) Left, population of cells stained with Lysotracker (green), uniformly green aggregates are formed. Center, population of cells stained with Chormeo Mitochondrial Staining (cyan), uniformly cyan aggregates are formed. Right, two independently stained populations of cells (green or cyan) are mixed and dual color aggregates are formed, indicating that cells from different origins aggregate to each other. (B) Total number of cells per day in control cells, aphidicolin-treated cells and hydroxyurea-treated cells. Note that cell division is blocked by both aphidicolin and hydroxyurea. Aggregate formation was evaluated under each condition. All cells, even those treated with aphidicolin or hydroxyurea, developed aggregates. A representative aggregate is shown for each condition. Scale bars= 10 μm.
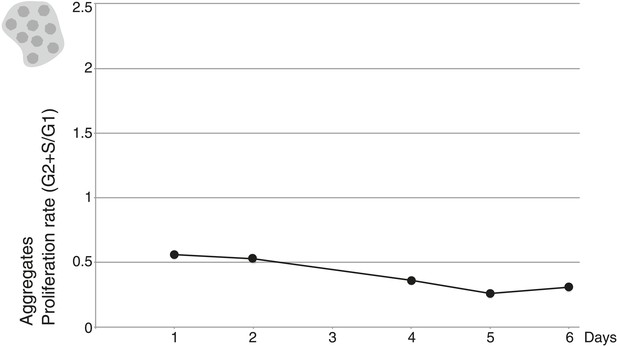
Proliferation rate per day of aggregative cells.
https://doi.org/10.7554/eLife.01287.010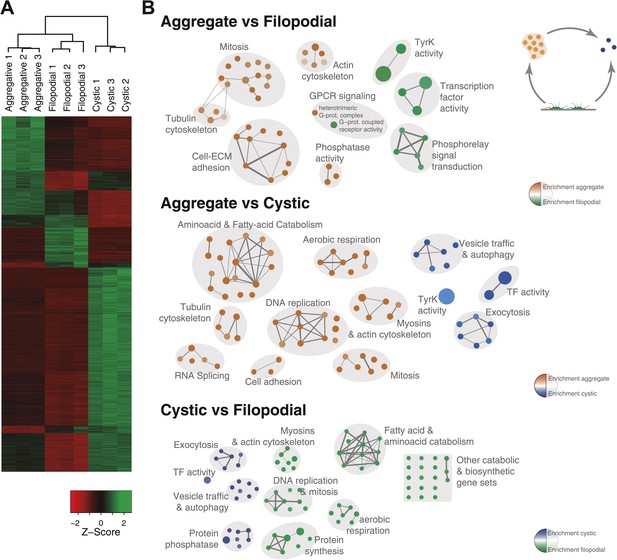
Differential gene expression in C. owczarzaki.
(A) Heatmap showing differential gene expression in the different biological replicates of each stage. Only genes with cRPKM ≥ 5 in at least one sample and with a 2-fold expression change in at least one pair-wise comparison are shown. (B) Gene set enrichment analysis (GSEA) for the different cell stages (‘Materials and methods’). Orange represents enrichment in the aggregative stage, blue in the cystic stage, and green in the filopodial stage, with color intensity proportional to enrichment significance. The node size is proportional to the number of genes associated to the GO category, and the width of the edges is proportional to the number of genes shared between GO categories. Groups of functionally related GOs are manually circled and assigned a label.
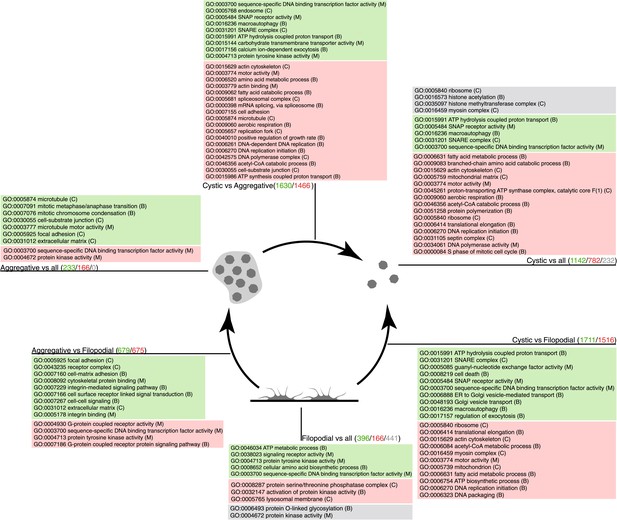
GO enrichment in sets of differentially expressed genes.
Pairwise (Aggregative vs Filopodial, Cystic vs Aggregative and Cystic vs Filopodial) and one-versus-all comparisons are indicated. The significantly overrepresented GO categories (‘Materials and methods’) are shown for sets of overexpressed (green) and downregulated (red) genes and for genes with differential intron retention (gray). The number of genes included in each set is indicated with the same color code.
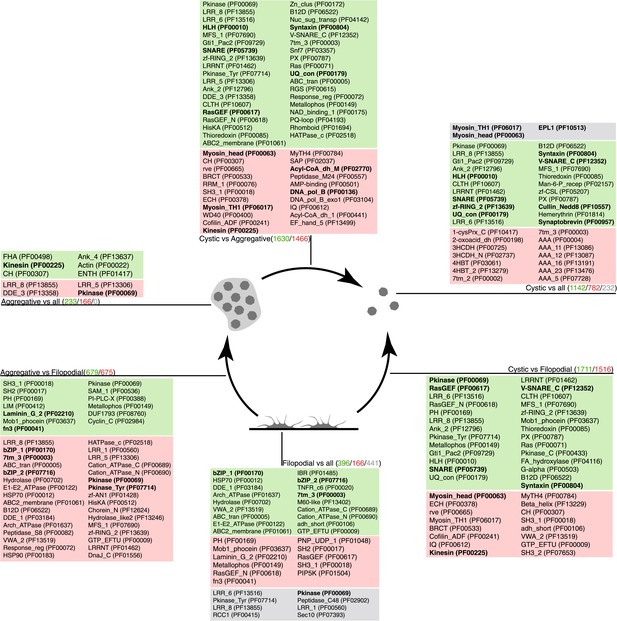
Pfam protein domain enrichment in sets of differentially expressed genes.
Pairwise (Aggregative vs Filopodial, Cystic vs Aggregative and Cystic vs Filopodial) and one-versus-all comparisons are indicated. Significantly overrepresented Pfam domains (‘Materials and methods’) are shown for sets of overexpressed (green) and downregulated (red) genes and for genes with differential intron retention (gray). The number of genes included in each set is indicated with the same color code. Those Pfam domains mentioned in the text are shown in bold.
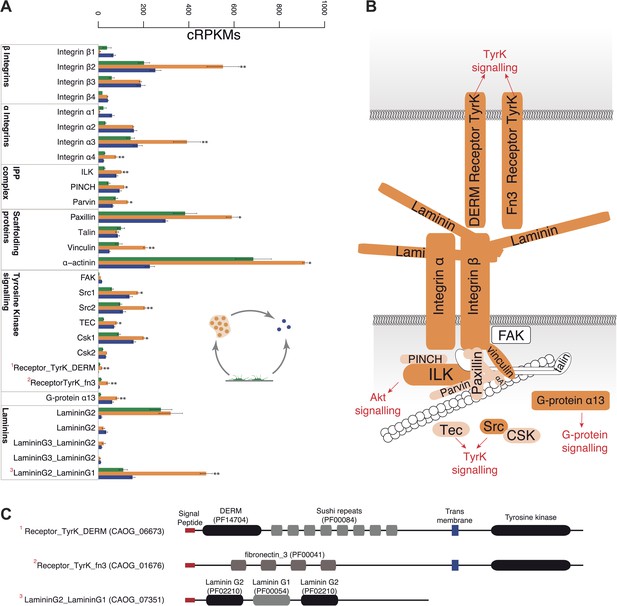
Expression of cell–ECM adhesion genes.
(A) Barplot of the expression values of each gene in the different stages, showing overexpression of most components in the aggregate stage (orange). Asterisks indicate that the gene is significantly differentially expressed in both (two asterisks) or only one (one asterisk) pair-wise comparison (agg. vs fil. and agg. vs cyst.). Bars show standard error. (B) Schematic representation of the putative C. owczarzaki integrin adhesome and putative associated signalling proteins, colored according to overexpression in aggregates as shown in the barplot (dark orange, two asterisks; light orange, one asterisk; and white, no differences in expression). (C) Specific protein domain architectures for the fibronectin and DERM receptor tyrosine kinases (CAOG_01676 and CAOG_06673) and for the laminin protein (CAOG_07351).
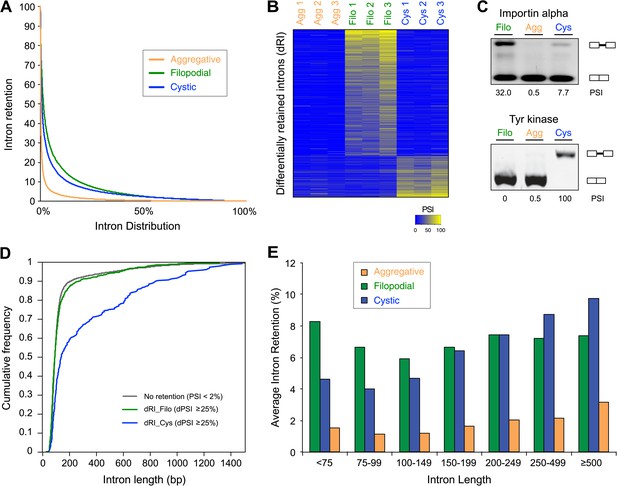
Regulated alternative splicing in C. owzarzaki.
(A) Plot of percentage of intron inclusion by intron in rank order for the three studied cellular stages. Filopodial (green) and cystic (blue) stages show higher intron retention levels than the aggregative stage (orange) (p<2.2e-16, Wilcoxon Rank Sum test). (B) Heatmap of PSIs of filopodial- and cystic-specific differentially retained introns across three replicates for each cellular stage. (C) Examples of stage-specific intron retention. (D) Intron length distributions for differentially retained introns in cystic (blue), filopodial (green), and weakly retained introns (gray). (E) Relationship between intron length and retention. Percentage of average intron retention in each of the three cellular stages for different bins of intron size. In the cystic stage, the percentage of intron retention increased with intron length.
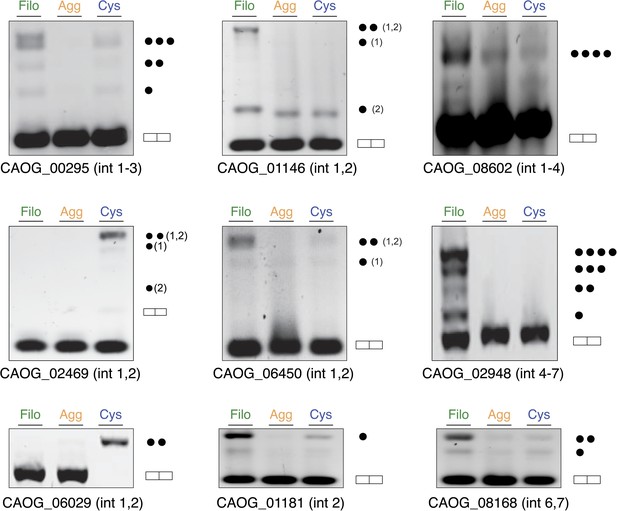
Intron retention validation (see ‘Materials and methods’).
Genes with confirmed intron retention events are indicated using gene IDs. Each dot corresponds to the inclusion of one intron (introns in C. owczarzaki usually have very similar lengths, and thus different combinations of the same size introns cannot be differentiated).
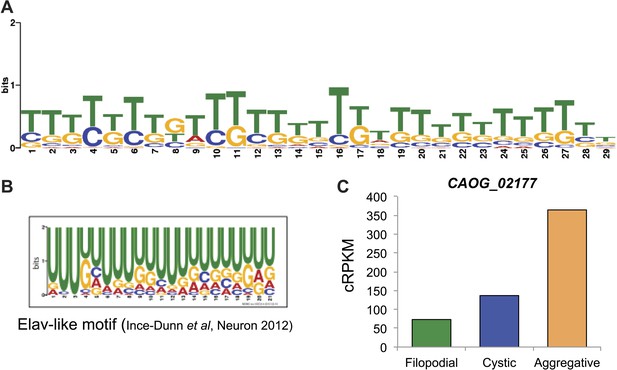
Possible role for an Elav-like ortholog in the negative regulation of filopodial stage-specific dRIs.
(A) Most significantly enriched motif in filopodial stage-specific dRIs, obtained by MEME. (B) Consensus motif obtained by CLIP-Seq data for an Elav-like member in mammals by Ince-Dunn et al. (2012) and that closely resembles the motif in (A). T∼U. (C) Expression (measured as cRPKMs) of CAOG_02177, a Elav-like ortholog from C. owczarzaki that shows lower expression in filopodial stage.
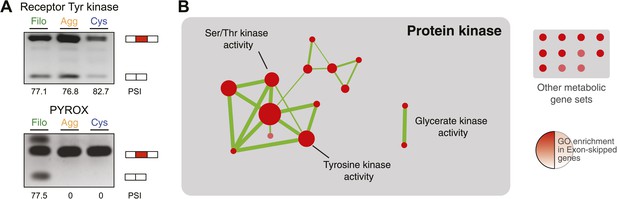
Regulated exon-skipping in C. owzarzaki.
(A) Examples of exon skipping. (B) Gene set enrichment analysis (GSEA) of the genes containing cassette exons that are differentially-regulated among cellular stages showing high enrichment for protein kinase-associated activities.
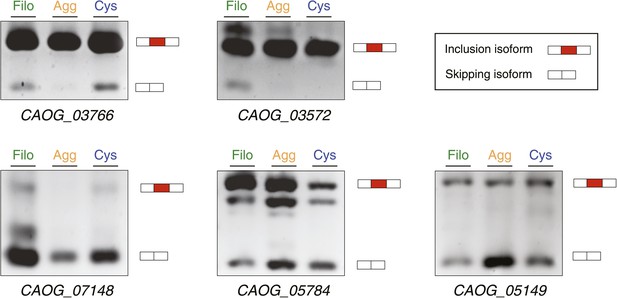
Exon Skipping validation by RT-PCR (‘Materials and methods’).
Note that DNA heteroduplexes between the two isoforms may be formed and appear as extra bands. Bars in the key correspond to the exons amplified: the alternative exon in red, and the adjoining constitutive exons in white.
Videos
C. owczarzaki filopodial amoeba stage cells crawling.
Dark and refringent vesicles can be observed inside each cell. Up to nine different cells can be observed in the video. Also available on YouTube: http://youtu.be/0Uyhor_nDts.
C. owczarzaki cells aggregation.
Also available on YouTube: http://youtu.be/83HB8srWQw4.
C. owczarzaki cells aggregation.
Also available on YouTube: http://youtu.be/OvI6BvBucrc.
Additional files
-
Supplementary file 1
Exon-skipping events in C. owczarzaki.
For each event, the table shows the affected gene, a reference of the skipping event, the coordinates of the skipped exon, its length and if it is multiple of three nucleotides, the full coordinates of the event (i.e., including the donor of the upstream constitutive exons and acceptor of the downstream constitutive exon), percentage of inclusion of the exon among all transcripts present at each stage, general protein domain structure and a brief description of the gene, and information on which region of the protein is affected by the exon-skipping event (in orange if impacting a structured domain). Protein kinases are highlighted in dark green and tyrosine kinases in light green.
- https://doi.org/10.7554/eLife.01287.020





