p53 activity is selectively licensed in the Drosophila stem cell compartment
Figures
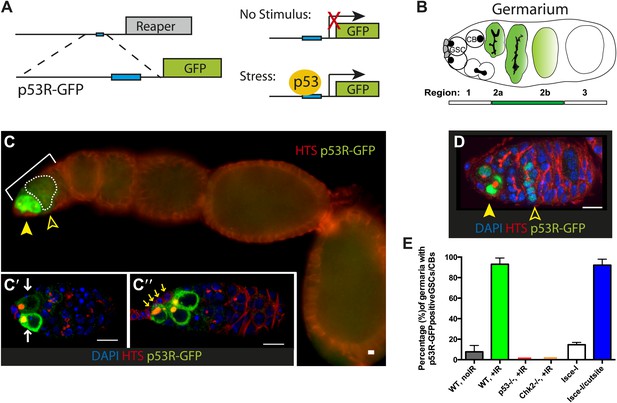
Genotoxic stress selectively triggers p53 activity in ovarian stem cells.
(A) Construction of p53 biosensors. A well-characterized p53 enhancer (black line) that contains a p53 consensus binding site (blue box) conserved from flies to humans resides upstream of the reaper locus (gray box) (Brodsky et al., 2000). A 150-bp fragment containing this enhancer was placed upstream of GFP (p53R-GFP). Transgenic fly strains are made with two reporter constructs, one contains a nuclear localization signal for GFP (p53R-GFPnls) and the other one does not (p53R-GFPcyt). Stimuli that trigger p53 activation induce GFP expression. These biosensors require wild-type p53 and are effective readouts for p53 function. (B) Germline stem cells (GSCs) are in contact with cap cells (in gray) at the apical tip of the germarium and undergo self-renewing division to produce a GSC and cystoblast (CBs) (Spradling et al., 2001). In unperturbed ovaries, programed activation of the p53R-GFP biosensor is triggered by meiotic recombination in region 2 of the germarium, marked by open arrowhead in (C) and (D) (Lu et al., 2010). (C) After radiation challenge (IR) the p53R-GFPcyt biosensor (green) is selectively induced in ovarian GSCs and CBs noted by a solid arrowhead. Bracket denotes the germarium. The open arrowhead and dotted line indicates p53 activation in region 2 prompted by meiosis. Insets (C’ and C’’) are confocal images from different irradiated germaria counterstained with DAPI (blue). p53R-GFPcyt induction (green) initiates in GSCs that exhibit rounded fusomes (C’ white arrows) labeled by α-HTS (Hu li tai shao, red) and are in contact with cap cells (C’’ yellow arrows). Cells that activate p53 in (C’ and C’’) were confirmed to be germ cells by α-Vasa staining (shown in Figure 1—figure supplement 2C–D′′). (D) An engineered DNA double-stranded break (DSB) mediated by I-SceI (see texts and ‘Materials and methods’) induces the p53R-GFPnls biosensor (green) in GSCs/CBs, noted by a solid arrow. Open arrow indicates meiotic p53. The germarium is counterstained with α-HTS (red) and DAPI (blue). (E) Quantifies the percentage of germaria activated for the p53 biosensors in GSCs and their immediate progeny. Note that the perturbation-dependent responses reported here are all highly penetrant. Selective activation is IR (green) and I-SceI (blue) dependent at the 0.001 significance level. Note that biosensor activation did not occur in p53−/− (red) or chk2−/− (orange) mutants (see Figure 1—figure supplement 2A′,A′′). Sample sizes are combined from at least two independent trials (available in Figure 1—source data 1). All scale bars represent 10 μm. In panels C–C’’ the p53R-GFPcyt reporter was used. In panel D, the p53R-GFPnls biosensor was used.
-
Figure 1—source data 1
Validation of the p53R-GFP biosensors.
In A and B reporter activation is quantified as indicated. (A) Reporter activation in female GSCs/CBs is p53 dependent and Chk2 dependent but independent of the topoisomerase, Spo11. Reporter activation in testis is also stimulus dependent and p53 dependent. p53, Chk2, or Spo11 status are noted in column 1. The reporter used (nuclear or cytoplasmic) is indicated in column 2. Column 3 shows unirradiated controls in which the percent reporter activation in GSCs/CBs is provided, as well as the total number of germaria or testis that were assayed. Column 4 shows reporter activation in irradiated tissue at 24 hr post-irradiation with percentage of germaria or testis with GFP positive GSCs/CBs and the number of samples assayed. Quantification of reporter activation is from three independent trials in the ovary and two independent trials in the testis. (B) Quantification of p53-GFPnls in region 1 of flies containing I-SceI endonuclease by itself or with the I-SceI cutsite. Reporter activation in I-SceI expressing animals that also have the I-SceI cutsite is comparable to wild-type irradiated flies (A). Quantification of reporter activation is from two independent trials. (C) Quantification of p53-GFPnls in GSCs and follicle cells of flies heterozygous (ATR+/−) or mutant for ATR (ATR−/−). After irradiation challenge, p53 activation is highly penetrant in both ATR+/− and ATR−/− genotypes. ATR mutants show a robust induction of reporter activation in follicle cells after irradiation.
- https://doi.org/10.7554/eLife.01530.004
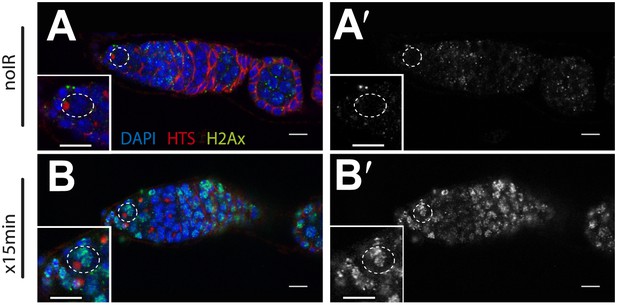
Wide-spread DNA breaks after irradiation.
Unirradiated (A) and irradiated (B) WT germaria were stained for the Drosophila counterpart of H2Ax, designated pH2Av (green). A′ and B′ show the pH2Av channel alone (white) from A to B respectively. DNA double-stranded breaks visualized by pH2Av (green) appear throughout germarium within 15 min after irradiation. Germaria in A and B are counterstained with α-HTS (red) and DAPI (blue). Note that panels A and A′ are the same as in Figure 4—figure supplement 2 A-A′. Panels B and B′ are the same as in Figure 4—figure supplement 2 C-C′. Scale bar, 10 μm.
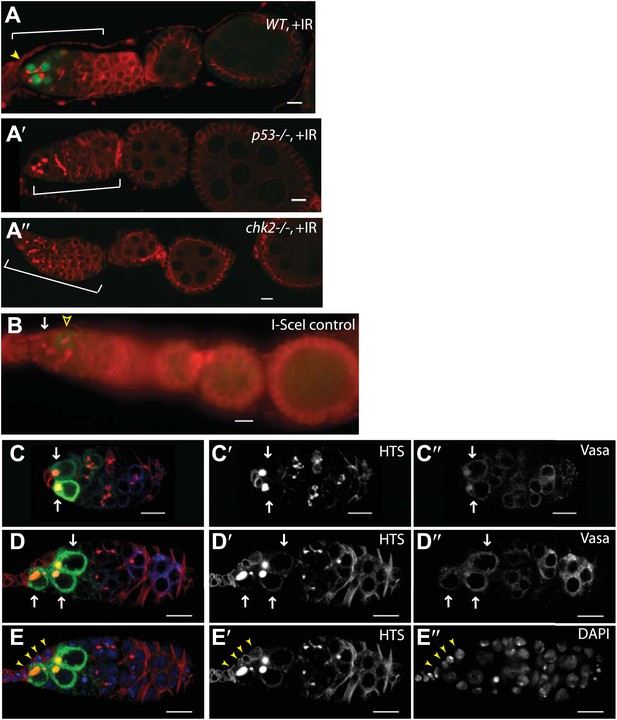
Selective p53 action in germline stem cells is detected using a p53 biosensor.
Irradiated ovaries in panels (A–A′′) validate the p53R-GFP biosensor, establishing that wild-type p53 and chk2 are required for GFP activation in the germarium (brackets). (A) p53R-GFPnls is selectively activated in GSC/CBs (arrowhead) upon irradiation. Reporter activation is not seen in p53 mutants (A′ p53R-GFPnls;p53−) or in Chk2 mutants (A′′ Chk2−;p53R-GFPcyt). Panel (B) is a control for Figure 1D showing that p53R-GFPnls activation is not observed in stem cells (white arrow) that express the I-SceI enzyme but lack the I-SceI restriction site. Note that programed meiotic p53 activation is still observed in these ovaries (open arrowhead). See Figure 1E and Figure 1—source data 1 for quantification. In panels (C–D′′) irradiated germarium are stained with α-HTS (Lin et al., 1994) and α-VASA (Hay et al., 1990) to confirm GFP expression in the GSCs/CBs. (C and D) p53R-GFPcyt activation (green in C and D, white arrows in C–D′′) is restricted to the germline, identified by Vasa staining (white in C′′ and D′′). p53R-GFP activation is also restricted to GSCs and CBs identified by round fusomes (HTS staining, red in C and D, white in C′ and D′). Rounded yellow fusomes validate co-incidence of p53R-GFP and rounded HTS. (E–E′′) p53R-GFPcyt activation in GSCs (green in E) was seen in cells that contact the cap cells (yellow arrowheads), which are visualized by location and size of DAPI stained nuclei (blue in E, white in E′′). Note that the sample in C is also shown as the C′ inset in Figure 1C stained with DAPI. Likewise, D is also shown as the C′′ inset in Figure 1C with single channels of α-HTS and DAPI shown in D′ and D′′ respectively. Also, D and E are the same sample but D highlights Vasa positive cells while (E) highlights contact with cap cells. In panels A′′, C, D, and E the p53R-GFPcyt reporter was used and in panels A, A′, and B the p53R-GFPnls reporter was used. All scale bars are 10 μm.
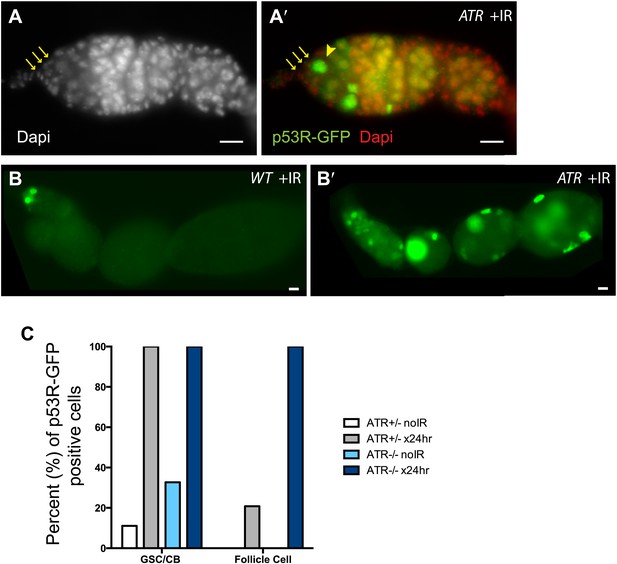
ATR is not rate limiting for p53 activation in the germline.
p53R-GFPnls activation (green) was examined in ATR mutants (A–B′) after irradiation. (A–A′) GSCs (yellow arrowhead) that are in contact with cap cells (yellow arrows in A and A′) identified by DAPI staining (white in A, red in A′). These observations show that ATR is not rate limiting for p53 activation in GSCs. Panels (B–B′) show that induction of the p53R-GFPnls biosensor is not selective in ATR mutants (B′) when compared to WT controls (B). Genotype for (A–A′) is mei-41[D3]/[D3] and for (B′) is mei-41[D3]/[RT]. The p53R-GFPnls biosensor was used for panels A–B′. (C) Quantification of p53R-GFPnls reporter in GSCs/CBs and follicle cells in ATR heterozygous controls and ATR mutants with and without irradiation. Both the ATR+/− control and ATR−/− show a robust induction of p53R-GFPnls in GSCs after irradiation. ATR mutants also show a robust induction of reporter activation in follicle cells after irradiation. Scale bars, 10 μm.
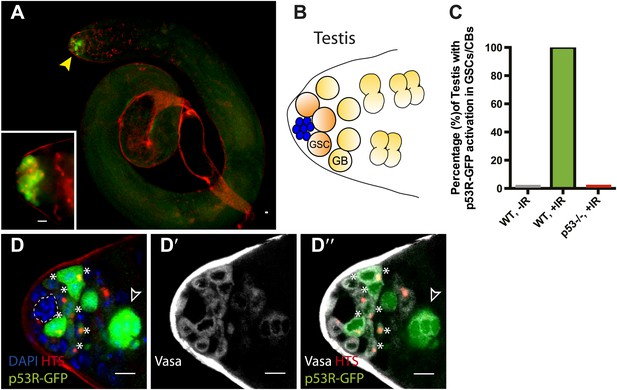
Selective p53 activity occurs in male germline stem cells.
(A) p53R-GFPcyt (green) is induced at the apical tip of an irradiated testis (arrowhead), where stem cells are located (see B). α-HTS co-staining (red) highlights early stages of germline development. The inset in panel (A) shows a higher magnification view from a different irradiated testis. (B) Male GSCs are in contact with cap cells (blue flower pattern) at the apical tip of the testis and divide to produce a gonioblast daugther (GB). (C) Quantifies the percentage of testis activated for the p53 biosensors in GSCs and their immediate progeny. Selective activation is IR (green) dependent and conditional upon p53 since p53R-GFP activation did not occur in p53−/− mutants (red bar). (D–D’’) Confocal images from other irradiated testes confirmed that stem cells induced for p53R-GFPcyt (green, D and D’’) are also positive for rounded HTS staining (red, D and D’’) and the germline specific marker Vasa (white, D’ and D’’) as expected. The hub (dotted line, D) was routinely identified by the characteristic nuclei pattern as illustrated in B (blue cells) and by negative Vasa staining (D’ and D’’). Asterisks mark p53R-GFP positive cells that are adjacent to the hub and Vasa positive or Vasa positive with rounded fusomes. Also note that the hub was identified by α-Armadillo staining (Figure 2—figure supplement 1C). Open arrowhead in (D and D’’) is likely a dying cyst as indicated by pyknotic and condensing nuclei and irregular HTS (Figure 2—figure supplement 1D–E′). In panels A, D–D’’ the p53R-GFPcyt reporter was used. All scale bars represent 10 μm.
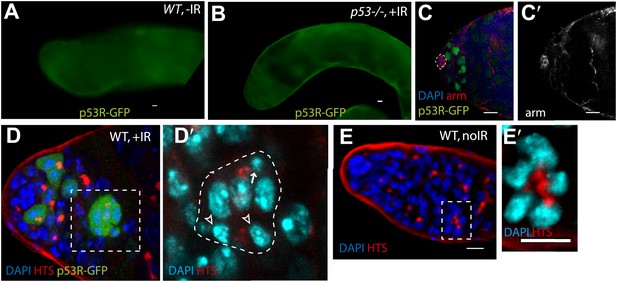
p53 Reporter activation in the male germline (seen in Figure 2A) is conditional upon irradiation (A) and is p53 dependent (B).
Compare A and B to Figure 2A) (C–C′) p53R-GFPnls (green) activation in testis after irradiation. The hub is identified here by α-Armadillo staining (red in H, white in H′), which is noted by a dotted line in (H). (D–D′) Image in (D) is a different z projection from the same irradiated testis shown in Figure 2D, where HTS (red) and DAPI (blue) are used to identify cells. The p53R-GFP positive dying cyst (green) indicated by the open arrowhead in Figure D–D′′′ exhibits pyknotic nuclei (D′ arrow), condensing nuclei (D′ open arrowheads) and irregular HTS. Image in (D′) represents a magnified view of the dashed box in (D). Compare D and D′ to the unirradiated WT testis control in (E–E′) that shows a branched fusome detected by α-HTS (red in E and E′) and nuclei of similar size by DAPI (blue in E and E′). In panels A, B, and E the p53R-GFPcyt reporter was used. In panels C, the p53R-GFPnls reporter was used. All scale bars are 10 μm.
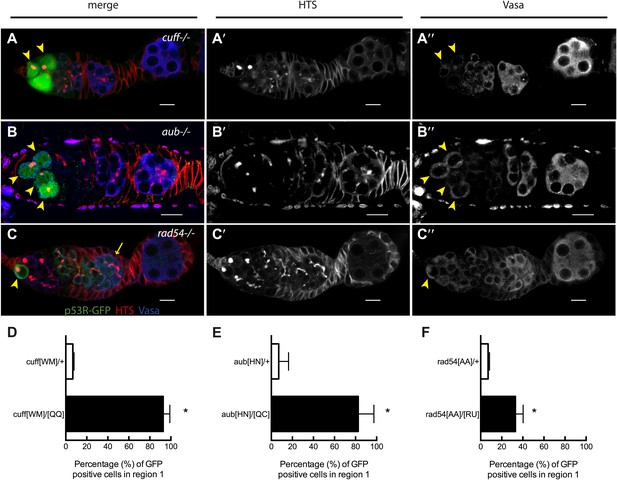
Stem cell associated p53 activity in defective DNA repair and retrotransposon silencing mutants.
(A–B) Activation of the p53 biosensor (green) in the germarium of piRNA mutants, (A) cutoff[QQ}/[WM] and (B) aubergine[HN]/[QC]. (C) Activation of the p53 biosensor in rad54, a meiotic DNA repair mutant. (D–F) Germaria were found to express p53R-GFPcyt in GSCs/CBs with a penetrance of 90% for cutoff mutants (D, p<0.0001), 80% for aubergine mutants (E, p=0.0018), and 33% for rad54 mutants (F, p=0.0039). Asterisks indicate significant differences between heterozygous controls and homozygous mutants. GSCs/CBs were identified by rounded fusomes detected with α-HTS (red in merge A, B, C and white in A’, B’, C’). Arrowheads indicate that p53R-GFP positive cells are also germ cells identified by Vasa staining (blue in A, B, C and white split channel in A’’, B’’, C’’). Note that this particular α-Vasa antibody cross-reacts against the muscle sheath that surrounds each ovariole. If the sheath is not fully dissected and removed, then background staining is evident, as seen in Figure 2B′′. Control genotypes were cuff[WM]/CyO, aub[HN]/CyO, rad54[AA]/CyO. Note that aub and rad54 mutants occasionally showed p53 activation beyond region 2 of the germarium (arrow in C), quantified in Figure 3—figure supplement 1, Figure 3—source data 1. All scale bars represent 10 μm. In panels A, B, and C, the p53R-GFPcyt reporter was used.
-
Figure 3—source data 1
Quantification of p53 activation in defective DNA repair and retrotransposon silencing mutants.
Mutants defective for (A) meiotic repair (rad54 and rad50) and (B) retrotransposon silencing (aubergine and cutoff) have increased spontaneous reporter activation compared to heterozygous controls. The percentage of ovarioles positive for p53R-GFP in the regions indicated (GSC/CB, region 3, stage 2–8 egg chamber) was calculated and the number of ovarioles assayed per region is indicated. Quantification of reporter activation is from three independent trials.
- https://doi.org/10.7554/eLife.01530.011
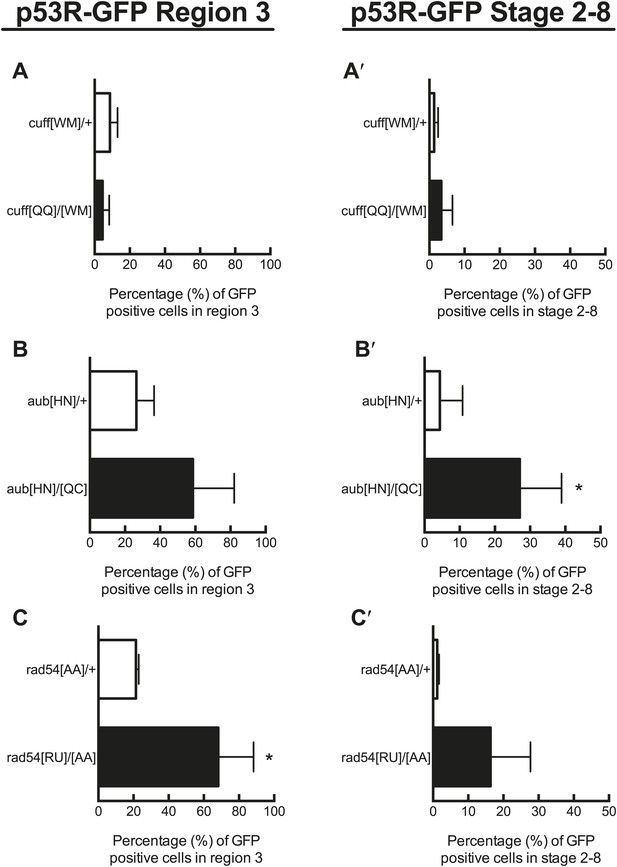
Quantification of p53 activation in defective DNA repair and retrotransposon silencing mutants in region 3 and stage 2–8 egg chambers.
A, B, and C show quantification of reporter activation in region 3 of the germarium. For rad54 mutants, p53 activation was significantly different from the heterozygous control for region 3 (C′, p=0.0157). A′, B′, C′ show quantification of p53 reporter activation in stage 2–8 egg chambers. For aubergine mutants, p53 activation was significantly different from the heterozygous control for stage 2–8 egg chambers (B′′, p=0.0437). Control genotypes were cuff[WM]/CyO, aub[HN]/CyO, rad54[AA]/CyO. See Figure 3—source data 1 for number of ovarioles quantified. Samples shown here are combined from three independent trials. The p53R-GFPcyt reporter was used for these studies.
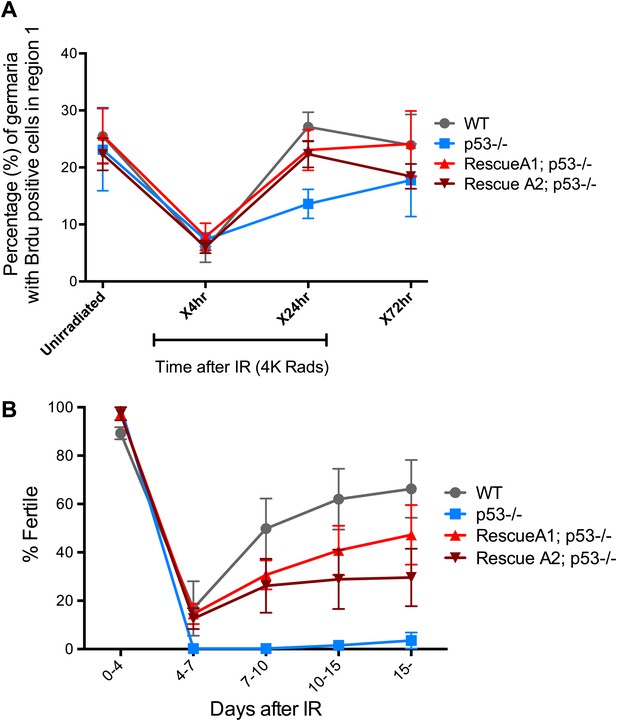
p53 mutants exhibit impaired fertility and delayed re-entry into the cell cycle after irradiation.
(A) BrdU incorporation in GSCs after 4 krad of IR. The percentage of germaria containing BrdU positive GSCs/CBs was plotted on the Y axis. WT and p53−/− GSCs arrest with similar kinetics but p53−/− GSCs were significantly delayed for re-entry into the cell cycle. Error bars represent standard deviation from tests of three independent cohorts. WT and two rescue strains are significantly different from p53−/− at the 0.05 level at the x24 hr time point. Percentages and number of germaria assayed are included in Figure 4—source data 1B. In panels A and B, p53−/− represents animal transheterozygous for two p53 null alleles, p53ns and p53K1. (B) Fertility in wild-type (WT) and p53−/− females was measured after exposure to 11.5 krad of IR (see ‘Materials and methods’), which induces persisting sterility in p53 mutants. WT fertility is significantly different from p53−/− during time points 7–10, 10–15, and 15- at the 0.05 level (see ‘Materials and methods’). Two rescue strains showed partial restoration of fertility. Rescue 1A strain showed restored fertility is significantly different from p53−/− at the 0.05 level at days 10–15 and 15-. Note that after 15 days post irradiaton, fertility was monitored for at least 9 more days as indicated by 15-. Error bars represent standard deviation from five independent trials.
-
Figure 4—source data 1
Quantification of proliferative potential and apoptosis of germaria challenged with irradiation.
(A) Quantification of germaria that have cleaved-caspase3 (CC3) positive GSCs/CBs. Columns indicate the time points after irradiation. Rows indicate the genotype. The percentage of germaria that have CC3 positive GSCs/CBs and the number of germaria is quantified from three independent trials for the x4 hr time point and from two independent trials for the non-irradiated, x2 hr, x8 hr, and x24 hr time points. (B and C) Quantification of germaria that have BrdU positive cells in region 1. The percentage of germaria that have BrdU positive cells in region 1 and the number of germaria assayed are quantified from three independent trials in B and two trials in C per time point.
- https://doi.org/10.7554/eLife.01530.014
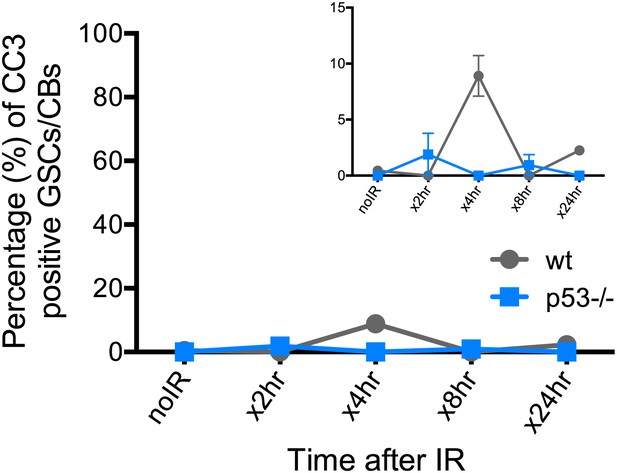
Reporter activation after irradiation does not lead to purging of GSCs through apoptosis.
Time course analysis of stainings for cleaved-caspase3 (CC3) in GSCs/CBs after 4.0 krad of irradiation. The percent of germaria with CC3 positive GSCs/CBs are plotted on the Y axis. The highest incidence of CC3 was only 8% at 4 hr post irradiation in wild-type flies. This is considerably different from the incidence of p53R-GFP positive stem cells after irradiation (∼90%, see Figure 1E). The inset is a magnified view of the same graph to better appreciate the error bars. Error bars represent standard deviations from two trials for no irradiation, 2 hr, 8 hr, 24 hr and three trials for 4 hr for both genotypes. Percentages and number of germaria assayed are provided in Figure 4—source data 1B.
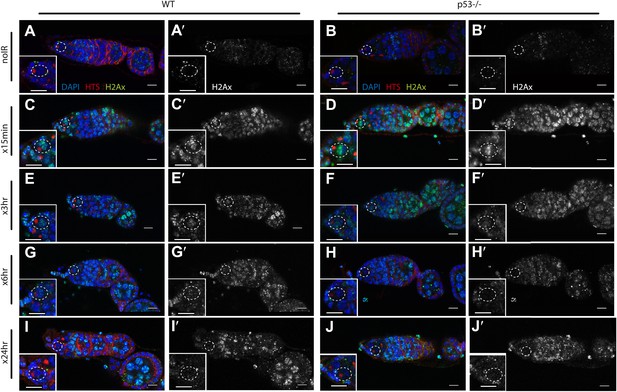
Radiation-induced DNA double-stranded breaks appear and disappear with similar kinetics in WT and p53−/− GSCs.
Time course of α-pH2Av (green) clearance after irradiation of wild-type (A, C, E, G, I) and p53 mutants (B, D, F, H, J) at a dose of 4 krad. Little or no pH2Av staining is observed in unirradiated WT (A) or p53−/− stem cells (B). Similar pH2Av staining is observed in WT and p53−/− stem cells 15 min after irradiation (C and D). In both cases damage was generally cleared from GSCs within 24 hr (I and J). Note that many cells are damaged after irradiation (compare A to G) yet p53 biosensor activation is restricted to GSCs/CBs (Figure 1). White circles indicate stem cells. Insets are magnified views of tip of the germarium from the same image for better GSC visualization. HTS (red) and DAPI (blue) are used to highlight cells in the germarium. Note that the samples in A, A′, C and C′ are included in Figure 1—figure supplement 1 A-B′. Scale bars, 10 μm.
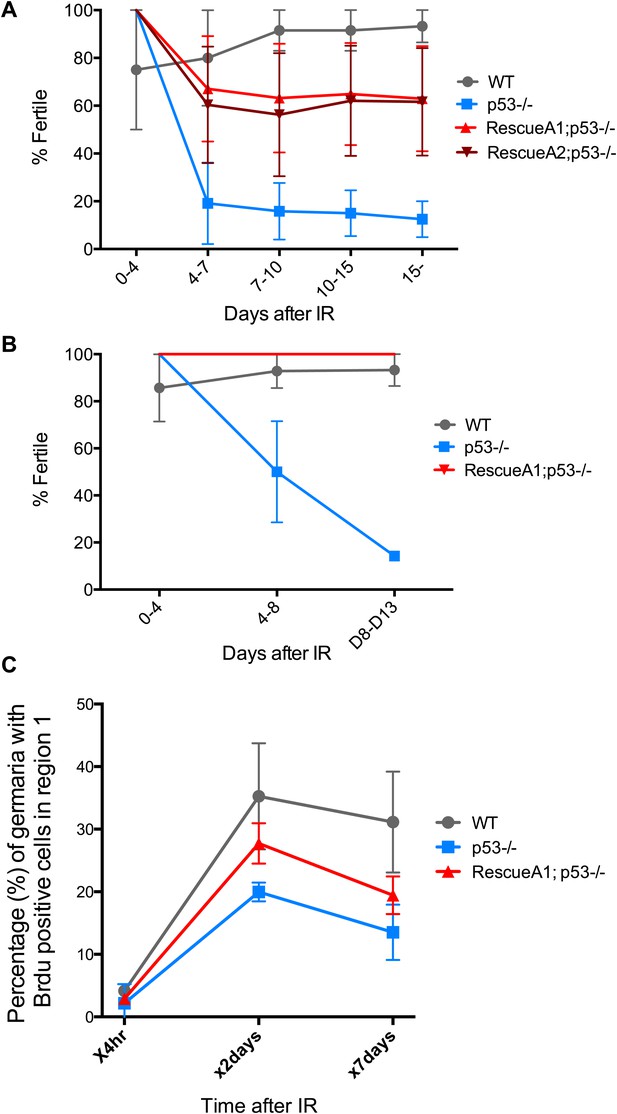
Fertility recovery correlates with proliferation by GSCs and their progeny.
(A) Fertility recovery after 9.0 krad of IR. A similar pattern is observed as seen at higher doses (Figure 4B, 11.5 krad). WT fertility is significantly different from p53−/− at all time points (p<0.05). The A1 Rescue and p53−/− are significantly different at 10–15 and 15- days after IR (p<0.05). Error bars represent standard deviation from four independent trials. Note that fertility was monitored for a total of 25+ days after irradiation as indicated by 15−. To link the fertility defect to cell cycle kinetic differences we observe at lower doses (Figure 4A, 4.0 krad), we performed 2 trials where we assayed fertility (B) and BrdU incorporation in region 1 of the germarium (C) after 9.0 krad of irradiation. Panel (C) shows that GSCs and CBs in p53−/− flies have a reduced proliferation potential at 2 and 7 days post irradiation (Figure 4—source data 1C). Fertility recovery suggests a radiation sensitivity phenotype since p53−/− flies recover fertility in a dose dependent manner (compare to Figure 4B).
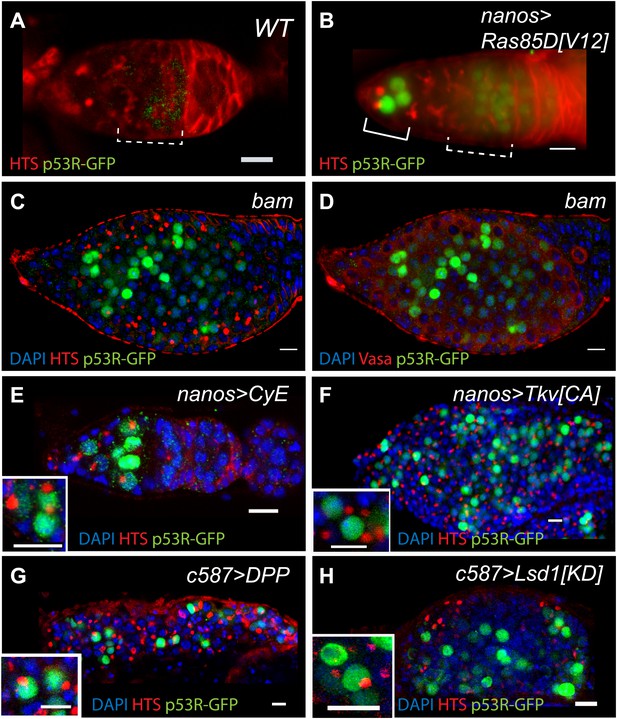
Deregulated growth in the stem cell compartment provokes p53 action.
(A) In an unperturbed wild-type (WT) germarium, the p53R-GFPnls biosensor is absent from GSCs/CBs, marked here by rounded fusomes stained with α-HTS (red). The modest signal in region 2 reflects meiotic p53 activity (dotted bracket) (Lu et al., 2010). When perturbed by RasV12 (B) the p53 biosensor (green) is induced in GSCs/CBs (solid bracket in B, see Figure 5—source data 1). Perturbation from failed differentiation programs caused by the bam mutation (C–D) or Cyclin E over-expression (E) provokes similar p53 biosensor activity. Likewise, increased DPP signaling caused by a constitutively active Tkv receptor (F) or ectopic DPP ligand expression (G) also prompts induction of the p53 reporter. Induction of the p53 reporter is also seen, when the stem cell niche is expanded by silencing of Lsd1 (H) (Eliazer et al., 2011). Insets in panels E–H are magnified views of tumor cysts showing that p53R-GFP positive cells exhibit stem-like properties with rounded fusomes detected by α-HTS co-staining (red). Note in panels B, E and F, the indicated UAS transgenes were expressed using the germline specific driver, nanos-GAL4VP16 (Rorth, 1998). For panels G and H, expression was achieved by the driver c587-GAL4 in somatic cells of the ovariole tip (Song et al., 2004). All images shown are immunostainings for the p53R-GFPnls biosensor (green), HTS (red), and/or DAPI (blue) except for panel D which was co-stained with α-Vasa (red) to show that p53 activated cells retain the germline marker in bam mutants. All other panels (A–C, E–H) were stained with α-HTS (red). Note that panel D stained with α-Vasa is the same bam ovariole shown in C with α-HTS. Relevant quantification including the nanosGAL4 driver alone is shown in Figure 5—source data 1. Scale bars = 10 μm.
-
Figure 5—source data 1
Quantification of biosensor activity in germline tumors.
This table quantifies the number of p53R-GFPnls positive stem-like cells associated with a rounded fusome (α-HTS). Ovaries containing the nanosGAL4 driver alone (control) or the UAS oncogene indicated (Rasv12 or CyclinE) and the same GAL4 driver were scored. Note that the numbers of stem cells activated for p53 is much greater when either Rasv12 or CyclinE are present when compared to the control alone.
- https://doi.org/10.7554/eLife.01530.019
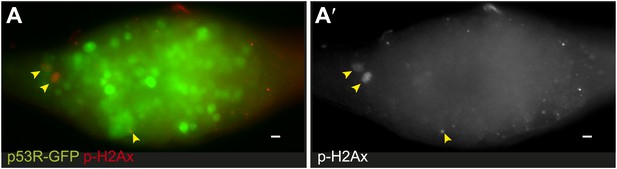
Reporter induction during forced proliferation signals is independent of DNA damage.
(A) Immunostaining for α-pH2Av (red) and p53R-GFPnls (green) in bamΔ86 ovaries. (A′) shows α-pH2Av channel from (A). Note the incidence of pH2Av (arrows) is rare and infrequently colocalizes with GFP.
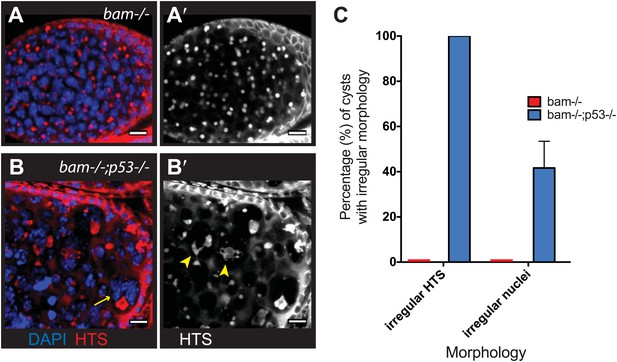
Abnormal fusomes and irregular nuclei are seen in bam−/−p53−/− tumors.
(A–A’) Cells in bam−/− tumors have rounded fusomes normally associated with the undifferentiated GSC fate. These are detected by α-HTS staining (red in B, white in B’). The nuclei of these cells counterstained with DAPI have diameters less than 10 μm (blue in B). (B–B’) bam−/−;p53−/− tumors frequently exhibit disorganized fusomes detected here by α-HTS staining (red in C, white in C’, yellow arrowhead). These tumors also have many fragmented and enlarged nuclei with a diameter significantly greater than 10 μm (blue in C, yellow arrow). (C) Quantification of altered fusome structure and irregular nuclei in bam−/− and bam−/−;p53−/− tumors. Note that in panel C, counts for irregular nuclei do not include micronuclei. A total of 14 cysts were assayed in bam−/−;p53−/− and 8 cysts were assayed for bam−/−. All scale bars, 10 μm.
Tables
p53 status impacts expression profiles in bam−/− tumors
| Downregulated by p53 | Upregulated by 53 | |||
|---|---|---|---|---|
| Gene symbol | Fold change | Gene symbol | Fold change | |
| 1 | CG31681 | 8.7 | CG31809 | −7.2 |
| 2 | CG5156 | 8.0 | CG31810 | −5.6 |
| 3 | LysX | 7.9 | CG2177 | −5.2 |
| 4 | CG31901 | 7.6 | CG7106 | −5.1 |
| 5 | CG16762 | 7.5 | CG1504 | −4.5 |
| 6 | CG32277 | 7.3 | CG15614 | −4.3 |
| 7 | CG17239 | 7.2 | unpg | −4.2 |
| 8 | CG17012 | 7.1 | CG7329 | −4.2 |
| 9 | CG9897 | 7.1 | CG15236 | −4.1 |
| 10 | Ser12 | 6.8 | CG9294 | −4.1 |
| 11 | CG2191 | 6.8 | esg | −3.9 |
| 12 | CG33258 | 6.6 | Ugt36Ba | −3.7 |
| 13 | CG18125 | 6.5 | CG14297 | −3.6 |
| 14 | CG12780 | 6.4 | CG17129 | −3.6 |
| 15 | CG4783 | 6.3 | Cyp6a14 | −3.6 |
| 16 | Cyp6a18 | 6.3 | CG5568 | −3.4 |
| 17 | CG17234 | 6.2 | CG1077 | −3.3 |
| 18 | CG18063 | 6.2 | CG11226 | −3.3 |
| 19 | CG9568 | 6.1 | CG33105 | −3.1 |
| 20 | CG32834 | 6.0 | CG3328 | −3.1 |
-
We performed microarray analysis on bam−/− and bam−/−;p53−/− tumors. The genes that are altered by p53 status in bam−/− tumors are recorded. Listed on the left are the top 20 genes whose abundance is directly or indirectly suppressed by p53. Listed on the right are the top 20 genes whose abundance is directly or indirectly induced by p53. The gene symbol is listed on the left and the fold change in gene expression between bam−/− and bam−/−;p53−/− tumors is listed on the right. Many genes listed here are dramatically affected when p53 is absent.
-
Table 1—source data 1
Expression features of the top 20 genes suppressed by p53.
The top 20 genes that were suppressed by p53 in bam−/−;p53−/− tumors (see Table 1) were examined using GEXC (Seita et al., 2012) to identify enriched pathways. Using this collection we observed a mild enrichment for genes that were absent in embryos or absent in adult somatic tissues relative to all genes in the fly genome.
- https://doi.org/10.7554/eLife.01530.023





