Mitochondrial Rab GAPs govern autophagosome biogenesis during mitophagy
Figures
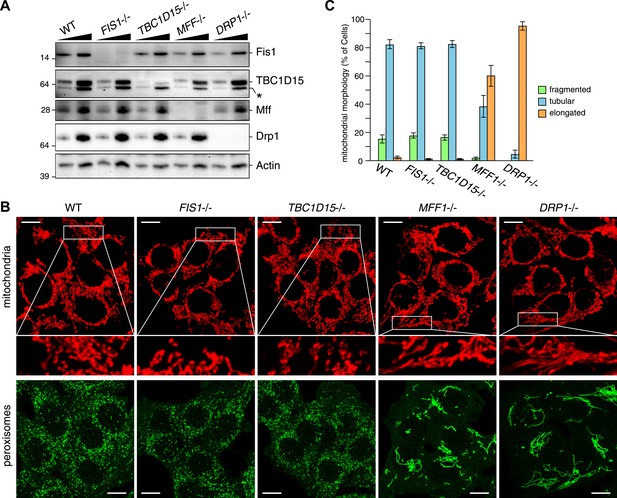
TBC1D15 is dispensable for mitochondria and peroxisome morphologies.
(A) Total cell lysates prepared from the indicated HCT116 cell lines were analyzed by immunoblotting. For comparison, different amounts of proteins (1:3 ratio) were applied. An asterisk indicated non-specific crossreactive bands. (B) The indicated cell lines were analyzed by immunofluorescence microscopy using anti-Cytochrome c antibody for mitochondria and anti-PMP70 antibody for peroxisome staining. Images are displayed as z-stacks of 6 confocal slices. Magnified images are also shown for mitochondrial morphologies. Scale bars, 10 μm. (C) Quantification of mitochondrial morphologies in (B). Percentages of cells harboring fragmented, tubular, or elongated mitochondria are shown. Tubular and elongated denote normal tubular mitochondria seen in WT cells and highly connected mitochondrial network, respectively. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each of three replicate wells.
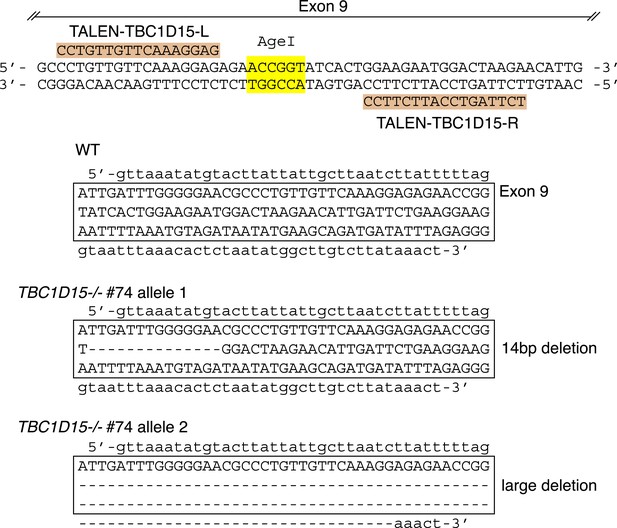
TBC1D15 gene knock out by TALENs.
Highlighted TALEN-TBC1D15-L and TALEN-TBC1D15-R indicate the TALEN binding pair that targets exon 9 of the TBC1D15 gene. DNA sequencing confirmed the nucleotide deletions in both alleles of TBC1D15−/− #74 clone.
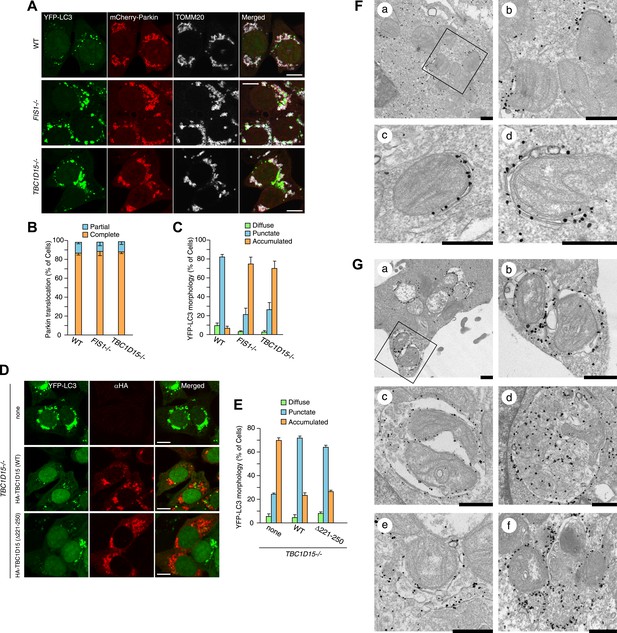
Loss of Fis1 or TBC1D15 causes LC3 accumulation during mitophagy.
(A) The indicated cell lines stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin for 3 hr and subjected to confocal immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 10 μm. (B) Quantification of mCherry-Parkin translocation to mitochondria after 3 hr of valinomycin treatment. Partial or complete translocation to mitochondria in each cell was scored as separate phenotypes. Partial and complete denote that Parkin translocates to some of or all mitochondria, respectively. The error bars represent ±SD from three independent experiments. Over 100 cells were counted in each of three separate wells. (C) YFP-LC3 morphologies in (A) were quantified. Percentages of cells harboring diffuse, punctate or accumulated YFP-LC3 are shown. The error bars represent ±SD from three independent replicates. Over 100 cells were counted in each replicate. For the criteria of LC3 morphology, see Figure 2—figure supplement 1. (D) YFP-LC3 and mCherry-Parkin stably expressing TBC1D15−/− cells in the absence or presence of HA-tagged TBC1D15 WT or Δ221-250 mutant were treated with valinomycin for 3 hr. Cells were subjected to immunofluorescence microscopy with anti-HA antibody. Scale bars, 10 μm. (E) The YFP-LC3 morphology of cells in (D) was quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each well. (F and G) YFP-LC3 and mCherry-Parkin stably expressing WT (F) and TBC1D15−/− (G) cells were treated with valinomycin for 3 hr and then subjected to immunoelectron microscopy with anti-GFP antibody. The square in panel a shows enlarged areas in panel b Scale bars, 500 nm.
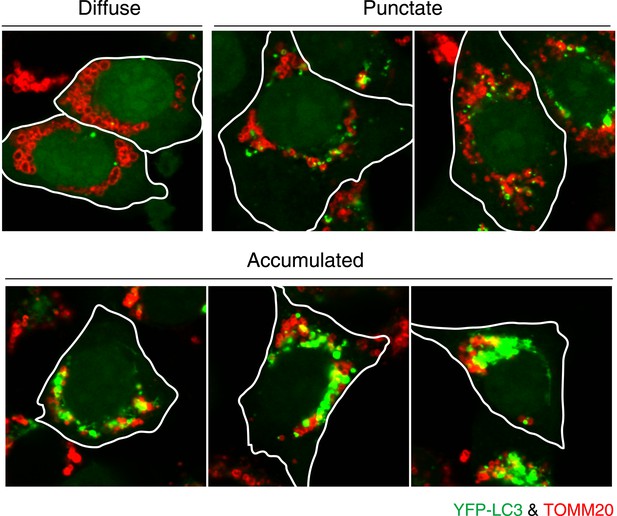
Image examples of different LC3 morphologies.
HCT116 cells stably expressing YFP-LC3 (green) and mCherry-Parkin were treated with valinomycin for 3 hr and then subjected to immunostaining with anti-TOMM20 (red) antibody. Diffuse, YFP-LC3 distributes throughout the cytosol with few dots that are not on mitochondria; Punctate, many discrete YFP-LC3 dots, crescent-shaped, or circular signals on or near mitochondria; Accumulated, highly interconnected YFP-LC3 accumulation or large YFP-LC3 clumps on or near mitochondria.
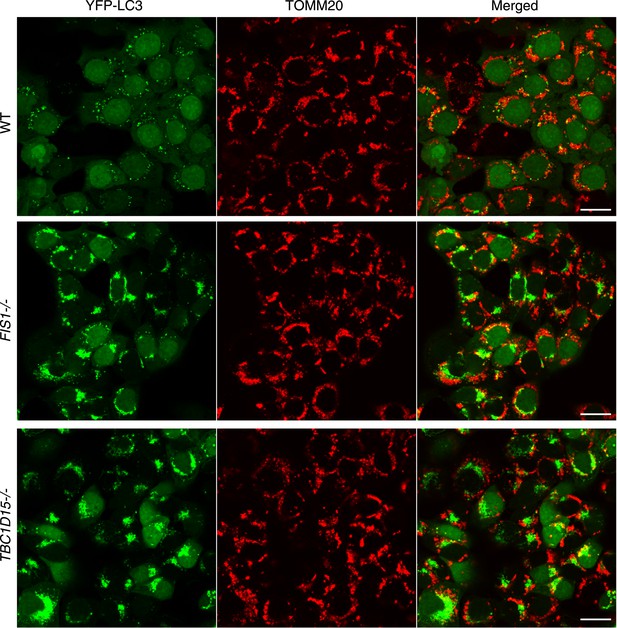
YFP-LC3 accumulation in FIS1−/− and TBC1D15−/− cells during mitophagy.
Low magnification images of cells stably expressing YFP-LC3 and mCherry-Parkin treated with valinomycin for 3 hr. The cells were subjected to immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 20 μm.
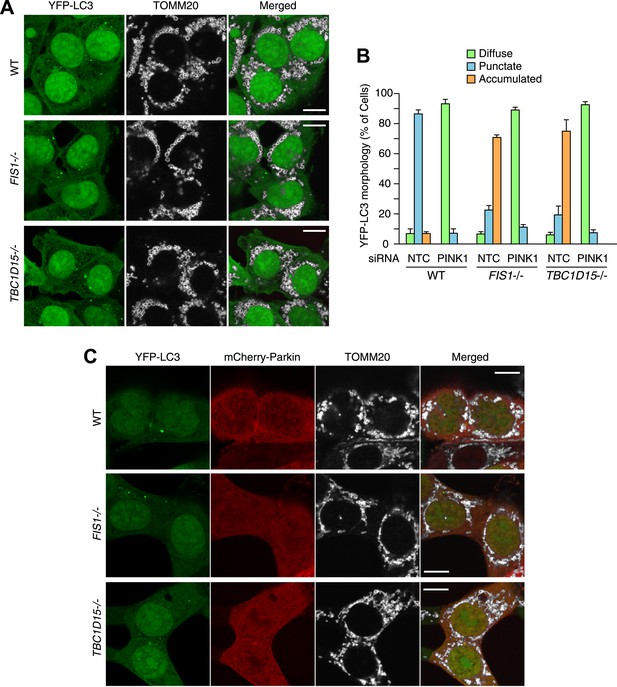
Excessive LC3 accumulation depends on PINK1/Parkin-mediated mitophagy.
(A) The indicated HCT116 cells stably expressing YFP-LC3 (without mCherry-Parkin) were treated with valinomycin for 3 hr followed by immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 10 μm. Of note, HCT116 cells do not express enough endogenous Parkin to induce mitophagy. (B) WT, FIS1−/− and TBC1D15−/− cells stably expressing YFP-LC3 and mCherry-Parkin were treated with control (NTC) or PINK1 siRNA. The percentages of cells harboring diffuse, punctate or accumulated YFP-LC3 after 3 hr of valinomycin treatment were quantified. The error bars represent ±SD from three independent replicates. Over 100 cells were counted in each of three replicates. (C) The indicated cells stably expressing YFP-LC3 and mCherry-Parkin without valinomycin treatment were subjected to immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 10 μm.
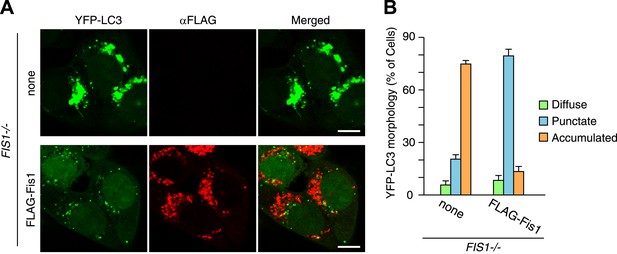
LC3 accumulation in FIS1−/− cells was rescued by Fis1 re-expression.
(A) YFP-LC3 and mCherry-Parkin stably expressing FIS1−/− cells with or without 3xFLAG-Fis1 were treated with valinomycin for 3 hr. Cells were subjected to immunofluorescence microscopy with anti-FLAG antibody. (B) YFP-LC3 morphologies in cells in (A) were quantified. Percentages of cells harboring diffuse, punctate or accumulated YFP-LC3 are shown. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate.
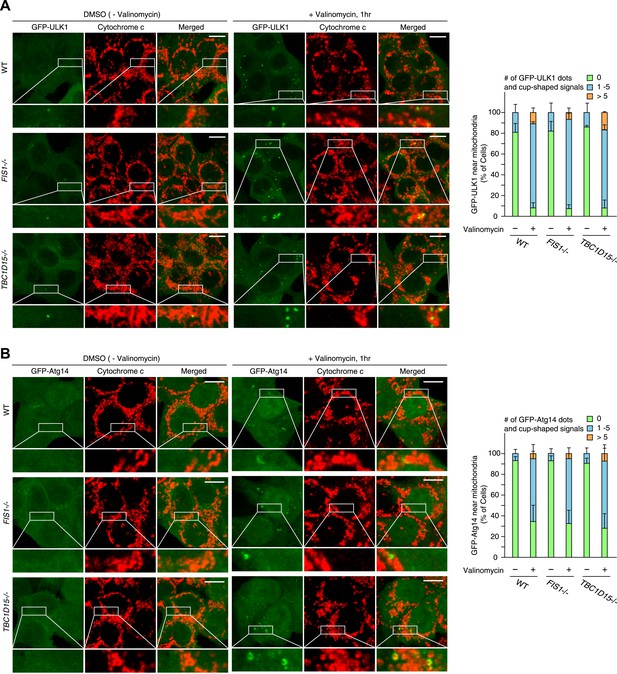
ULK1 and Atg14 recruitment in WT, FIS1−/− and TBC1D15−/− cells during mitophagy.
The indicated cell lines stably expressing GFP-mouseULK1 (A) or GFP-Atg14 (B) and mCherry-Parkin were treated with or without valinomycin for 1 hr and subjected to immunofluorescence microscopy with anti-Cytochrome c antibody. Images are displayed as z-stacks of 6 confocal slices. Magnified images are also shown. Scale bars, 10 μm. The right graph shows the number of GFP-ULK1 or GFP-Atg14 dots and/or cup-shaped signals near mitochondria. The error bars represent ±SD from three replicates. Over 50 cells were counted in each well.
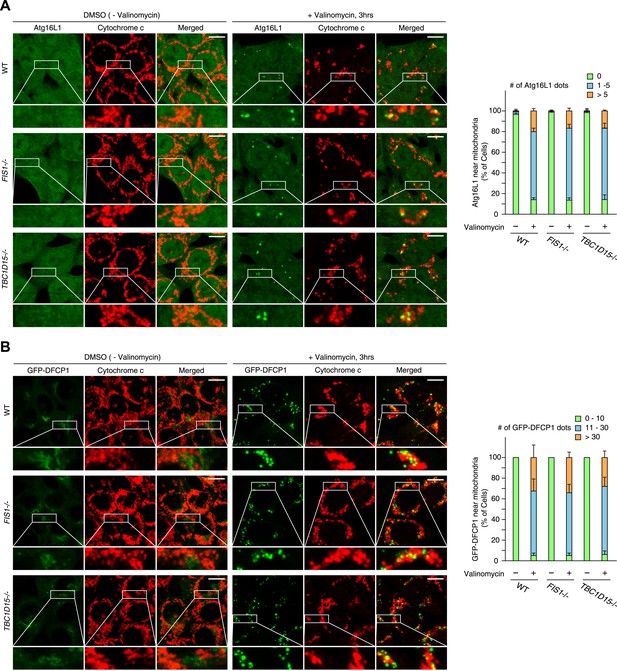
Atg16L1 and DFCP1 recruitment in WT, FIS1−/− and TBC1D15−/− cells during mitophagy.
(A) The indicated cell lines stably expressing mCherry-Parkin were treated with or without valinomycin for 3 hr and subjected to immunofluorescence microscopy with anti-Atg16L1 and anti-Cytochrome c antibodies. (B) The indicated cell lines stably expressing GFP-mouseDFCP1 and mCherry-Parkin were treated with or without valinomycin for 3 hr and subjected to immunofluorescence microscopy with anti-Cytochrome c antibody. Images are displayed as z-stacks of 6 confocal slices. Magnified images are also shown. Scale bars, 10 μm. The right graph shows the number of Atg16L1 or GFP-DFCP1 dots near mitochondria. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate.
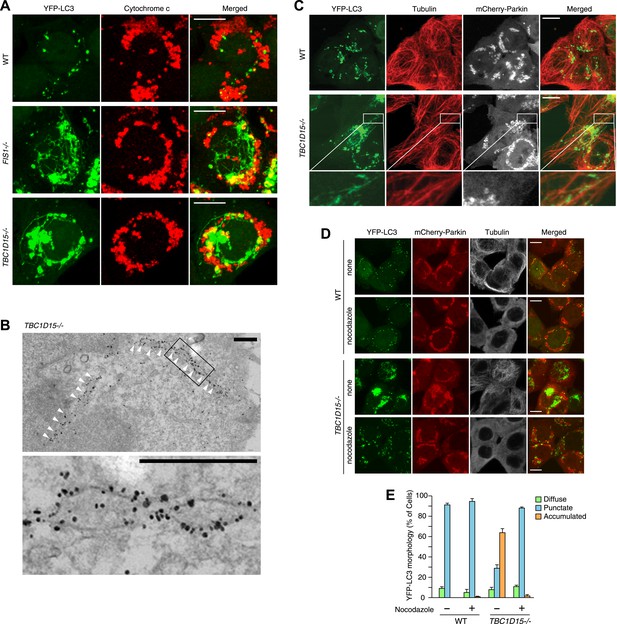
Tubular LC3 expands along microtubules.
(A) The indicated cells stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin for 3 hr followed by immunofluorescence microscopy with anti-Cytochrome c antibody. Confocal images were acquired as z-stacks comprising 6 sequential sections with 0.8 μm z-intervals. Scale bars, 10 μm. (B) TBC1D15−/− cells prepared as in (A) were subjected to immunoelectron microscopy with anti-GFP antibody. White arrowheads indicate YFP-LC3-labeled tubules. High magnification image is shown in the lower panel. Scale bars, 500 nm. (C) The indicated cells prepared as in (A) were subjected to immunostaining with anti-Tubulin antibody. YFP-LC3 and Tubulin staining are merged in the right panels. Magnified images are shown for TBC1D15−/− cells. Scale bars, 10 μm. (D) The indicated cells stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin in the presence or absence of nocodazole for 3 hr. YFP-LC3 and mCherry-Parkin images are merged in the right panels. Scale bars, 10 μm. (E) YFP-LC3 morphologies of cells in (D) were quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each well.
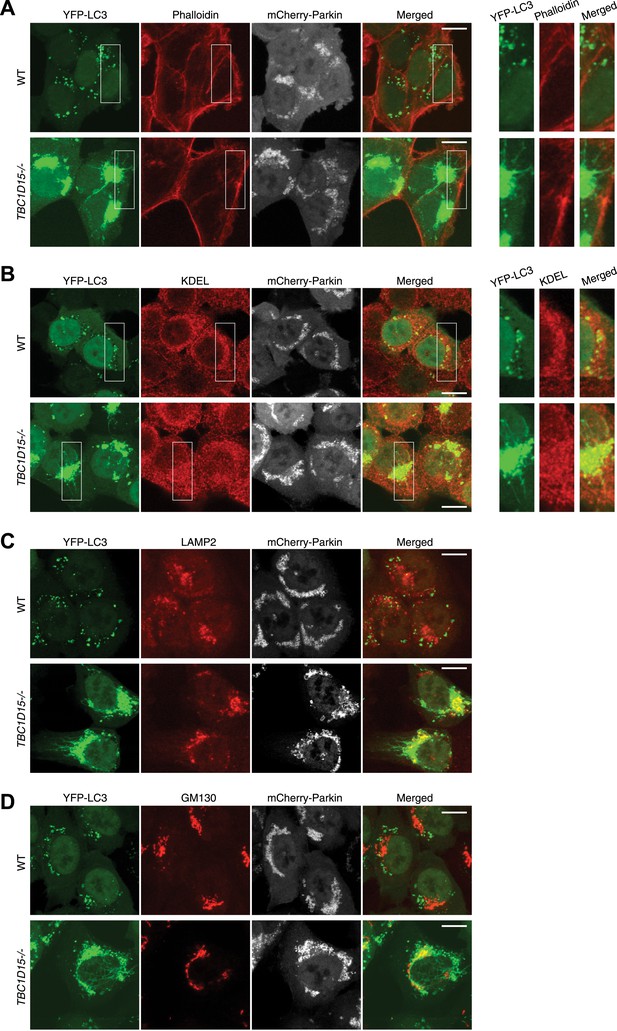
Tubular YFP-LC3 morphologies in TBC1D15−/− cells.
WT and TBC1D15−/− cells stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin for 3 hr. Cells were stained with (A) Alexa Fluor 635 Phalloidin for actin, (B) anti-KDEL antibody for the ER, (C) anti-LAMP2 antibody for lysosomes, and (D) anti-GM130 antibody for Golgi staining. Z-stack confocal images are shown. YFP-LC3 and organelle staining are merged in the right panels. Scale bars, 10 μm. Magnified images are also shown for Actin and ER.
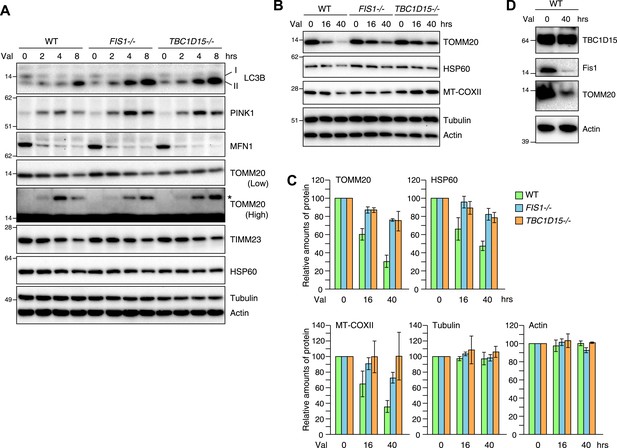
Loss of Fis1 or TBC1D15 impedes clearance of damaged mitochondria.
(A and B) YFP-Parkin stably expressing cells were treated with valinomycin for indicated times. Total cell lysates were subjected to immunoblotting. I and II denote cytosolic and lipidated LC3B, respectively. An asterisk indicates ubiquitinated TOMM20. (C) Indicated protein amounts as in (B) were quantified. The amount of protein without valinomycin treatment was set to 100%. The error bars represent ±SD from three independent experiments. (D) YFP-Parkin stably expressing WT HCT116 cells were treated with or without valinomycin for 40 hr. Total cell lysates were analyzed by immunoblotting.
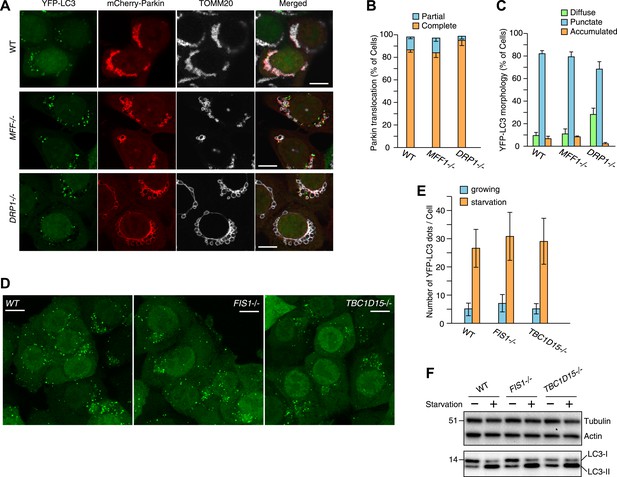
Effect of mitochondrial fission and starvation induction on LC3 accumulation in WT, FIS1−/− and TBC1D15−/− cells.
(A) WT, MFF−/− or DRP1−/− cells stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin for 3 hr and subjected to immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 10 μm. (B) Quantification of mCherry-Parkin translocation to mitochondria after 3 hr of valinomycin treatment. Partial and complete denote that Parkin translocates to some of and all mitochondria, respectively. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate. (C) YFP-LC3 morphologies of cells in (A) were quantified. Percentages of cells harboring diffuse, punctate or accumulated YFP-LC3 are shown. The error bars represent ±SD from three independent replicates. Over 100 cells were counted in each replicate. (D) WT, FIS1−/−, and TBC1D15−/− cells stably expressing YFP-LC3 were grown in starvation media. Z-stacks of confocal images are shown. Scale bars, 20 μm. (E) The number of YFP-LC3 dots in cells under growth or starvation conditions was quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each well. (F) Total cell lysates from cells grown in normal or starvation media for 6 hr were subjected to immunoblotting. LC3-I and LC3-II denote cytosolic and lipidated LC3B, respectively.
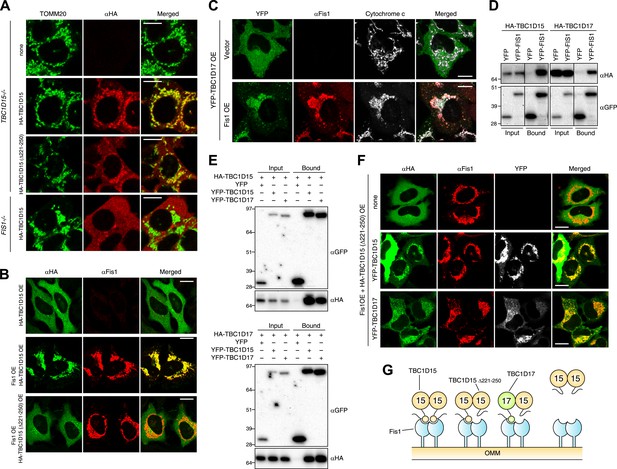
Identification of TBC1D17 as a Fis1 and TBC1D15 binding protein.
(A) TBC1D15−/− cells and those stably expressing HA-TBC1D15 WT and HA-TBC1D15 (Δ221-250), and FIS1−/− cells stably expressing HA-TBC1D15 WT were subjected to immunostaining with anti-TOMM20 and anti-HA antibodies. Scale bars, 10 μm. (B) HA-TBC1D15 WT or HA-TBC1D15 (Δ221-250) with or without Fis1 was transiently overexpressed (OE) in HeLa cells. Cells were subjected to immunostaining with anti-HA and anti-Fis1 antibodies. Scale bars, 20 μm. (C) YFP-TBC1D17 together with pcDNA vector or Fis1 was transiently overexpressed (OE) in HeLa cells. Cells were subjected to immunostaining with anti-Fis1 and anti-Cytochrome c antibodies. Scale bars, 20 μm. (D) YFP or YFP-Fis1 was co-overexpressed with HA-TBC1D15 or HA-TBC1D17 in HEK293 cells. The cell extracts were subjected to pull down assays with GFP-Trap. 5% input and bound fractions were analyzed by immunoblotting with anti-HA (upper panel) and anti-GFP (lower panel) antibodies. (E) YFP, YFP-TBC1D15, or YFP-TBC1D17 was co-overexpressed with HA-TBC1D15 (upper panel) or HA-TBC1D17 (lower panel) in HEK293 cells. The cell extracts were subjected to pull down assays with GFP-Trap. 5% input and bound fractions were analyzed by immunoblotting with anti-GFP and anti-HA antibodies. (F) HA-TBC1D15 (Δ221-250) together with Fis1 and YFP-TBC1D15 WT or YFP-TBC1D17 WT were transiently overexpressed (OE) in HeLa cells. Cells were subjected to immunostaining with anti-HA and anti-Fis1 antibodies. Images of HA and Fis1 staining were merged in the right panels. Scale bars, 20 μm. (G) Schematic model of Fis1, TBC1D15, and TBC1D17 binding. Homo- or hetero-dimer of TBC1D15 can interact with Fis1 dimer on the mitochondrial outer membrane (OMM).

TBC1D17 gene knock out by TALENs.
Highlighted TALEN-TBC1D17-L and TALEN-TBC1D17-R indicate TALEN binding pair that targets the intron–exon 5 junction of TBC1D17 gene. DNA sequencing confirmed nucleotide deletions in both alleles of TBC1D17−/− #74 clone and of TBC1D15/17 DKO #58 clone.
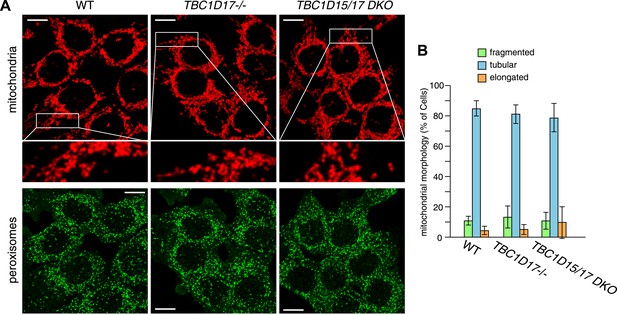
TBC1D17 is dispensable for normal mitochondria and peroxisome morphologies.
(A) The indicated cell lines were analyzed by immunofluorescence microscopy using anti-Cytochrome c antibody for mitochondria and anti-PMP70 antibody for peroxisome staining. Images are displayed as z-stacks of 6 confocal slices. Magnified images are also shown for mitochondrial morphologies. Scale bars, 10 μm. (B) Quantification of mitochondrial morphologies in (A). Percentages of cells harboring fragmented, tubular or elongated mitochondria were shown. Tubular and elongated denote normal tubular mitochondria seen in WT cells and highly connected mitochondrial network, respectively. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate.
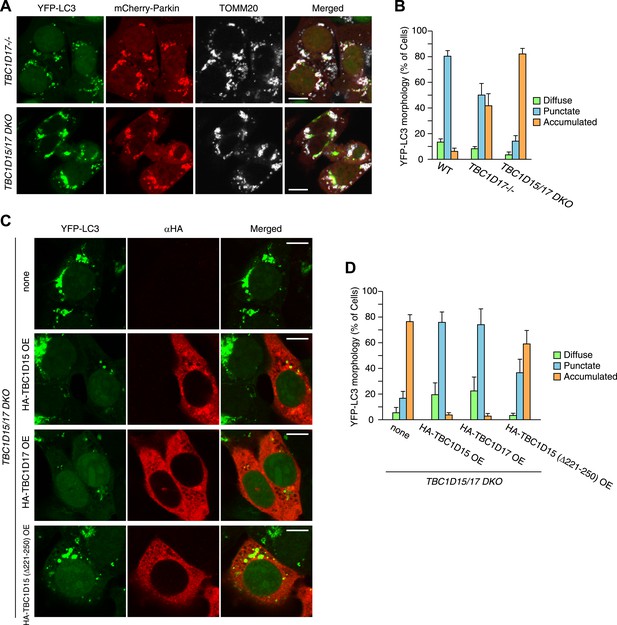
YFP-LC3 accumulation in TBC1D17−/− and TBC1D15/17 DKO cells during mitophagy.
(A) TBC1D17−/− and TBC1D15/17 DKO cells stably expressing YFP-LC3 and mCherry-Parkin were treated with valinomycin for 3 hr and subjected to immunofluorescence microscopy with anti-TOMM20 antibody. Scale bars, 10 μm. (B) YFP-LC3 morphologies in (A) were quantified. Percentages of cells harboring diffuse, punctate or accumulated YFP-LC3 are shown. The error bars represent ±SD from three independent replicates. Over 100 cells were counted in each well. (C) The indicated proteins were transiently overexpressed (OE) in TBC1D15/17 DKO cells stably expressing YFP-LC3 and mCherry-Parkin. The cells were treated with valinomycin for 3 hr followed by immunofluorescence microscopy with anti-HA antibody. Scale bars, 10 μm. (D) YFP-LC3 morphologies in (C) were quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each well.
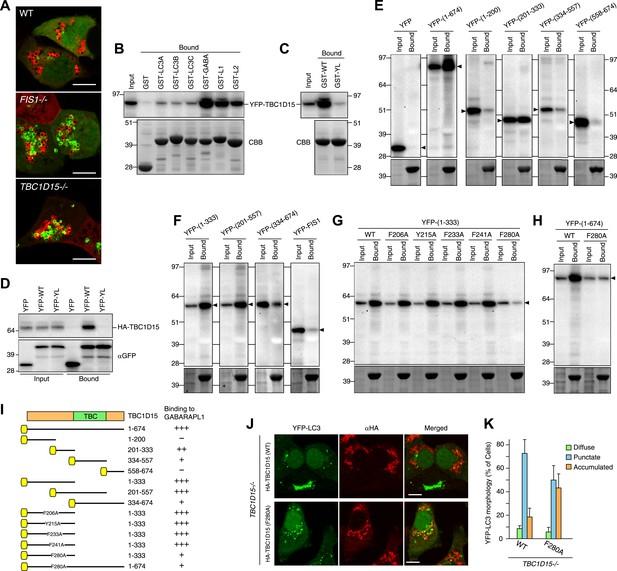
TBC1D15 binds ATG8 family proteins.
(A) The indicated cells transiently expressing YFP-GABARAPL1 (green) and mCherry-Parkin (red) were treated with valinomycin for 3 hr. Scale bars, 10 μm. (B) YFP-TBC1D15 overexpressed in HEK293 cells was subjected to binding assays with GST-fused proteins (GABA, L1, and L2 represent GABARAP, GABARAPL1, and GABARAPL2, respectively). 5% input and bound fractions were analyzed by immunoblotting with anti-GFP antibody (upper panel). Coomassie brilliant blue (CBB) staining shows GST-fusion proteins in bound fractions (lower panel). (C) Binding assay carried out as in (B) with GST-GABARAPL1 WT (GST-WT) or its Y49A/L50A mutant (GST-YL). Immunoblotting with anti-GFP antibody (upper panel) and CBB staining (lower panel) are shown. (D) Cell extracts from HEK293 overexpressed HA-TBC1D15 and YFP, YFP-GABARAPL1 (YFP-WT), or its Y49AL50A mutant (YFP-YL) were subjected to pull down assays with GFP-Trap. 5% input and bound fractions were analyzed by immunoblotting with anti-HA (upper panel) and anti-GFP (lower panel) antibodies. (E–H) The indicated YFP-tagged TBC1D15 full-length, truncated, or point-mutant protein or YFP-Fis1 overexpressed in HEK293 cells were subjected to binding assays with recombinant GST-GABARAPL1. 5% input and bound fractions were analyzed by immunoblotting with anti-GFP antibody (upper panel) and CBB staining (lower panel). (I) Summary of binding abilities of truncated or point-mutated TBC1D15 constructs. –, +, ++, and +++ indicates binding of recombinant GST-GABARAPL1 to less than 1%, 1–5%, 5–10%, and over 10%, respectively of the total YFP-TBC1D15 fragment. Yellow boxes indicate YFP tags. (J) YFP-LC3 and mCherry-Parkin stably expressing TBC1D15−/− cells in the presence of HA-tagged TBC1D15 WT or F280A mutant were treated with valinomycin for 3 hr. Cells were subjected to immunofluorescence microscopy with anti-HA antibody. Scale bars, 10 μm. (K) The YFP-LC3 morphology of cells in (J) was quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each well.
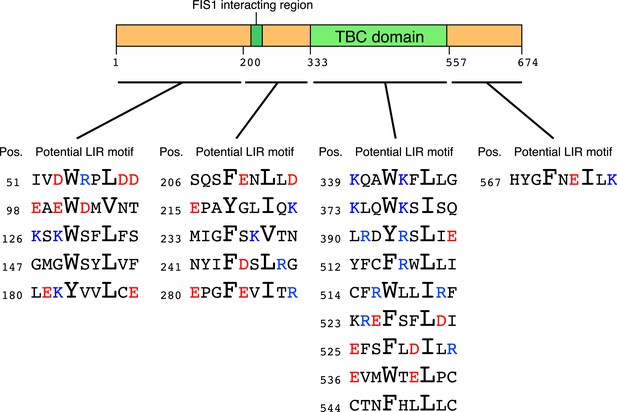
Schematic representation of potential LIR motifs of TBC1D15.
The amino acid position (Pos.) of the conserved aromatic residue in the potential LIR motifs in human TBC1D15 is indicated. Red, acidic residues; Blue, basic residues.
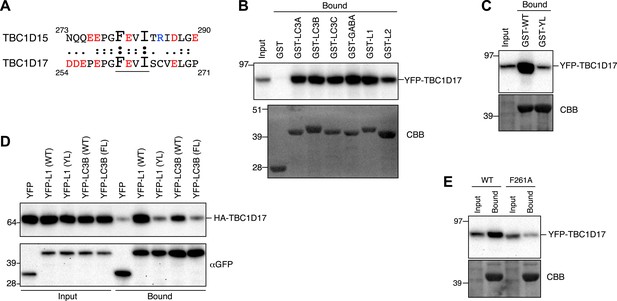
TBC1D17 has the ability to bind Atg8 homologues.
(A) Protein sequence comparison of the LIR motif in TBC1D15 with that of TBC1D17. Underlined sequence denotes consensus LIR motif. Red, acidic residues; Blue, basic residues. (B) Overexpressed YFP-TBC1D17 was subjected to binding assays with GST-fused Atg8 proteins (GABA, L1, and L2 represent GABARAP, GABARAPL1, and GABARAPL2, respectively). 5% input and bound fractions were analyzed by immunoblotting with anti-GFP antibody (upper panel) and coomassie brilliant blue (CBB) staining (lower panel). (C) Binding assays of YFP-TBC1D17 were carried out with GST-fused GABARAPL1 WT (GST-WT) or its Y49AL50A mutant (GST-YL). Immunoblotting with anti-GFP antibody (upper panel) and CBB staining (lower panel) are shown. (D) Extracts from cells overexpressing both HA-TBC1D17 and YFP, YFP-GABARAPL1 (YFP-L1), its Y49AL50A mutant (YL), YFP-LC3B or its F52A/L53A mutant (FL) were subjected to pull down assays with GFP-Trap. 5% input and bound fractions were analyzed by immunoblotting with anti-HA (upper panel) and anti-GFP (lower panel) antibodies. (E) YFP-TBC1D17 WT and F261A mutant overexpressed in HEK293 cells were subjected to a binding assay with recombinant GST-LC3B. 5% input and bound fractions were analyzed by immunoblotting with anti-GFP antibody (upper panel) and CBB staining (lower panel).
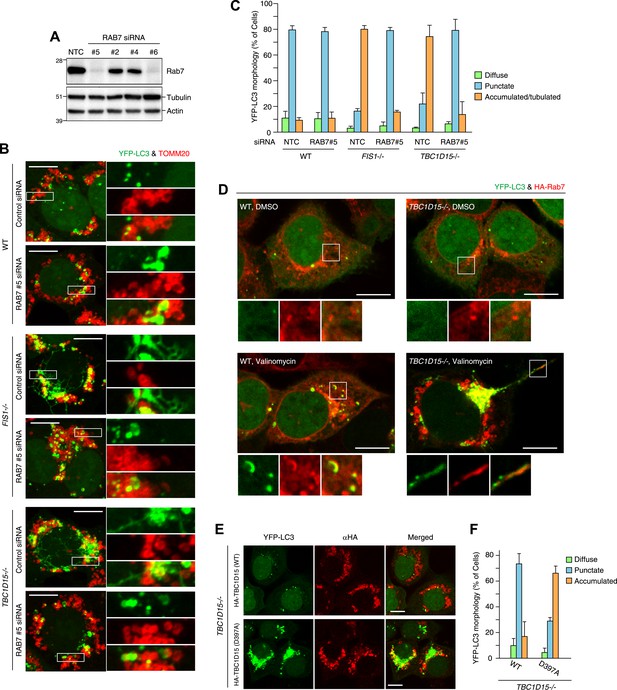
Rab7 is involved in autophagosome fusion during mitophagy.
(A) Total cell lysates from HCT116 treated with control (NTC) or RAB7A siRNA were analyzed by immunoblotting. (B) The indicated cells stably expressing YFP-LC3 (green) and mCherry-Parkin were treated with control (NTC) or Rab7_#5 siRNA. After 3 hr valinomycin treatment, cells were analyzed by immunofluorescence microscopy with anti-TOMM20 antibody (red). Z-stacks of confocal images are shown. Magnified images are also shown. Scale bars, 10 μm. (C) YFP-LC3 morphologies of cells in (B) were quantified. Percentages of cells harboring diffuse, punctuate or accumulated/tubulated YFP-LC3 are shown. Data and error bars were obtained from at least 50 cells in each of three independent replicates. (D) The indicated cells stably expressing YFP-LC3 (green), mCherry-Parkin, and 2HA-Rab7 (Red) were treated with or without valinomycin for 3 hr and analyzed by immunofluorescence microscopy with anti-HA antibody. Magnified images are also shown. Scale bars, 10 μm. (E) YFP-LC3 and mCherry-Parkin stably expressing TBC1D15−/− cells in the presence of HA-tagged TBC1D15 WT or the D397A mutant were treated with valinomycin for 3 hr. Cells were subjected to immunofluorescence microscopy with anti-HA antibody. Scale bars, 10 μm. (F) The YFP-LC3 morphology of cells in (E) was quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate.
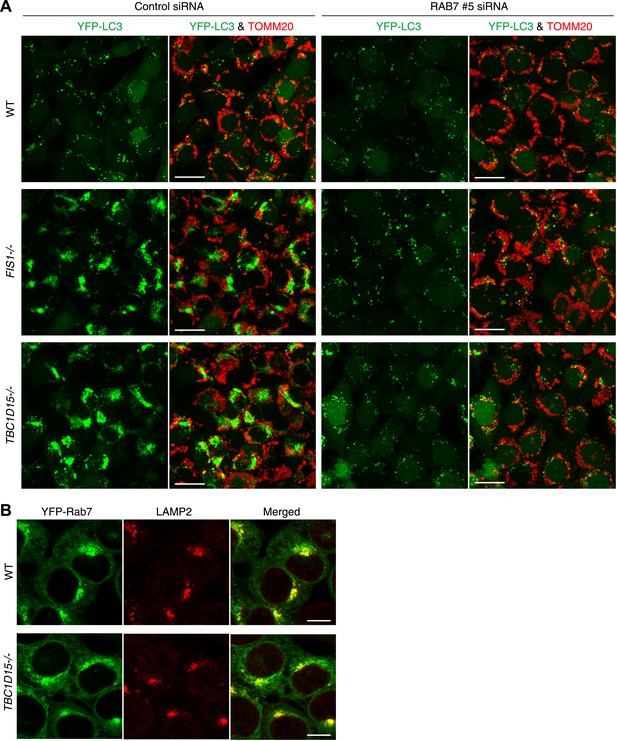
Rab7 is involved in LC3 accumulation of FIS1−/− and TBC1D15−/− cells.
(A) Low magnification images of cells stably expressing YFP-LC3 and mCherry-Parkin treated with valinomycin for 3 hr. Cells were subjected to immunofluorescence microscopy with anti-TOMM20 antibody. Z-stacks of confocal images are shown. Scale bars, 20 μm. (B) The indicated cells stably expressing YFP-Rab7 and mCherry-Parkin were analyzed by immunostaining with anti-GFP and anti-LAMP2 antibodies. Scale bars, 10 μm.
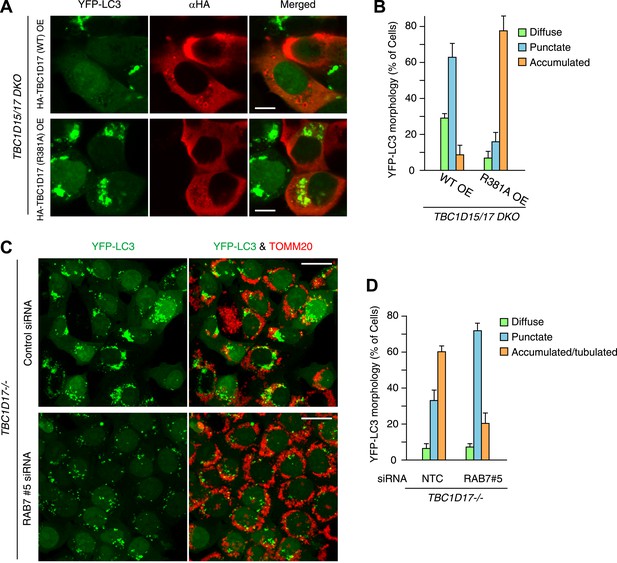
TBC1D17 GAP activity is important for LC3 accumulation through Rab7.
(A) HA-tagged TBC1D17 WT or the R381A mutant was transiently overexpressed (OE) in TBC1D15/17 DKO cells stably expressing YFP-LC3 and mCherry-Parkin. The cells treated with valinomycin for 3 hr were subjected to immunofluorescence microscopy with anti-HA antibody. Scale bars, 10 μm. (B) The YFP-LC3 morphology of cells in (A) was quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each replicate. (C) TBC1D17−/− cells stably expressing YFP-LC3 (green) and mCherry-Parkin were treated with control (NTC) and Rab7_#5 siRNAs for 48 hr. The cells treated with valinomycin for 3 hr were then subjected to immunofluorescence microscopy with anti-TOMM20 antibody (red). Z-stacks of confocal images are shown. Scale bars, 20 μm. (D) The YFP-LC3 morphology of cells in (C) was quantified. The percentages of cells harboring diffuse, punctate or accumulated/tubulated YFP-LC3 were quantified. The error bars represent ±SD from three independent replicates. Over 50 cells were counted in each of three replicate.
Videos
Tubular LC3 movement in TBC1D15−/− cells.
TBC1D15−/− cells stably expressing YFP-LC3 (green) and mCherry-Parkin (red) were treated with valinomycin for 3 hr and then took z-stack pictures.





