Designed α-sheet peptides inhibit amyloid formation by targeting toxic oligomers
Figures
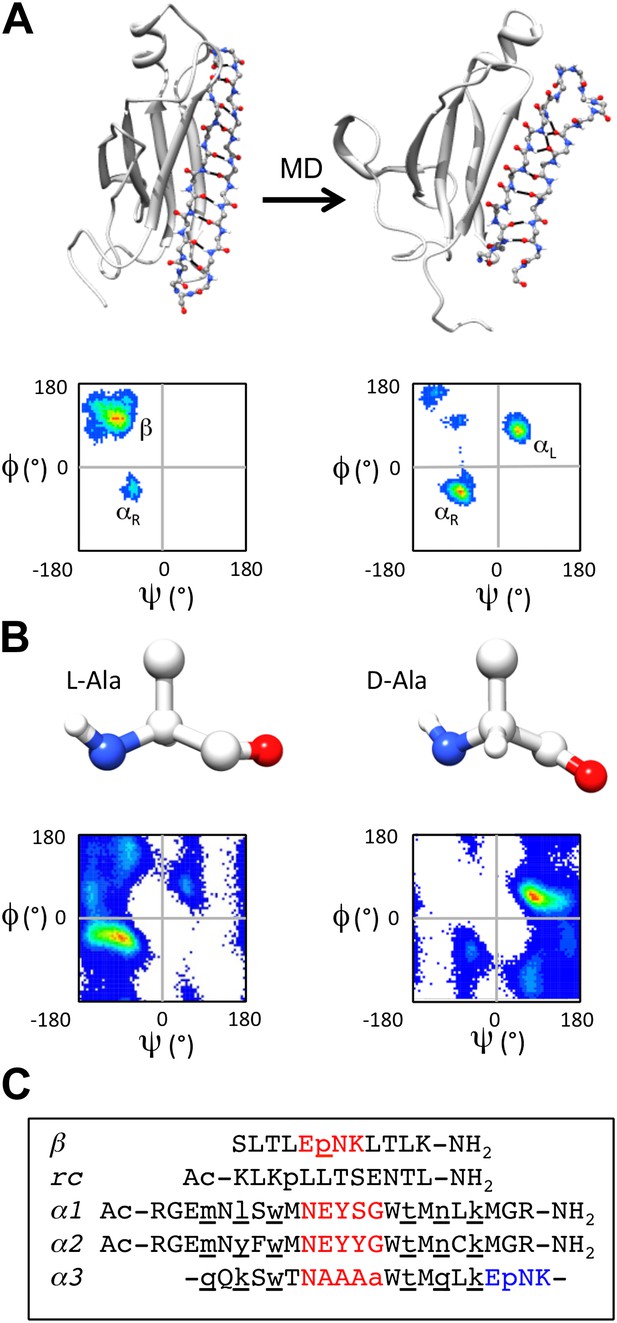
α-sheet conversion, conformational properties and peptide designs.
(A) β- to α-sheet conversion of transthyretin (as reported by Armen et al., 2004a, 2004b). The protein backbone is shown in cartoon representation with the region of interest (residues 105–121) shown as sticks. At 0 ns (top left) the residues of interest form a β-hairpin. The dihedral angles for 1 ns of dynamics of these residues are found mainly in the β-region of the Ramachandran plot (top left quadrant, lower left panel; increasing frequency of occupancy is shown from blue through red) with several turn residues in the αR conformation (bottom left quadrant). After 30 ns (top right) the β-sheet has converted to an α-sheet. The dihedral angles for 1 ns of dynamics of the same residues reveal that the majority of residues have moved from the β-region to the αL (top right quadrant) or αR region of the Ramachandran plot (lower right panel). (B) Intrinsic residue propensities for L- and D-alanine were calculated from 100 ns of MD simulations of a GGXGG peptide system (Beck et al., 2008) (D-alanine was simulated using the same protocol). The backbone structure (upper panels) as well as the Ramachandran plot of the conformation of the alanine residue during the entire simulation, demonstrate the conformational preference for L-alanine to adopt the αR conformation and for D-alanine to favor the αL conformation (lower panels). (C) Peptide designs reported in this study. All designs are single turn hairpins, with the exception of α3, which contains a cyclic peptide backbone resulting in two turns. Hairpin peptides are N- and C-terminally acetylated and amidated, respectively, except for β, which had a free N-terminus. D-amino acids are denoted by lower case and are underlined, and turn residues are colored red in the linear peptides and red and blue in the cyclic design.
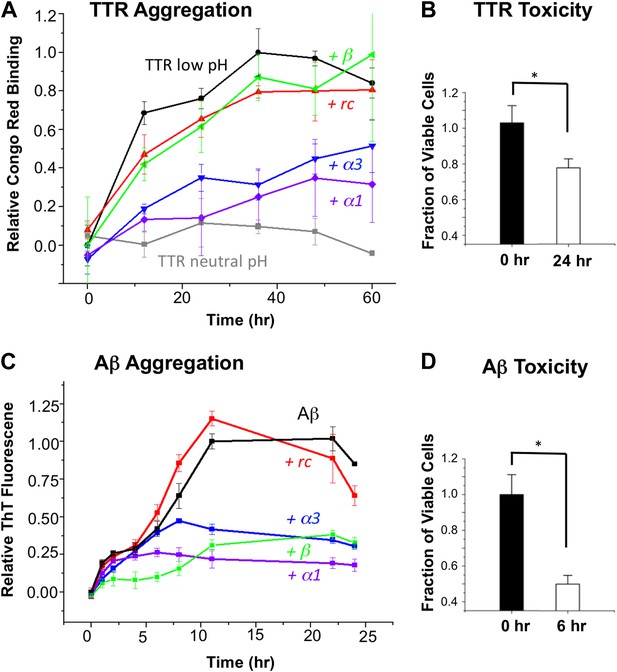
α-Sheet designs inhibit amyloid formation and selectively bind toxic species.
(A) Peptide designs (800 μM) were co-incubated at pH 4.5 with 40 μΜ TTR (monomer) at 37°C and aggregation was monitored by Congo red binding. Error bars indicate the standard deviation. (B) Toxicity of the TTR solution after 24 hr pre-incubation at pH 4.5 against the human neuroblastoma cell line SH-SY5Y in a MTT metabolic viability assay. (C) ThT monitoring of 10 μM Aβ aggregation and inhibitory effects of 100 μM peptides present from the beginning of the aggregation at 37°C. Inhibition values were taken at 12 hr due the decay in ThT fluorescence, particularly for uninhibited samples, which has been described elsewhere (Yamin et al., 2009). (D) Aβ toxicity after 6 hr of aggregation, as probed using the MTT assay and the SH-SY5Y cell line. All data represent average ± SD (* indicates p<0.05, determined using Student's t test).
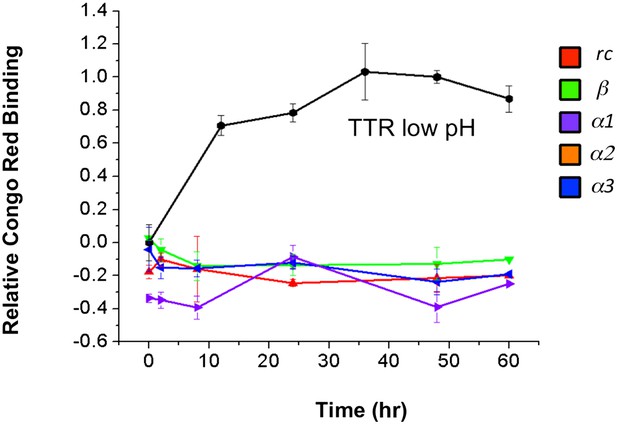
Increase in Congo red binding is not due to peptide aggregation.
Congo red binding of peptides incubated under identical aggregation conditions reveals that contributions to dye binding do not come from the designs.
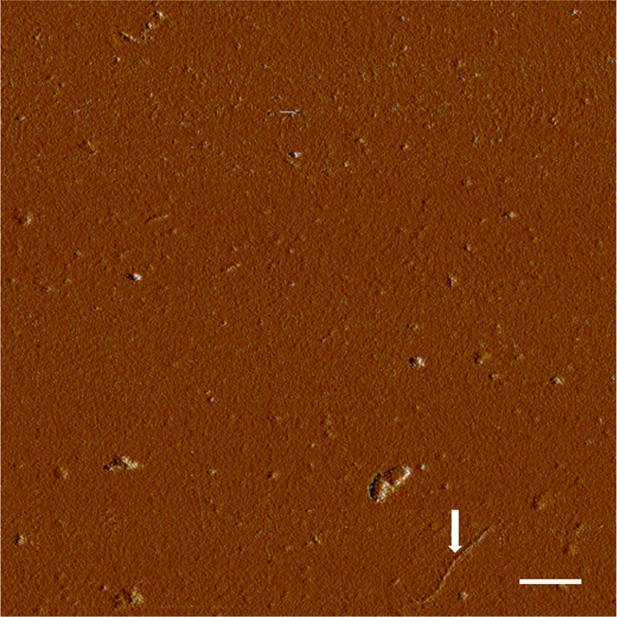

AFM spectroscopy reveals aggregation conditions ultimately result in fibrils.
Congo red binds multiple soluble species in addition to fibrils, so presence of fibrils was confirmed visually. Application of a TTR solution aggregated for 72 hr revealed the presence of small aggregates and fibrils. Fibrils are indicated with an arrow. Scale = 200 nm.
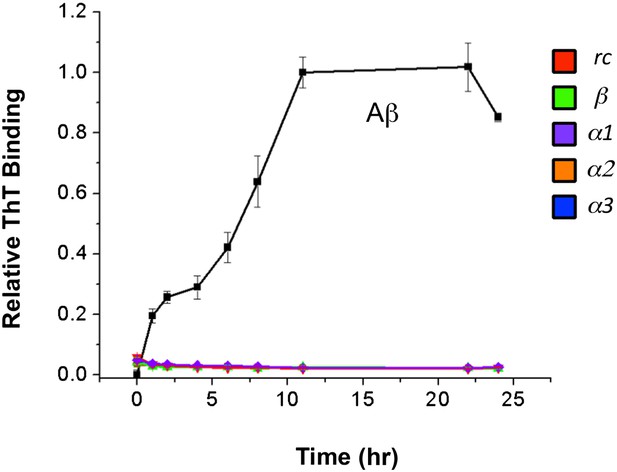
Increase in ThT fluorescence is not due to peptide aggregation.
ThT fluorescence of peptides incubated under identical aggregation conditions indicate that the peptides do not contribute to the observed increase in fluorescence.
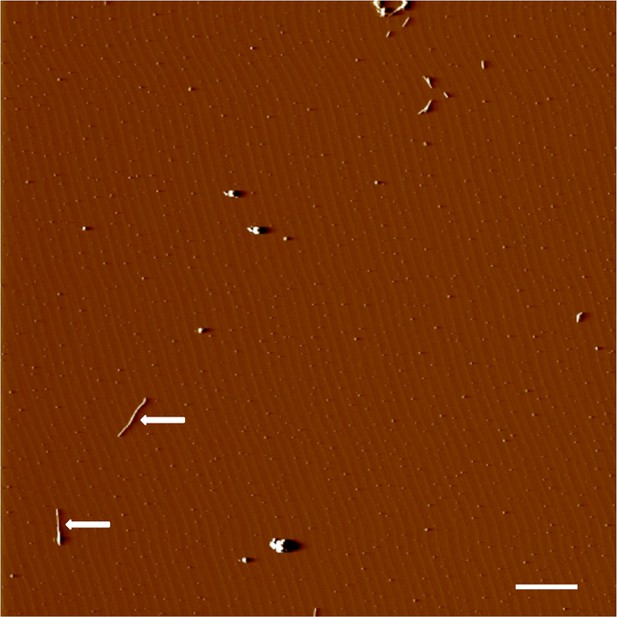

AFM spectroscopy confirms an increase in fibrillar products.
Application of an aggregated Aβ solution reveals the presences of aggregates and fibrils. Fibrils are indicated with arrows. Scale = 200 nm.
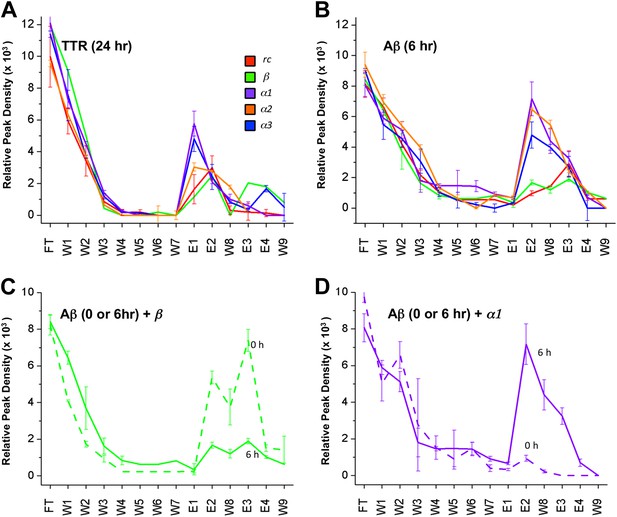
Immobilized designs bind toxic soluble oligomer from solution.
Peptide designs were immobilized onto agarose beads and their ability to bind TTR or Aβ from solution at various stages of aggregation was probed using dot blot analysis. The presence of TTR or Aβ in the initial flow through (FT), sequential buffer washes (W), and sequential guanidine hydrochloride elution steps (E1–E2 and E3–E4) (x-axes) was detected by the integrated peak density of the dot blot analysis (y-axes). E1–W9 are within the linear range of the immunochemistry. (A) All three α-sheet designs, α1, α2 and α3 more strongly bound species from the 24 hr pre-aggregated, toxic TTR solutions than did either control. (B) Similar results were observed with the α-sheet designs binding to toxic Aβ solutions pre-aggregated for 6 hr. Despite the inhibitory effects seen with the β design in the Aβ fibrillization assay, little Aβ from a pre-aggregated solution bound to the immobilized β design. (C) Comparison of binding from a fresh (0 h), or pre-aggregated (6 hr), toxic Aβ solution. β is the only design that preferentially bound fresh Aβ over the aggregated toxic form, indicating that the inhibition observed was due to interactions with monomeric Aβ. (D) In contrast to the β control, α1 preferentially bound the pre-aggregated, toxic form of Aβ compared with fresh monomeric Aβ.
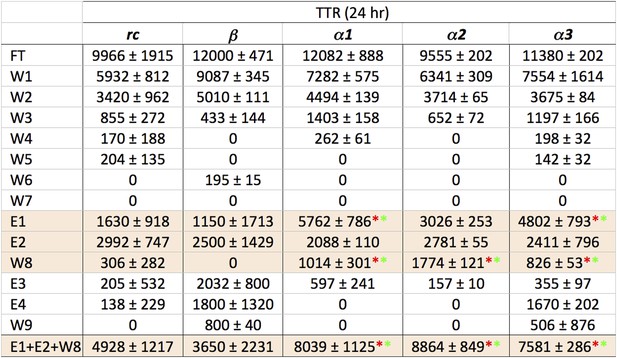
Statistical analysis of data presented in Figure 3A.
Relative peak density values from dot blot analyses are given as average values from three independent experiments, with standard deviations. Statistical analysis for E1, E2 and W8 were performed against the corresponding values for rc and β. t tests that returned a value p≤0.05 are indicated with an asterisk (red when tested against the corresponding value from rc, green when tested against the same value for β) and were considered significant. Values for E1, E2 and W8 were also summed and standard deviations propagated (bottom line), and the t test repeated against the corresponding values for rc and β. These results indicate that α1, α2 and α3 all bind species present in a toxic TTR solution to a greater extent than the random coil or β-sheet control peptides.
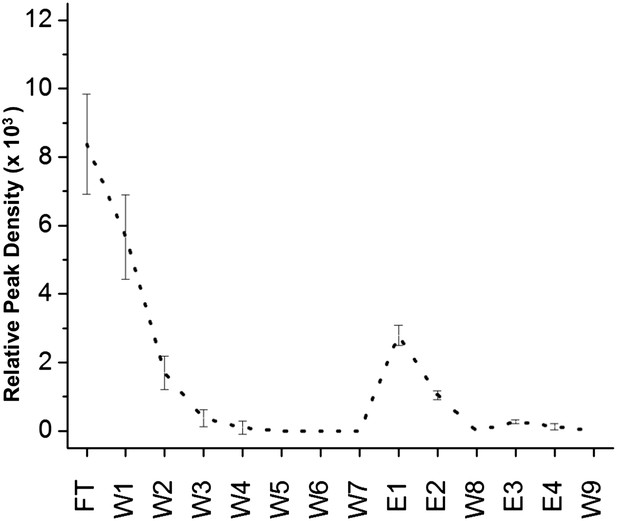
Nonspecific binding of nonnative TTR to column matrix.
While the column-binding assay is not quantitative, nor was it expected to be, we can estimate the confidence in the peak densities by seeing to what extent TTR binds to the column matrix in the absence of coupled peptides. TTR aged for 24 hr was applied to agarose beads blocked only with Tris to establish the effect of the column matrix. As can be seen, the extent of binding to the column matrix (E1 and E2) is comparable to that observed with the rc and β controls. Note also that aggregated TTR elutes earlier from columns blocked with Tris, presumably due to the increased hydrophobicity of TTR as it aggregates and the less favorable interactions with the more hydrophilic Tris matrix.
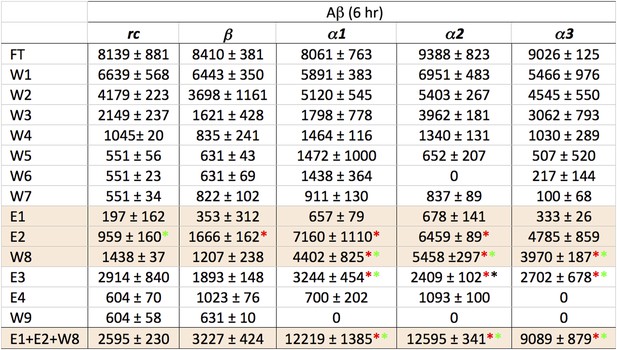
Statistical analysis of data presented in Figure 3B.
Relative peak density values from dot blot analyses are given as average values from three independent experiments, with standard deviations. Statistical analysis for E1, E2 and W8 were performed against the corresponding values for rc and β. t-tests that returned a value p≤0.05 are indicated with an asterisk (red when tested against the corresponding value from rc, green when tested against the same value for β) and were considered significant. Values for E1, E2 and W8 were also summed and standard deviations propagated (bottom line), and the t test repeated against the corresponding values for rc and β. These results indicate that α1, α2 and α3 all bind species present in a toxic Aβ solution to a greater extent than the random coil or β-sheet control peptides.
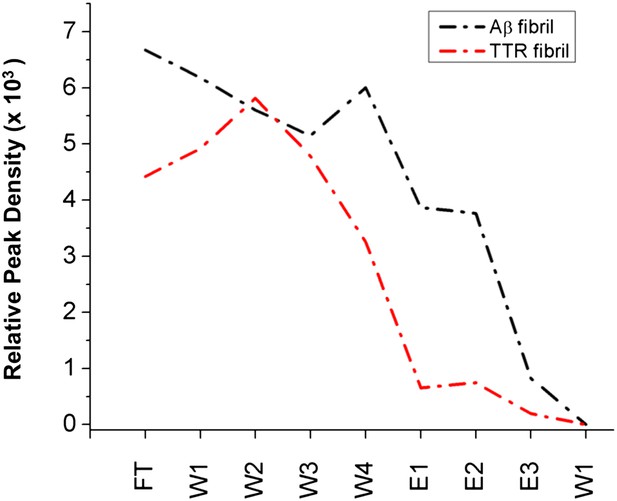
Immobilized α-sheet designs do not bind fibrils.
TTR (red) or Aβ (black) fibrils were applied to agarose beads coated with the α1 design. The elution profile does not show any increase in elution when guanidine is applied to the column, indicating that there are no specific interactions between the fibrils and the immobilized designs.
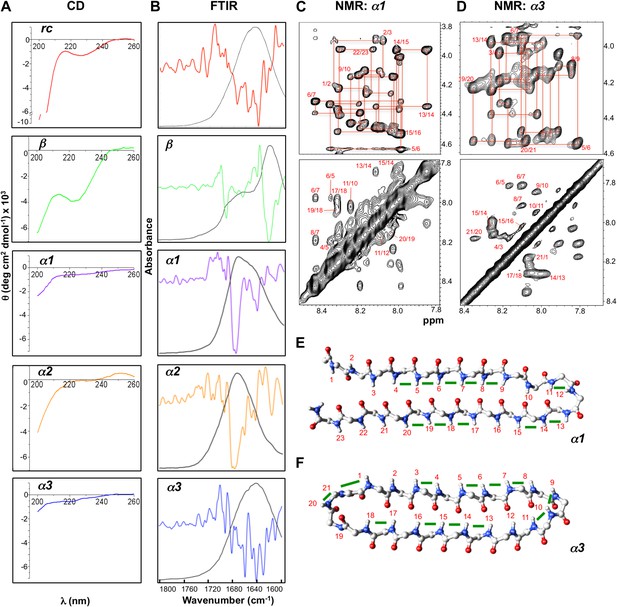
Designed peptides display unique spectroscopic signatures expected for α-sheet.
(A) CD spectra for peptide designs reveal a random coil structure for rc and β-structure spectrum with a bit of random coil for β. In contrast, α1, α2 and α3 have largely featureless CD spectra with some random coil content expected to arise from the turns and tail residues. Note the different scales for the y-axes. All spectra are presented as molar ellipticity, highlighting the difference in intensity of the random coil component for each design compared with the rc spectrum. (B) FTIR spectra of the peptide designs, displayed as both absorbance (black line) and the second derivative (colored line), correlate well with the CD spectra. The β design shows a strong signal at 1632 cm−1, as expected for β-structure. The α-sheet designs have signals near 1640 and 1675–1680 cm−1 and the absorption is more intense for α1 and α2. (C) Fingerprint (top) and NH region (bottom) of the 1H NOESY spectra for α1. Sequential assignments are shown in red and multiple sequential NOEs are observed and labeled. (D) Fingerprint (top) and NH region (bottom) of the 1H NOESY spectra for α3. The NH region reflects the predominance of NH–NH interactions and lack of other main-chain interactions characteristic of the common secondary structures. Mapping backbone NOEs on computational models as green bars (E, α1 and F, α3) reveal in-plane alignment of the peptide groups along the majority of the sheet. NOEs in the turn regions determined whether the carbonyl or amide hydrogens pointed up in the structures as oriented in the figure (N-terminus top left). Cα, C, N, H and O atoms are shown in gray, gray, blue, white and red, respectively.
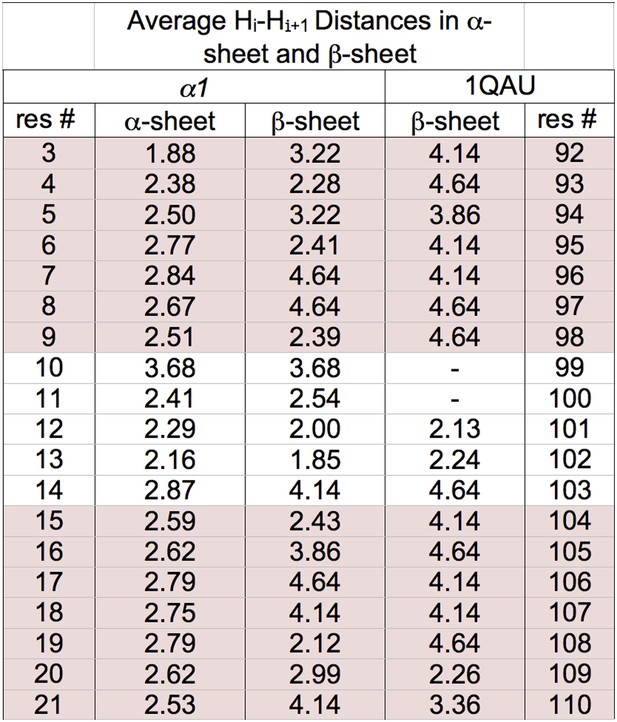
Distances corresponding to dNN NOEs calculated from MD simulations.
r−6 weighted distances calculated from MD simulations of α1 in α-sheet and in β-sheet conformation. α1 is not stable as a β-sheet and did not retain the structure well even in these short simulations. Consequently a β-hairpin of the same size within a protein was also used as a control for expected H–H distances within a dynamic but stable hairpin. Specifically, residues 105–124 of the antiparallel β-hairpin neuronal nitric oxide synthase (PDB:1QAU) was used from a 50 ns simulation at 298 K. Triplicate simulations of the isolated hairpins were performed at 298 K for 20 ns. r−6 intensities, which should be proportional to the NOE intensity, were calculated step by step from the distance then averaged across the pooled simulations and time points and converted back to distance (<r−6 >−1/6). Residue 117 of 1QAU is proline, preventing measurement for itself and the preceding residue. The lower values observed for α1 in a β-hairpin conformation, similar to those of the anti-parallel β-hairpin in 1QAU suggest to us that our α1 does not adopt β-structure. Sheet regions are highlighted.
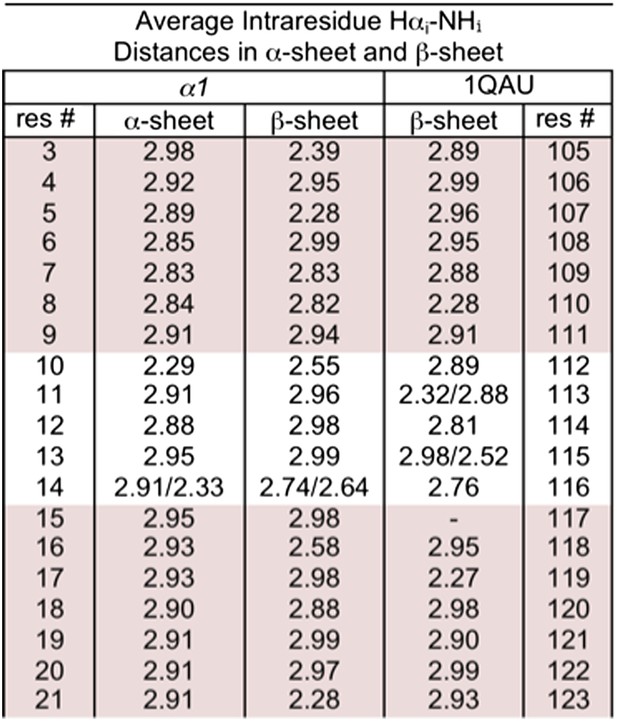
Distances corresponding to intraresidue dαN NOEs calculated from MD simulations.
r−6 weighted distances calculated from MD simulations of α1 in α-sheet and in β-sheet conformation. r−6 intensities, which should be proportional to the NOE intensity, were calculated for each structure then averaged across the pooled simulations and time points and converted back to distance (<r−6 >−1/6). Residue 117 of 1QAU is proline, preventing measurement for itself. Residues 113 and 115 of 1QAU, and residue 14 of α1 are glycines, resulting in two measurements for Hαi–HNi. Sheet regions are highlighted.
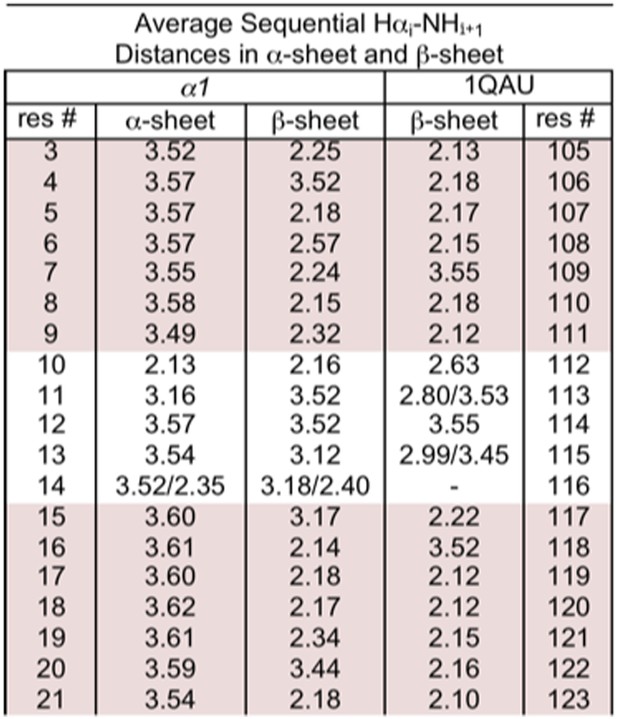
Distances corresponding to sequential dαN (i–i+1) NOEs calculated from MD simulations.
r−6 intensities, which should be proportional to the NOE intensity, were calculated for each structure and then averaged across the pooled simulations and time points and converted back to distance (<r−6 >−1/6). Residue 117 of 1QAU is proline, preventing measurement for the preceding residue. Residues 113 and 115 of 1QAU, and residue 14 of α1 are glycines, resulting in two measurements for Hαi–HNi+1. Sheet regions are highlighted.





