Reconstitution of dynein transport to the microtubule plus end by kinesin
Figures
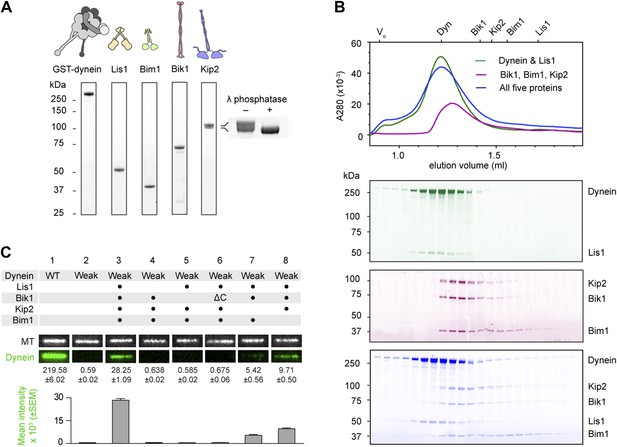
Purification and analysis of the putative dynein plus-end transport machinery.
(A) SDS-PAGE and diagrams of purified proteins. Kip2 from S. cerevisiae migrates as a doublet of bands, corresponding to differently phosphorylated isoforms as verified by phosphatase treatment (right panel). (B) Analysis of sub-complex formation by size-exclusion chromatography. Elution volumes of the individual proteins are shown on the top axis. V0: void volume. Dynein and Lis1 co-elute as a complex (green trace), as do Bik1, Bim1 and Kip2 (magenta trace). A mixture of all five proteins elutes as dynein-Lis1 and Bik1-Bim1-Kip2 sub-complexes, rather than one stable assembly (blue trace). SDS-PAGE analysis of the fractions is shown below, pseudo-colored to match the traces. (C) TIRF microscopy microtubule recruitment assay. Fluorescently labeled wild-type (WT) dynein or a mutant with weak microtubule affinity (K3116A, K3117A, E3122A, R3124A: Weak) were incubated with microtubules in the presence of the indicated proteins. Representative microtubule images and the corresponding dynein channel are shown in each condition. The width of each panel corresponds to 5.4 μm. Mean dynein fluorescence intensity ± SEM is shown below (N = 30–50 microtubules).
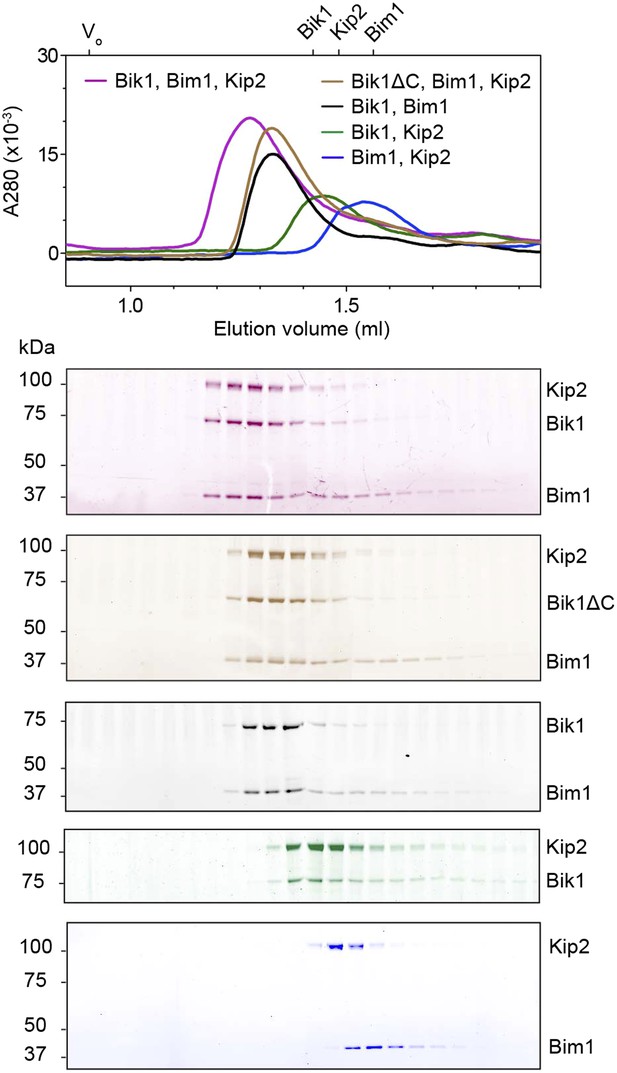
Analysis of interactions between Bik1, Bim1 and Kip2 by size-exclusion chromatography.
The void volume (V0) and elution volumes of the individual proteins are shown on the top axis. SDS-PAGE analysis of the corresponding fractions is shown below, pseudo-colored to match the traces. Bik1 and Bim1 co-elute as a complex that is upshifted relative to the individual proteins, in agreement with Blake-Hodek et al. (2010). There is a partial upshift and co-elution of Kip2 with Bik1, whereas Bim1 and Kip2 do not co-elute as a stable complex. However, all three proteins (Bik1, Bim1 and Kip2) together co-elute as an upshifted ternary complex. Removal of Bik1's C-terminal zinc knuckle domain (Bik1ΔC) does not prevent its ability to form a complex with Bim1 and Kip2.
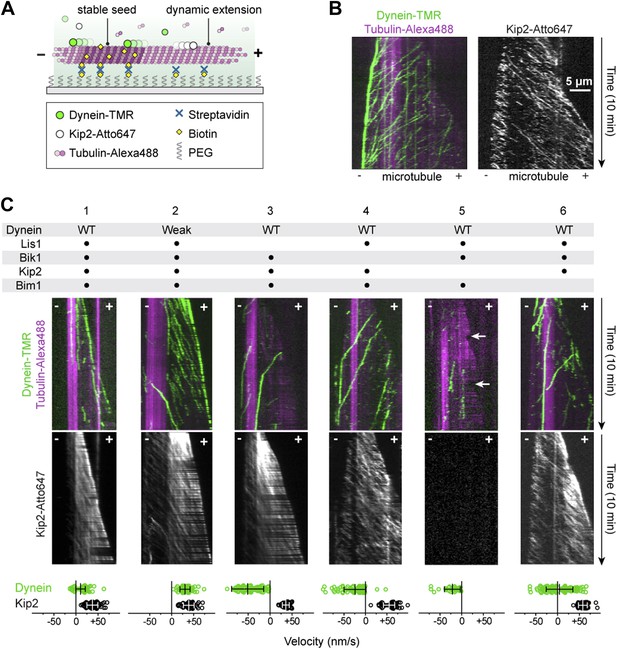
Reconstitution of dynein transport to the microtubule plus end.
(A) Diagram of dynamic microtubule assay, after Bieling et al. (2007). A bright, biotinylated, GMP-CPP-stabilized microtubule seed is attached to the coverslip via streptavidin and biotin-PEG. Dimly labeled microtubule extensions grow from this seed in the presence of tubulin and GTP. Tubulin, dynein and Kip2 are visualized using TIRF microscopy, and motor protein movement on the dynamic extensions is analyzed. (B) Dynein and Kip2 motility on dynamic microtubules. Kymographs of the dynein and microtubule channels are overlaid, and the Kip2 channel is shown separately for clarity. Plus (+) and minus (−) denote microtubule polarity. Dynein moves to the minus end and Kip2 moves to the plus end. Scale bar, 5 μm. (C) In the presence of Lis1, Bik1, Bim1 and Kip2, dynein moves to the plus end of the microtubule (lane 1). Weakening dynein’s microtubule affinity via mutagenesis (Figure 2—figure supplement 2) increases its plus-end velocity and accumulation (lane 2). In the absence of Lis1, Bik1 or Kip2, dynein resumes minus-end-directed motion (lanes 3–5). Arrows mark microtubule catastrophe events seen in the absence of Kip2 (lane 5). When Bim1 is omitted, dynein displays both plus- and minus-end-directed movements (lane 6). Mean velocities ± SD for dynein and free Kip2 (without dynein bound) are shown at bottom (N = 36–122).
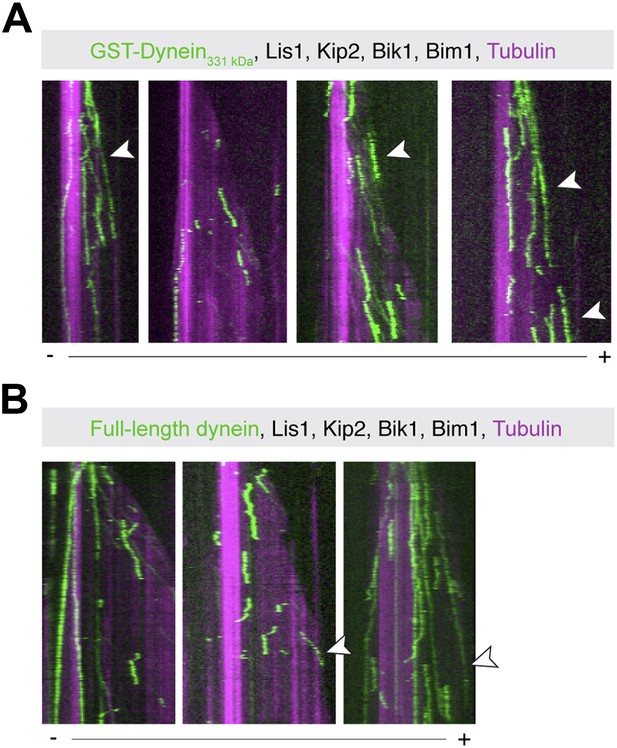
Examples of plus-end-directed motion of GST-Dynein331 kDa and full-length dynein.
Dynein at the growing microtubule plus end is marked with arrowheads. Plus (+) and minus (−) denote microtubule polarity.
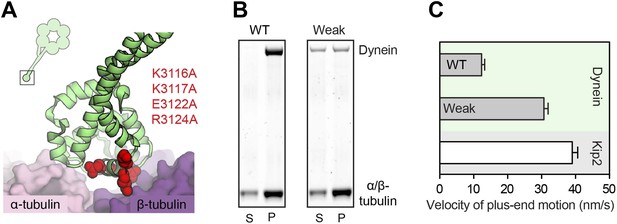
Weakening dynein's microtubule affinity increases the velocity of its plus-end-directed transport.
(A) Four charged amino acids were mutated to weaken dynein's microtubule affinity (K3116A, K3117A, E3122A and R3124A). These residues are depicted in red sphere representation using the cryo-EM derived model of dynein's microtubule-binding domain bound to an α/β-tubulin dimer (PDB 3J1T) (Redwine et al., 2012). The inset shows the location of dynein's microtubule-binding domain (boxed) in relation to the rest of the molecule. (B) These mutations substantially weaken, but do not abolish, the binding of S. cerevisiae dynein to the microtubule in the absence of nucleotide, as shown by a microtubule co-sedimentation assay. Under conditions in which wild-type (WT) dynein is fully bound to the microtubule ([α/β-tubulin]: 600 nM), the weak binding mutant only partially co-sediments with the microtubule in the pellet (P) and partially remains in the supernatant (S). See also Figure 1C. (C) In the presence of Lis1, Bik1, Bim1 and Kip2, both the WT and weak binding dynein constructs are transported by Kip2 to the plus-end of the microtubule (Figure 2C). However, the plus-end velocity of the weak-binding construct is increased relative to the wild-type dynein (p<0.0001, Student's t test), becoming close to that of free Kip2. This suggests that wild-type dynein can resist its own transport to the plus end through its microtubule-binding domain.
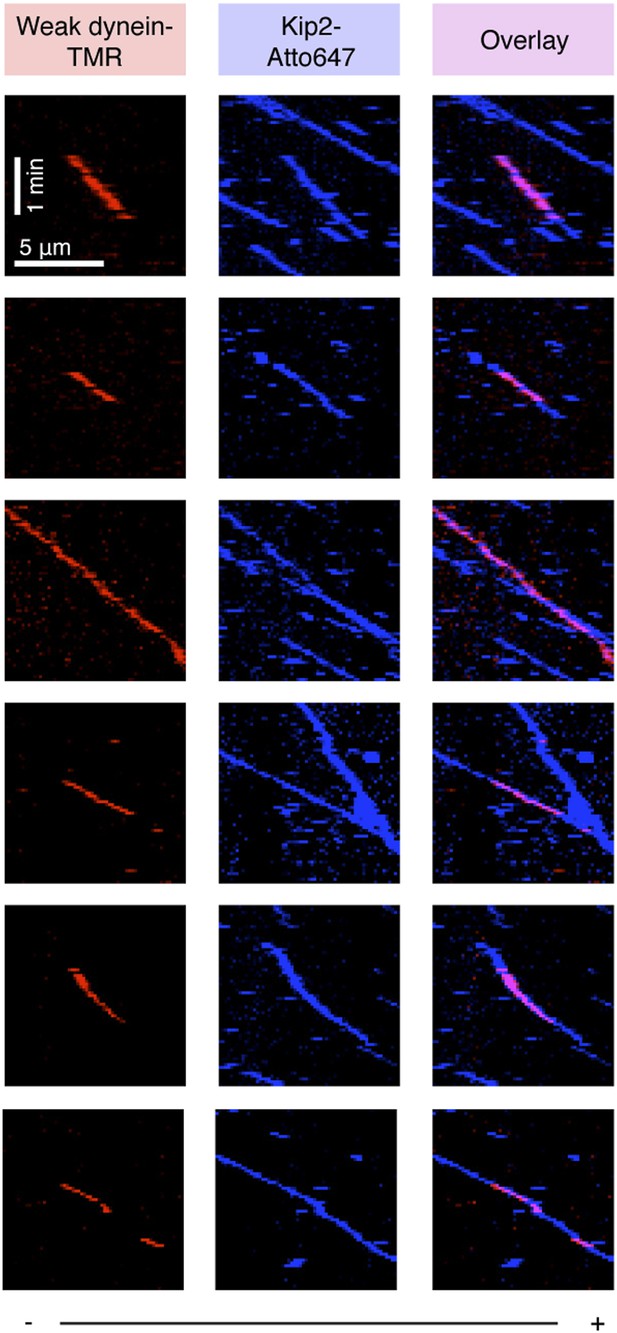

Kymographs showing colocalized, plus-end-directed runs of weak dynein and Kip2 in the presence of Bik1 and Lis1.
Plus (+) and minus (−) denote microtubule polarity.
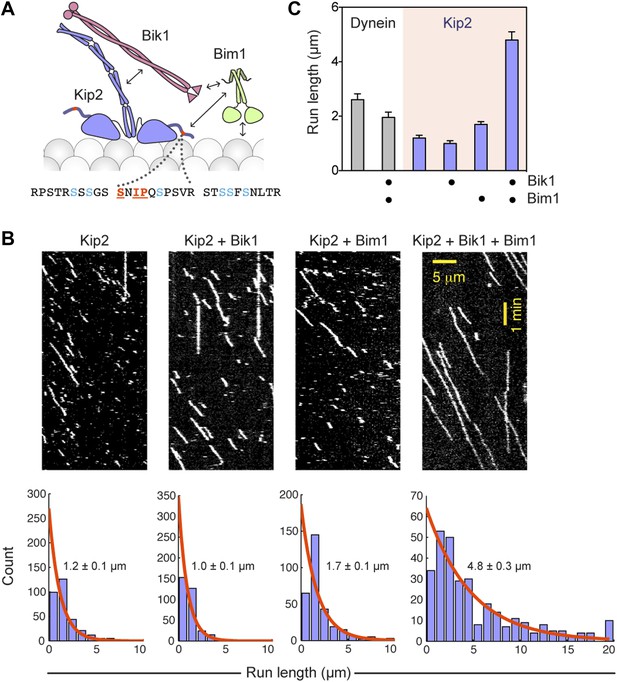
Bik1 and Bim1 are Kip2 processivity factors.
(A) Top: diagram of Kip2, Bik1 and Bim1 domain structure. Arrows indicate reported and putative interactions (see main text for details). Bottom: Kip2 contains an SxIP motif (orange) within an N-terminal extension to its kinesin motor domain. The SxIP motif is flanked by serine residues (blue) that can be phosphorylated (Holt et al., 2009; Bodenmiller et al., 2010). (B) Top: kymographs showing the movement of Kip2-Atto647 along Taxol-stabilized microtubules in the absence and presence of Bik1 and Bim1. Scale bars, 1 min and 5 μm. Bottom: histograms showing the distribution of Kip2 run lengths in each condition. Orange curves show single exponential fits, based on the cumulative distribution function of each dataset. Average run lengths were determined from the decay constant. Standard errors were determined by bootstrapping, with each dataset resampled 200 times. (C) Bar chart of run length ± SE of dynein and Kip2 toward the minus and plus end of the microtubule, respectively, in the indicated conditions (N = 233–331).
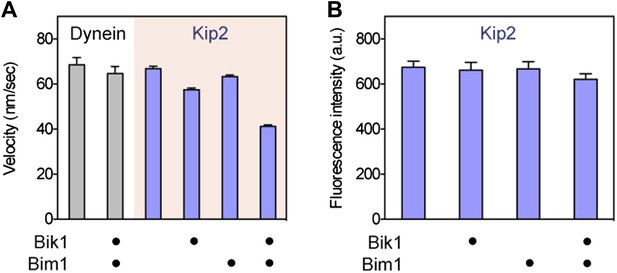
While increasing Kip2 processivity, Bik1 and Bim1 confer a small reduction in Kip2 velocity and do not change Kip2 copy number.
(A) Impact of Bik1/Bim1 on dynein and Kip2 velocity on Taxol-stabilized microtubules. Graphs show mean velocity ± SEM (N = 175–331 events per condition). Bik1 and Bim1 together confer a small reduction in the velocity of Kip2’s plus-end-directed movement (p < 0.0001, Student’s t test). (B) Bik1 and Bim1 do not change Kip2 copy number, as judged by average fluorescence intensity per Kip2 spot. Graphs show mean intensity ± SEM (N = 29–40 spots per condition). Fluorescence intensity per Kip2 spot does not change upon addition of Bik1 and Bim1 (P = 0.1529).
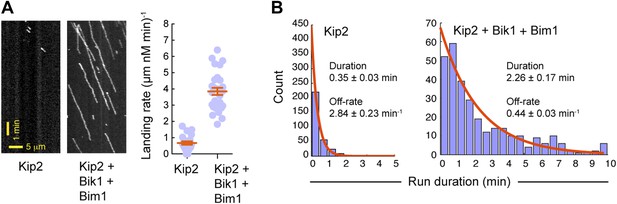
Bik1 and Bim1 increase Kip2’s microtubule on-rate and decrease its off-rate.
(A) Left: kymographs showing Kip2 at a fixed concentration in the absence or presence of Bik1 and Bim1. Right: quantification of Kip2 landing rate on the microtubule, expressed as events per micron of microtubule per nM Kip2 per minute. Bik1 and Bim1 increase Kip2's microtubule landing rate. Orange lines show mean ± SEM. (B) Histograms showing the duration of Kip2 runs in the absence or presence of Bik1 and Bim1. Orange curves show single exponential fits, based on the cumulative distribution function of each dataset. The average run duration and off-rate were determined from the decay constant. Standard errors were determined by bootstrapping, with each dataset resampled 200 times. Bik1 and Bim1 reduce Kip2's microtubule off-rate.
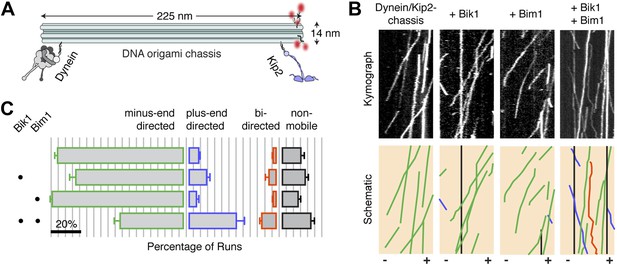
Bik1 and Bim1 regulate the outcome of a tug-of-war between dynein and Kip2.
(A) Coupling dynein and Kip2 to a DNA origami ‘chassis’ allows them to be pitted directly against each other in a tug-of-war. Single-stranded DNA oligonucleotides were attached to dynein and Kip2 via SNAP tags at their N- or C-terminus, respectively (Derr et al., 2012). These DNA oligonucleotides base pair with complementary sequences extending from the chassis. (B) Top: kymographs of dynein/Kip2-chassis motility on Taxol-stabilized microtubules in the presence and absence of Bik1 and Bim1. Bottom: Color-coded schematics, highlighting runs that are minus-end-directed (green), plus-end-directed (purple), bi-directed (orange) or non-mobile (black). (C) Quantification of the different types of dynein/Kip2-chassis behavior in the indicated conditions, expressed as the average percentage ± SEM (N = 3 separate dynein/Kip2-chassis assembly reactions, with 306–353 runs analyzed in each case). The addition of Bik1 and Bim1 causes the fraction of plus-end-directed runs to increase relative to dynein/Kip2-chassis alone (p < 0.05; Student’s t test). Scale bar, 20%.
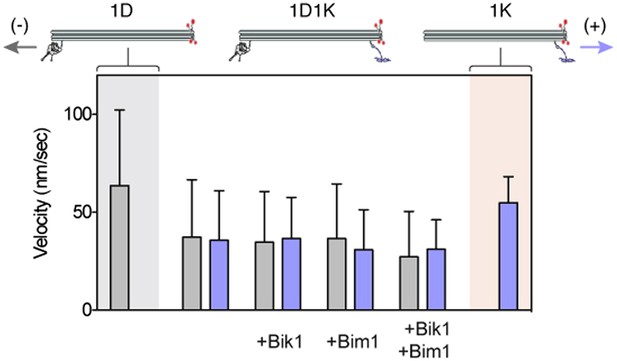
Dynein and Kip2 engage in a tug-of-war when coupled via the DNA origami chassis.
Graph shows the average velocity ± SD of chassis movements toward the microtubule minus end (gray bars) or plus end (purple bars). 1D denotes a chassis with one dynein-attachment site. 1K denotes a chassis with one Kip2-attachment site. 1D1K denotes a chassis with one dynein and one Kip2 attachment site. The velocity of the 1D1K chassis in the minus-end (dynein) direction is reduced compared to the 1D chassis, suggesting that Kip2 resists transport to the minus end (p < 0.0001, Student’s t test). The converse is also true for the velocity in the plus-end direction (p < 0.0001).
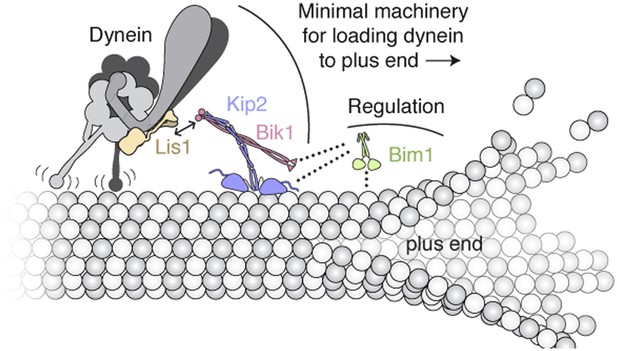
Model of the minimal machinery for transporting dynein to the microtubule plus end.
Based on results from this study and earlier work. Dynein is connected to the plus-end-directed motor Kip2 by Lis1 and Bik1. Specifically, Lis1's ß-propeller domain binds to the AAA4 subdomain of dynein's motor domain (Huang et al., 2012). The dynein/Lis1 complex interacts dynamically with the C-terminal zinc-knuckle domain of Bik1/Clip170 (Coquelle et al., 2002; Sheeman et al., 2003; Lansbergen et al., 2004; Markus et al., 2011) as indicated by the double arrow. Bik1's coiled coil is posited to interact with Kip2 (Newman et al., 2000; Carvalho et al., 2004). Bim1 is not strictly required for dynein's plus-end transport, but together with Bik1, it can enhance Kip2's processivity and help Kip2 overcome dynein's intrinsic minus-end-directed motility. As depicted with dotted lines, it is likely that the C-terminal tail of Bim1 interacts with the N-terminal Cap-Gly domain of Bik1 (Weisbrich et al., 2007), and the cargo-binding domain of Bim1 interacts with the SxIP motif of Kip2 (Honnappa et al., 2009). This arrangement would leave the calponin-homolgy domain of Bim1 (Slep and Vale, 2007) free to interact transiently with the microtubule lattice, and thus promote Kip2's microtubule association. A single copy of each molecule is shown for clarity; interplay with the numerous other + TIPs in S. cerevisiae (not shown) is expected.
Tables
Protein homolog names
| Generic | S. cerevisiae | S. pombe |
|---|---|---|
| Kinesin | Kip2 | Tea2 |
| Clip170 | Bik1 | Tip1 |
| EB1 | Bim1 | Mal3 |
| Lis1 | Pac1 | – |
Yeast strains used in this study
| Strain | Genotype | Used to purify | Source |
|---|---|---|---|
| RPY753 | PGAL1-ZZ-Tev-GFP-3xHA-GST-DYN1331kDa-gs-DHA-KanR; pac1Δ::klURA3; ndl1Δ::cgLEU2 | GST-dynein331 kDa | (Huang et al., 2012) |
| RPY780 | ZZ-Tev-3xHA-DYN1-gs-DHA-KanR nip100Δ pac1Δ::HygroR ndl1Δ::cgLEU2 | Full-length dynein | (Huang et al., 2012) |
| RPY816 | PGAL1-8HIS-ZZ-Tev-PAC1; dyn1D::cgLEU2; ndl1D::HPH | Lis1 | Julie Huang |
| RPY1084 | PGAL1-ZZ-Tev-GFP-3xHA-SNAP-gs-GST-DYN1331kDa-gs-DHA-KanR | Dynein for DNA labeling | (Derr et al., 2012) |
| RPY1099 | PGAL1-8HIS-ZZ-Tev-KIP2-g-FLAG-ga-SNAP−KanR | Kip2 | This work |
| RPY1235 | PGAL1-ZZ-Tev-GFP-3xHA-GST-DYN1E3197K-gs-DHA-KanR | Polarity marker dynein for chassis experiments | (Redwine et al., 2012) |
| RPY1536 | PGAL1-ZZ-Tev-GFP-3xHA-GST-DYN1K3116A, K3117A, E3122A, R3124A-gs-DHA-KanR | Weak dynein | This work |
-
All strains were made in the W303a background (MATa his3-11,15 ura3-1 leu2-3,112 ade2-1 trp1-1) with genes encoding the proteases Pep4 and Prb1 deleted (prb1Δ; pep4Δ::HIS5). DHA and SNAP refer to the HaloTag (Promega) and SNAP-tag (NEB), respectively. Tev indicates a Tev protease cleavage site. PGAL1 denotes the galactose promoter, which was used to induce protein expression. Amino acid spacers are indicated by g (glycine), ga (glycine–alanine), and gs (glycine–serine).
Additional files
-
Supplementary file 1
ImageJ macro used to calculate motor velocities and run lengths from kymographs.
- https://doi.org/10.7554/eLife.02641.017





