Fringe proteins modulate Notch-ligand cis and trans interactions to specify signaling states
Figures
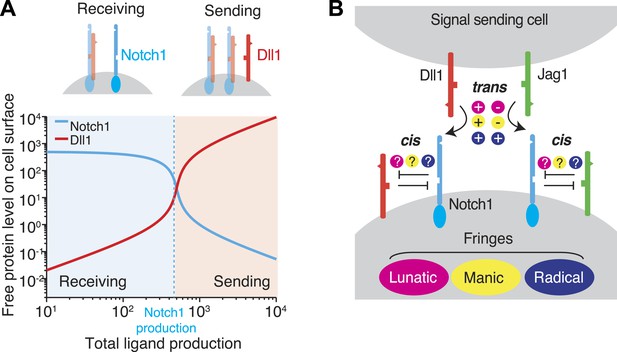
Cis interactions between receptors and ligands lead to exclusive sending and receiving signaling states.
(A) In the blue shaded region, receptor expression exceeds ligand expression (as indicated schematically above plot), so that mutual cis interactions leave mainly free receptors, allowing the cell to receive, but not efficiently send, signals. When ligand expression exceeds Notch expression, mutual cis interactions consume most of the Notch receptors, leaving an excess of free ligand, favoring sending over receiving. (B) There are multiple potential ways in which Notch1 could interact in cis and trans with Jag1 and Dll1 ligands, and in which Fringe proteins could modulate these interactions. Known interactions are indicated by + and − for positive and negative regulation, respectively. Unknown ways in which Fringe proteins could modulate these interactions are indicated by question marks.
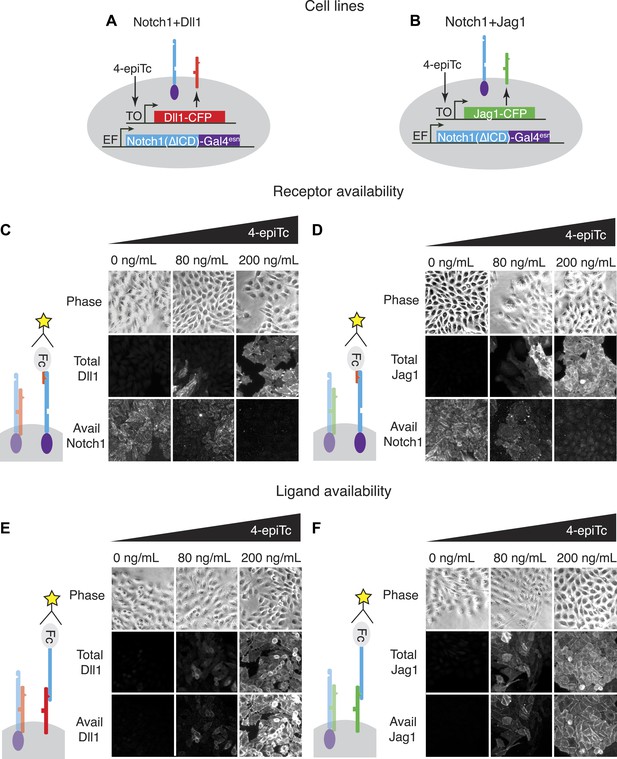
The availability assay labels receptors and ligands that can participate in trans signaling.
(A and B) Stable CHO-K1 cell lines constitutively express a Notch1-Gal4 chimeric receptor and a tetracycline-inducible Dll1 (A) or Jag1 (B) ligand fused to cerulean fluorescent protein. (C and D) In the receptor availability assay, soluble Dll1ext-Fc is bound to free Notch receptor on the surface of live cells. After fixation, bound Dll1ext-Fc is labeled with anti-Fc fluorescent reagents. Increasing ligand-Cerulean expression reduces receptor availability, as shown in these snapshots (C, D, bottom panels). (E and F) The ligand availability assay works similarly, except soluble Notch1ext-Fc fragments bind free ligands on the cell surface. Increasing ligand-Cerulean expression (E, F, middle panels), leads to increased ligand availability (E, F bottom panels). The surface ligand availability shows high spatial correlation with the total cellular ligand staining. Note that cells were plated at high cell density for illustration purposes. For quantitative analysis, cells were dissociated and plated at low density before staining (Figure 3—figure supplement 3).
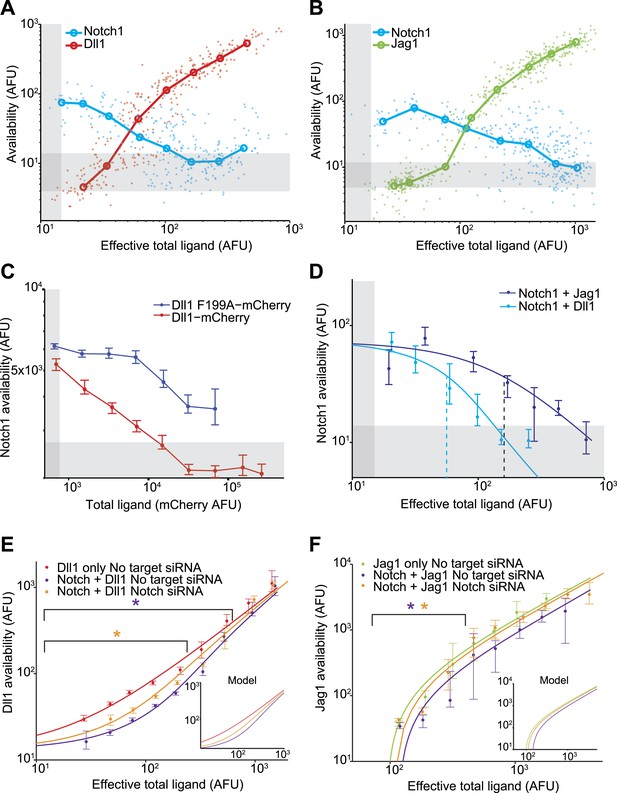
Dll1 and Jag1 exhibit mutual cis-inhibition with Notch1.
(A and B) Single cell data show decreasing receptor availability and increasing ligand availability with increasing ligand expression. Circles denote the medians of data points in logarithmically spaced bins along the x-axis. ‘Effective total ligand’ refers to the ligand availability observed at a given ligand-CFP fluorescence value in a cell line expressing only ligand. For receptor availability data in A, n = 299 and in B, n = 352. For the ligand availability data in A, n = 323 and in B, n = 530. Gray bars in all panels represent background levels, defined as the 25–75 percentile range of fluorescence from parental CHO-K1 cells that do not express Notch1 or ligand-Cerulean constructs. (C) Transient expression of wild-type Dll1-mCherry (n = 8817), but not the Dll1-mCherry F199A mutant (n = 14,292), reduced Notch availability to background levels in a Notch1 cell line. Cells were analyzed by flow cytometry. Error bars in all panels denote 95% confidence interval for the bootstrapped estimate of the median. (D) Comparison of Notch availability in the Notch1+Dll1 and Notch1+Jag1 cell lines. Lines are fits to a model of receptor-ligand cis-interactions (Supplementary). (E) Comparison of ligand availability in cell lines expressing Dll1. Ligand availability a cell line expressing Dll1 only (n = 1146). Notch1 reduces ligand availability (purple, n = 1131), and this effect is rescued by siRNA against Notch1 (orange, n = 972). In the purple starred region, cells differ significantly in ligand availability between Notch1 or no target siRNA samples, while the orange star denotes regions where Dll1 and Notch1+Dll1 cells transfected with no target siRNA differ significantly. Significance was determined by applying the Wilcoxon rank sum test. Inset shows the model behavior for parameters derived from the fit in D. Knockdown of Notch was measured to be 50%. (F) Comparisons of ligand availability in a cell line expressing Jag1 only (green, n = 733), Notch1+Jag1 (orange, n = 532), and Notch1+Jag1 with siRNA against Notch (purple, n = 1163). Starred regions indicate significance as in E. Inset shows model behavior using parameters measured in D. Knockdown of Notch was measured to be 70%.
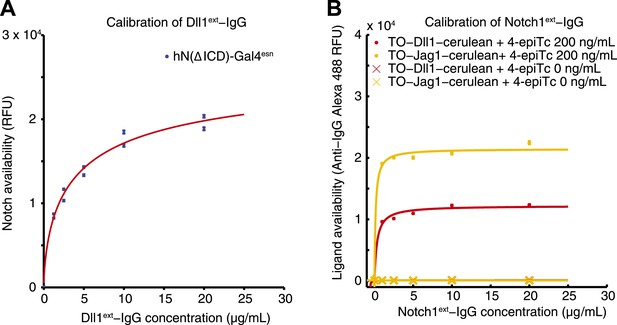
Calibration of availability assay reagents.
(A) The working concentration of Dll1ext-Fc was determined by incubating the parental hN1(ΔICD)-Gal4esn cells with increasing concentrations of soluble Dll1ext-Fc, and staining with secondary anti-IgG reagents. Blue points are results of two replicates. Data were fit (red line) to , where D is the concentration of soluble soluble Dll1ext-Fc, with a = 2.1 × 104 ± 0.17 × 104, and K = 2.27 ± 0.6 µg/ml (red line). (B) Similarly, the working concentration of N1ext-Fc was determined by incubating cells expressing fully induced TO-Jag1-cerulean (orange) and TO-Dll1-cerulean (red) with varying concentrations of N1ext-Fc, and staining with secondary reagents. Data were fit (red and orange lines) to . For the TO-Jag1-cerulean data, a = 2.1 × 104 ± 0.14 × 104, and K = 0.14 ± 0.16 µg/ml (orange line). For the TO-Jag1-cerulean data, a = 1.2 × 104 ± 0.11 × 104, and K = 0.33 ± 0.26 µg/ml (red line). Availability saturated at relatively low concentrations of the N1ext-Fc (<1 µg/ml), while un-induced cell lines showed no availability signal. Concentrations of secondary reagents were not limiting. The working concentration of both N1ext-Fc and Dll1ext-Fc reagents was set at a saturating level (10 μg/ml) based on these measurements.
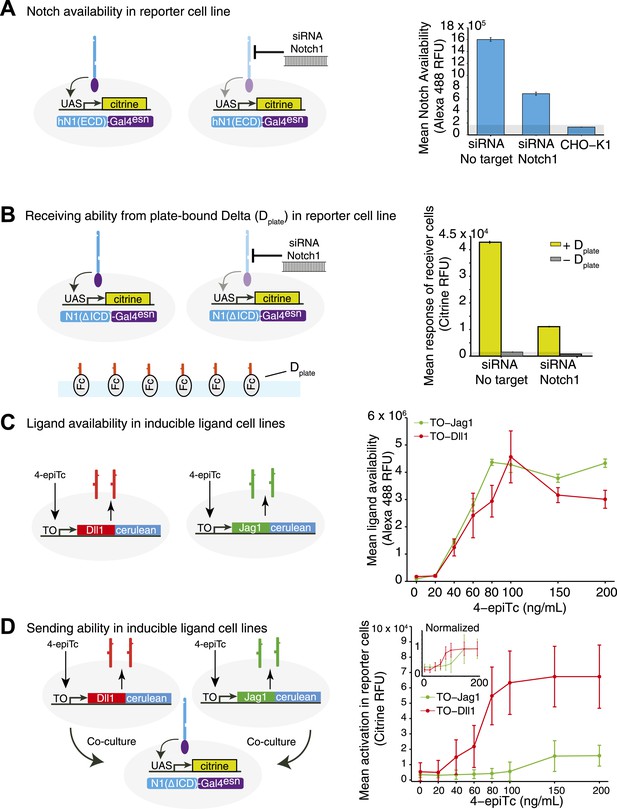
Receptor and ligand availability correlate with signal receiving and sending ability, respectively.
(A) To validate the Notch availability assay, we tested cells expressing hN1(ΔICD)-Gal4esn and observed high levels of availability fluorescence, while CHO-K1 cells lacking ectopic Notch expression showed minimal availability fluorescence (Anti-IgG Alexa 488). Bars show mean Notch1 availability, and error bars show standard error of the mean (SEM). (B) To show that receptor availability corresponds to receiving ability, we seeded a ‘receiver’ cell line expressing a diverted Notch receptor (hN1(ΔICD)-Gal4esn) and a fluorescent reporter for Notch signaling (UAS-H2B-citrine) on plates coated with Dll1 ligands. We observed a strong response from the reporter (compare ± Dplate). When we knocked down receptor expression using siRNA against the extracellular domain of Notch1, receiving ability decreased, coinciding with the decrease in Notch1 availability from A. Bars show mean reporter fluorescence, error bars are SEM. (C) Inducible ligand cells show increased ligand availability as ligand expression is induced with 4-epiTc. The TO-Dll1-cerulean and TO-Jag1-cerulean cell lines were incubated with increasing concentrations of 4epi-Tc and stained with ligand availability reagents. Points show the mean availability at each induction level, error bars show the SEM. (D) Inducible ligand cells from C are able to trans-activate Notch receiver cells in a dose-dependent fashion as ligand induction is induced with 4epi-Tc. Points are the mean of the top ten percent of receiver cells' YFP fluorescence, error bars are the standard deviation of these responders. Because ligand availability also increases with induction, ligand availability and sending ability are correlated. The Jag1 trans response in the receiver cells is weaker than the Dll1, suggesting that Jag1 is a less potent trans-activator of Notch1 than Dll1. In the inset, the same data normalized to the maximal activation elicited by each ligand. In all panels, cells were analyzed by flow cytometry.
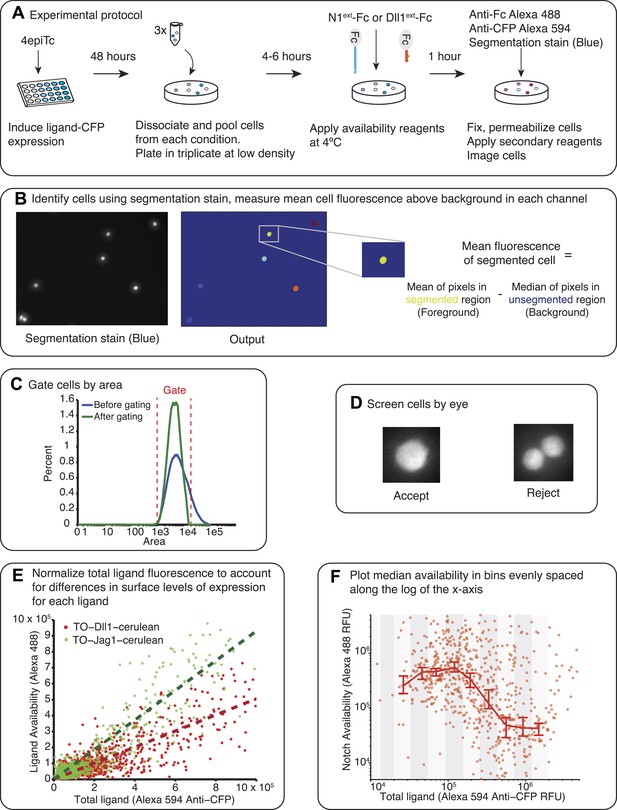
Availability protocol and analysis pipeline.
(A) Test cells were induced for 48 hr with varying concentrations of 4-epiTc. On the day of the experiment, the cells are dissociated with trypsin, split at low density and allowed to reattach to the cell culture plates. Cells are blocked with 2% FBS in PBS for 30 min at 37°C and then incubated with 10 μg/ml Dll1ext-Fc (for receptor availability) or N1ext-Fc (for ligand availability) for 1 hr at 4°C. Next, cells are washed, fixed, and permeabilized. To visualize the bound reagents, we incubated the cells with anti-Fc antibody conjugated to Alexa 488. We also added anti-CFP Alexa 594 to visualize the ligand expression. Finally, we added a blue cytoplasmic stain to identify individual cells. (B) After staining we imaged the cells on the microscope (‘Materials and methods’). The images are analyzed using a custom Matlab script to identify the cells and take the mean of the total fluorescence within each cell for each fluorescence channel. At this stage we subtract a background fluorescence value from each cell, defined as the median of the background (unsegmented) pixels in the neighborhood of the cell. (C) Next, we impose a gate on the cell area to filter out doublets and segmentation errors. (D) Then, all cells are screened by eye so that only single, isolated cells are included in the final analysis. (E) Next, we normalize the total ligand (x-axis) to account for differences in the surface expression of each ligand. In the plot in F, the same CFP fluorescence level results in different surface availability measurements for a cell line expressing only Jag1 compared to a cell line expressing only Dll1, with Jag1 showing higher surface expression. To adjust for this difference in efficiency of surface expression between the different ligands, we fit the total ligand vs ligand availability data from cells expressing ligand only with a linear fit. We use this fit to normalize the data for each ligand accordingly, allowing for comparison between different cell lines expressing different ligands. After this correction is applied, we refer to the total ligand as the ‘Effective total ligand’. (F) The single cell data from each replicate is pooled and then divided into evenly spaced bins along the log of the x (total ligand)-axis and the median availability level for each bin is plotted. We use a Matlab bootstrapping method to find the 95% confidence intervals for the estimate of the median.
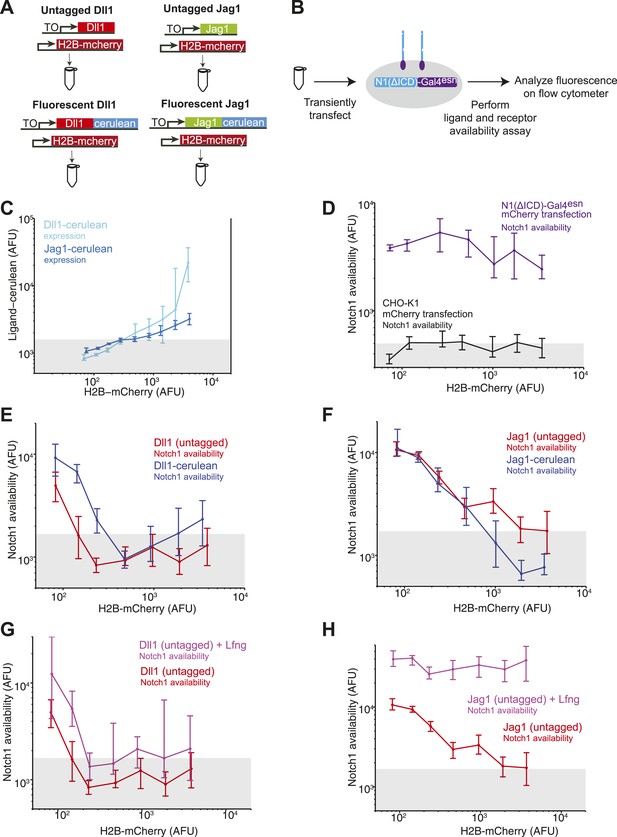
Untagged DSL ligands show similar effects to their fluorescently tagged counterparts.
(A) We compared the abilities of tagged and untagged DSL ligands to reduce Notch availability in cis. We transiently transfected the Dll1-cerulean plasmid (Figure 2A) or the Jag1-cerulean plasmid (Figure 2B) into the Notch-Gal4 receiver cell line. In parallel, we also transfected untagged Dll1 or Jag1. Because we cannot visualize the expression level of untagged ligand, we co-transfected each of the four ligands with a constitutive H2B-mCherry plasmid to label transfected cells. (B) We transfected each of the four ligands (co-transfected with H2B-mCherry) in parallel into receiver cells. After 24 hr, we performed the receptor availability assay, and analyzed the fluorescence by flow cytometry. (C) H2B-mCherry fluorescence correlated with both Dll1-cerulean expression and Jag1-cerulean expression. (D) Transfection of H2B-mCherry alone into receiver cells did not reduce Notch availability. (E) Based on the results from C, we used H2B-mCherry expression as a proxy for ligand expression level. We found that for increasing H2B-mCherry expression levels, Notch availability decreased, both for the sample transfected with Dll1-cerulean (blue), as we observed in Figure 3, and also for the sample transfected with untagged Dll1 (red). Notch availability decreased to background levels, as measured by CHO-K1 cells with availability reagents applied (gray bar denotes 10th to 90th percentile of background cells). (F) We found that increasing H2B-mCherry levels also led to a decrease in Notch availability, both for the sample transfected with Jag1-cerulean (blue), as also shown in Figure 3, and also for the sample transfected with Jag1 (red). (G) We then performed the experiments co-transfecting constitutive Lfng. We found that when we co-transfected Lfng with untagged Dll1, Notch availability decreased to background levels (magenta), just as it had without Lfng (red). (H) Finally, we compared Notch availability in a sample expressing untagged Jag1 and untagged Jag1 coexpressed with Lfng. We found that Notch availability did not decrease to background when Lfng was expressed, just as we had shown for Jag1-cerulean in Figure 4.
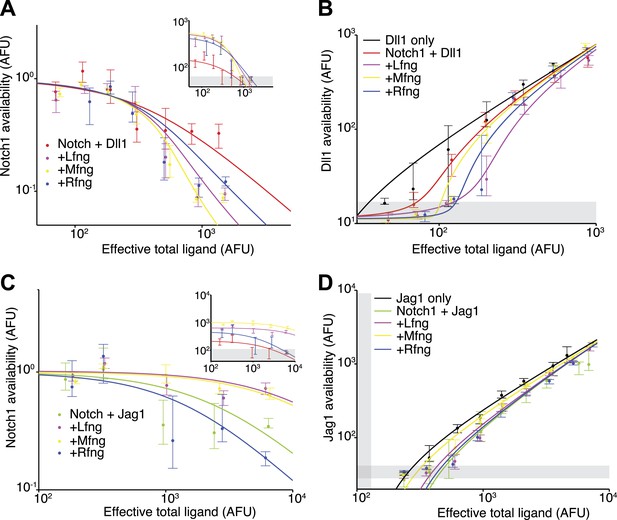
Fringe proteins show distinct effects on Jag1-Notch1 and Dll1-Notch1 cis interactions.
(A) Available Notch1 levels for the Notch1+Dll1 cell line without Fringe (red) or with Lfng (magenta), Mfng (orange), or Rfng (blue). Lines are fits to model (‘Materials and methods’). Addition of any of the three Fringes accelerates the drop-off of Notch1 availability. In the inset, the same data, but unnormalized, shows that addition of any of the three Fringe proteins does not prevent available Notch1 from reaching background levels. (B) Dll1 availability for the cell lines from A. (C) Similar to A, but for the Notch1+Jag1 cell lines. Addition of Lfng and Mfng prevents the depletion of Notch1 availability, while addition of Rfng accelerates depletion of Notch1 availability. In the inset, the unnormalized data shows that Lfng or Mfng, but not Rfng, can block the ability of Jag1 to reduce Notch1 availability to background levels. (D) Jag1 availability for the cell lines in C. In all panels, points represent medians of data points in evenly spaced bins taken along the log of the x-axis. Error bars are the 95% confidence intervals of the bootstrapped estimated of the bin medians. Solid lines are model fits to the single-cell data (see ‘Materials and methods’). Gray bars denote the 10–90th percentile fluorescence range of stained parental CHO-K1 cells that do not express Notch1 or ligands.
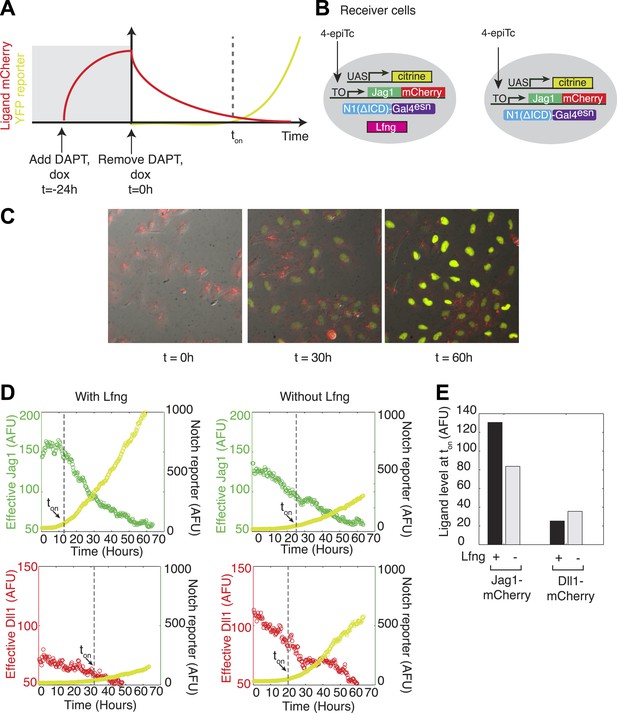
Lfng relieves Jag1-Notch1 but not Dll1-Notch1 cis-inhibition.
(A) Schematic design of time-lapse experiment to analyze cis-interactions. Before the video, cells are pre-induced to express high levels of ligand and then seeded on plates coated with Dll1-Fc. During the video, cell division dilutes the cis-ligand sufficiently to allow cells to respond to plate-bound ligands. (B) Cell lines expressing ‘diverted’ Notch1-Gal4 chimeric receptor, a fluorescent reporter for Notch signaling (UAS-H2B-citrine), and inducible Jag1-mCherry ligand, with (left) and without Lfng (right). Corresponding cell lines with the Dll1-mCherry ligand are not shown. (C) Typical video filmstrip. (D) Quantification of videos of the Notch1+Jag1 and Notch1+Dll1 cell lines, with and without Lfng. Points show the mean fluorescence of all cells in a single frame. The cell line with Lfng responds earlier than the cell line without Lfng, reflecting a weaker cis interaction between Jag1 and Notch1. The time when the YFP slope exceeds a threshold, defined as 10% of the final YFP slope, is marked as ton. (E) Quantification of the ligand levels at ton for each cell line. Values are the average of two videos. Notch activity occurs even at high cis-Jag1 levels in the +Lfng, but not the −Lfng, cell line. Notch responses occurred only at low ligand levels for the Dll1-mCherry cell lines, with and without Lfng.
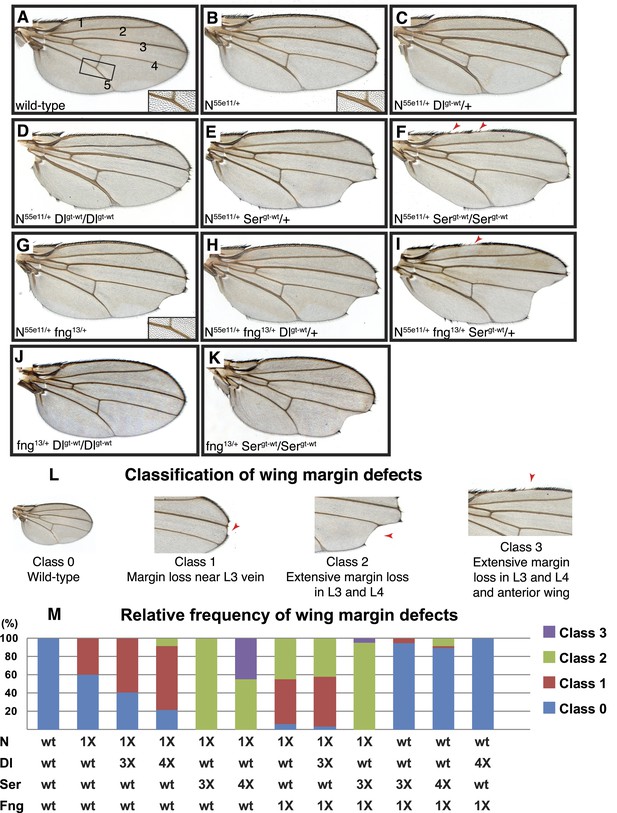
Delta and Serrate have distinct cis-inhibition phenotypes, and decreasing Fringe activity affects each of these phenotypes differently.
(A) A wild-type fly wing. The five wing veins are marked. Inset is a close-up of the area marked by the rectangle. (B) Loss of one copy of Notch leads to mild wing margin loss and wing vein thickening (inset). (C and D) One (C) or two (D) additional copies of Delta in the N55e11/+ background from B leads to mild enhancement of the wing margin defects and strong enhancement of vein thickening phenotypes. (E and F) One (E) or two (F) additional copies of Serrate in the N55e11/+ background from B results in strong enhancement of wing margin defects but only mild enhancement of vein thickening phenotypes. (G) Loss of one copy of fringe (fng13/+) in the N55e11/+ background from B enhances wing margin loss. (H) Addition of one copy of Delta to the N55e11/+ fng13/+ background from G does not further enhance margin loss and seems to suppress the vein thickening phenotype (compare to C). (I) Addition of one copy of Serrate to the N55e11/+ fng13/+ background further enhances wing margin loss in G, suggesting that Fringe usually works to counter the effects of Serrate overexpression. (J) Loss of one copy of fringe in a Delta overexpression background does not lead to any wing margin defects. (K) Loss of one copy of Fringe in a Serrate overexpression background leads to wing margin loss in some animals, suggesting that Fringe normally blocks the negative effects of Serrate on Notch signaling in animals with wild-type Notch expression levels. (L) Classification system used to quantify frequencies of mutant phenotypes. Class 0 denotes a wild-type fly wing morphology. Class 1 flies show mild wing margin loss adjacent to the L3 vein. Class 2 flies show more extensive margin loss extending to the L3 and L4 veins. Class 3 flies show margin loss in L3 and L4 and also in anterior regions of the wing. (M) Quantification of the phenotypes using the scoring system in J. At least 50 wings were scored for each genotype, except for the last two columns, for which we scored 48 and 34 wings, respectively. The most severe class 3 defects arise when Notch dosage is halved (1X Notch) and Serrate dosage is doubled (4X Serrate), a consequence of Serrate cis-inhibition. Class 3 defects also arise when Notch and fringe dosages are halved, and an additional copy of Serrate is added, suggesting that Fringe normally works to block the effects of Serrate cis-inhibition (Compare column 5 with column 9).
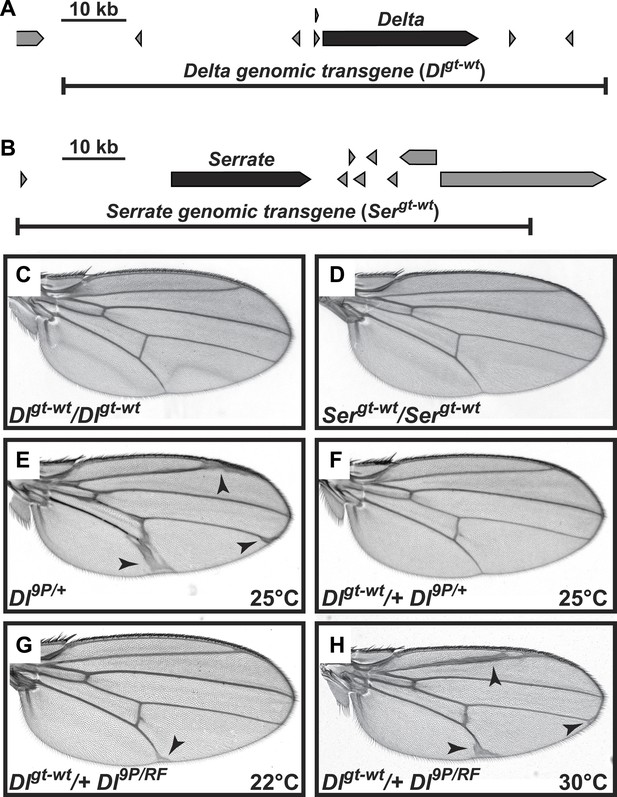
Genomic Delta and Serrate transgenes behave similarly to endogenous copies of Notch ligands in Drosophila.
(A and B) Schematic of genomic regions harboring Delta (A) or Serrate (B), their neighboring genes and their corresponding rescue transgenes. (C and D) Addition of two extra copies of Delta (C) or Serrate (D) does not alter wing margin or wing vein formation. (E) Loss of one copy of Delta results in an increase in wing vein material. (F) This phenotype is suppressed by one copy of the Dlgt-wt transgene. (G) When raised at room temperature (22°C), animals harboring Dl9P in trans to the temperature-sensitive allele DlRF (Bender et al., 1993) are lethal (not shown). However, one copy of the Dlgt-wt transgene rescues the lethality of the Dl9P/RF animals at 22°C. (H) The Dlgt-wt/+ Dl9P/RF animals raised at 30°C show a wing vein thickening phenotype similar to Dl9P/+ animals (E), most likely because increasing the temperature further decreases the activity of the DlRF allele and results in the appearance of Delta haploinsufficient phenotypes.
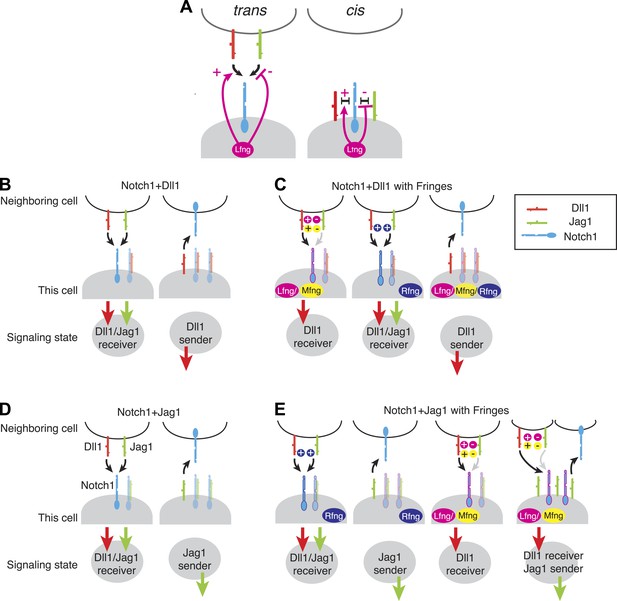
Fringe modulation of cis interactions generates a set of distinct Notch signaling states.
(A) Lfng or Mfng modification of Notch1 enhances trans activation from Dll1 and weakens trans activation from Jag1 (left). Lfng or Mfng modification of Notch1 enhances cis interactions with Dll1 and weakens cis interactions with Jag1. In (B–E), each cartoon denotes a distinct configuration of Notch pathway components, with the resulting signaling state indicated schematically below. We consider extreme endpoints, where Notch1 expression is much higher than ligand expression (Notch1 > ligand, light shaded panels), and where ligand expression is much higher than Notch1 expression (Ligand > Notch1, dark shaded panels). We also consider either low (A and C) or very high levels of Fringe expression (B and D). (A) A cell expressing Notch1 and Dll1 can be in a receiving state, where it can be activated by trans Dll1 or Jag1, when Notch1 levels surpass Dll1 levels, left. A Dll1-sending state occurs when Dll1 exceeds Notch1, right. (B) With the addition of Lfng or Mfng, the receiving state in A becomes sensitive to trans Dll1 but not trans Jag1 (left). Rfng enhances receiving from both ligands (middle). Any of the three Fringe proteins support the Dll1 sending state when Dll1 exceeds Notch1 (right). (C) Co-expression of Notch1 and Jag1 permits exclusive sending (right) or receiving (left) signaling states, similar to those in A. (D) Cells expressing Notch1, Jag1, and Rfng show exclusive send or receive signaling states as in A and C (first two panels). However, addition of Lfng or Mfng inhibits Notch1-Jag1 cis interactions. As a result, these cells can receive signals from Dll1 but not Jag1 when Notch1 expression exceeds Jag1 expression (third panel). Finally, when Jag1 exceeds Notch1, the cell can send with Jag1 and receive from Dll1 simultaneously (right panel). (E) The possible signaling states of the Notch pathway. Receptor ligand interactions prohibit cells from sending to themselves (no self arrows) and also disallow cycles.
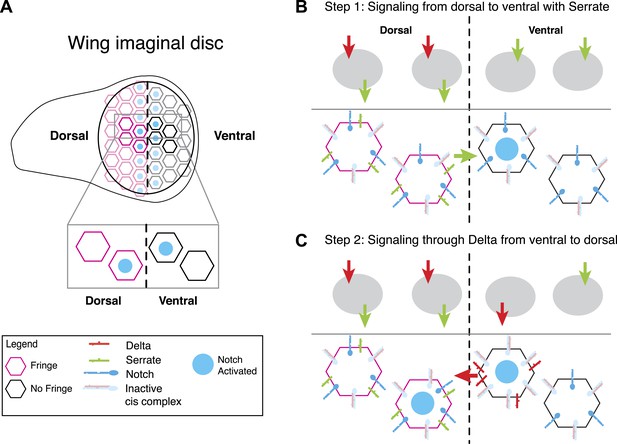
A model for Notch signaling states during dorsal-ventral boundary formation in the Drosophila wing disc.
(A) A schematic of the Drosophila wing imaginal disc during the third larval instar. During this stage of development, a sharp stripe of Notch signaling occurs at the interface of the dorsal and ventral wing compartments (blue nuclei), leading to upregulation of target genes that drive further wing development. (B) The first step of boundary formation, with signaling states indicated in cartoons above. Initially, ventral cells express Notch and Delta. Because ventral cells can receive signal, Notch must be in excess of Delta to achieve a receiving signaling state. Dorsal cells express Serrate, Notch, Delta, and Fringe (magenta outline denotes Fringe expression). Fringe promotes Delta-Notch cis interactions but weakens Serrate-Notch cis interactions, enabling dorsal cells to simultaneously receive signals from Delta while sending signals with Serrate. Thus, the first signaling step occurs from dorsal to ventral cells (green arrow). We cannot rule out that there may be some low level signaling from Delta expressed in dorsal cells as suggested by Lei et al. (2003); however, experiments suggest that Delta in ventral cells is dispensable for proper wing development. (C) The second step of signaling begins with upregulation of Delta in activated ventral cells. These cells switch to a sending state, and trans activate dorsal cells. We cannot rule out the possibility that these cells send signal to their ventral cell neighbors; however, because Delta cannot efficiently send in a Fringe-negative background, and because of the existence of feedback mechanisms at this stage, Notch activation is kept to low enough levels to prevent upregulation of target genes involved in wing margin formation (de Celis and Bray, 1997). The result is the observed pattern of Notch activation at the boundary of dorsal and ventral compartments.
Videos
Dilution video assay with a cell line expressing Notch1+Jag1.
Reporter activation shows a delay, as cell divisions are required to dilute out cis-Jag1. Frame rate is 10 fps, with a frame taken ever 20 min.
Dilution video assay with a cell line expressing Notch1+Jag1+Lfng.
Reporter activation is immediate despite high cis-Jag1 levels. Frame rate is 10 fps, with a frame taken ever 20 min.
Additional files
-
Supplementary file 1
(A) Cell lines used in this work. (B) Plasmids used in this work.
- https://doi.org/10.7554/eLife.02950.018





