Long non-coding RNAs as a source of new peptides
Figures
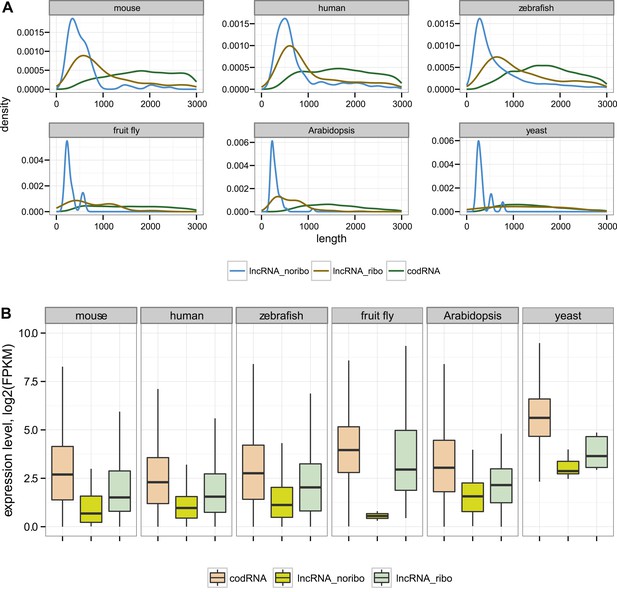
General characteristics of codRNA and lncRNA transcripts.
(A) Density plots of transcript length. (B) Box-plots of transcript expression level in log2(FPKM) units. lncRNA_ribo: lncRNAs associated with ribosomes; lncRNA_noribo: lncRNAs for which association with ribosomes was not detected. codRNA: coding transcripts encoding experimentally validated proteins except for zebrafish in which all transcripts annotated as coding were considered. The area within the box-plot comprises 50% of the data and the line represents the median value. In all studied species, codRNAs were expressed at higher levels than lncRNAs (Wilcoxon test, p < 10−5), and lncRNA_ribo at higher levels than lncRNA_noribo (Wilcoxon test, p < 0.005).
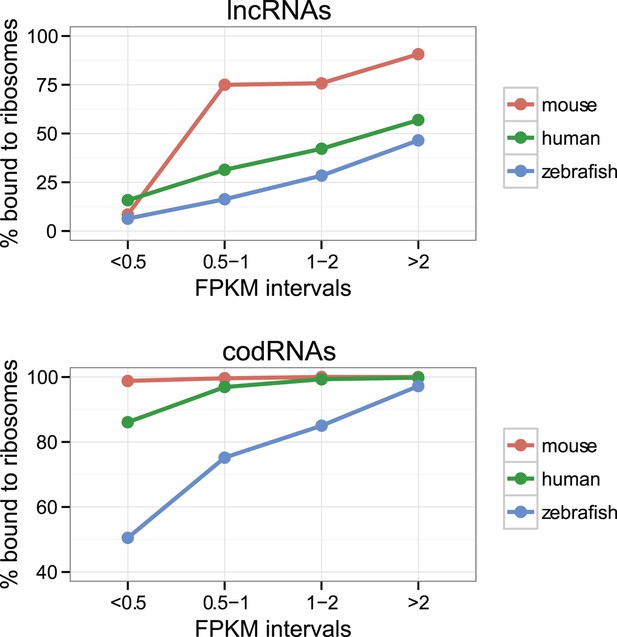
Effect of transcript expression level on the detection of ribosome association.
The percentage of transcripts associated with ribosomes is shown for several transcript expression intervals. codRNA: annotated coding transcripts encoding experimentally verified proteins (except in zebrafish for which all coding transcripts were considered). lncRNA: annotated and novel long non-coding RNAs. Only species with at least 20 transcripts in each expression bin were plotted. In the rest of species, the data were consistent with the trends shown.
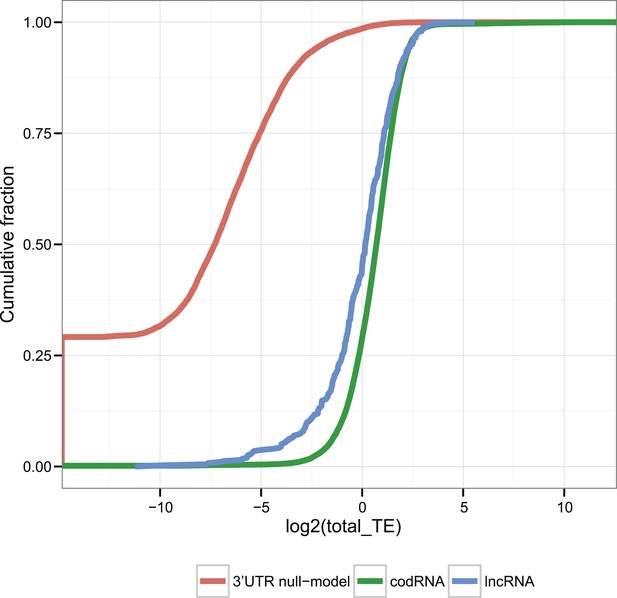
TE distribution in human transcripts and 3′UTRs (null-model).
Cumulative distribution of TE values in human codRNAs, lncRNAs, and 3′UTR sequences. We randomly selected 3′UTRs with a minimum length of 30 nucleotides to build a set of 3′UTR sequences with the same size distribution as the complete transcripts.
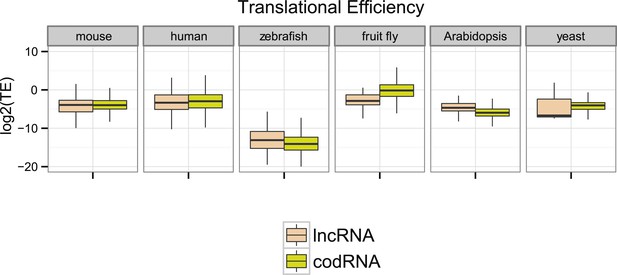
Ribosome association profiles for codRNAs and lncRNAs.
Box-plots of transcript translational efficiency (TE) in log2(TE) units. The area within the box-plot comprises 50% of the data, and the line represents the median value. lncRNA: lncRNAs for which association with ribosomes was detected. codRNA: coding RNAs transcripts encoding experimentally validated proteins except for zebrafish in which all transcripts annotated as coding were considered.
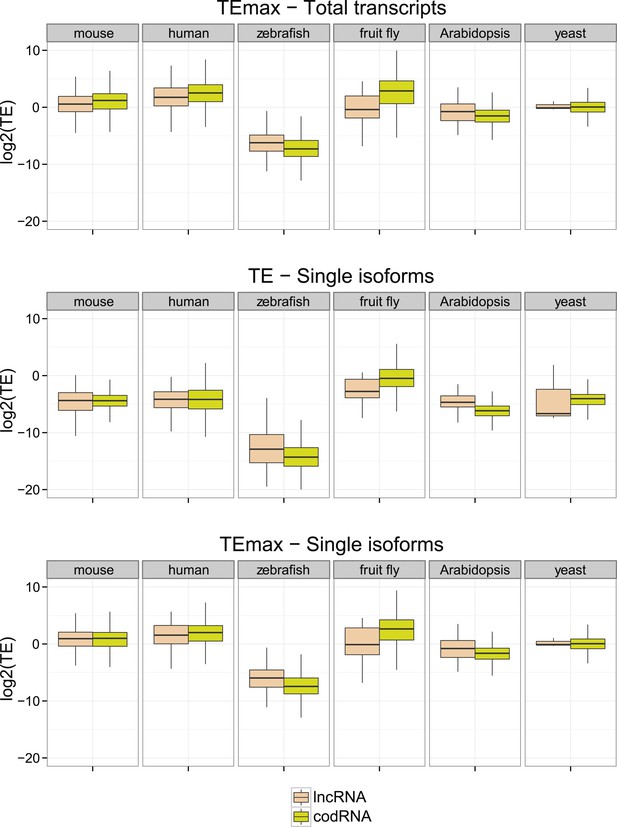
Additional translational efficiency (TE) measures.
Single isoforms correspond to data for genes with a single transcript. The number of such genes was 2961 codRNA and 246 lncRNA_ribo for mouse, 2853 codRNA and 150 lncRNA_ribo for human, 9352 codRNA and 412 lncRNA_ribo for zebrafish, 836 codRNA and 18 lncRNA_ribo for fruit fly, and 3024 codRNA and 92 lncRNA_ribo for Arabidopsis. In the case of yeast, all genes were taken. TE max is the maximum TE value taking 90 nucleotide windows.
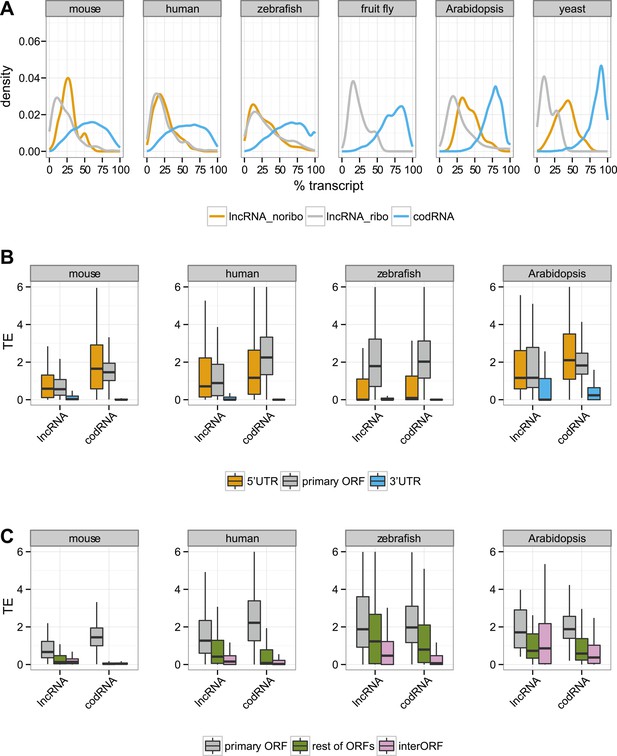
Ribosome association in different transcript regions.
(A) Density plot of the relative length of the primary ORF in lncRNA_ribo and codRNA with respect to transcript length. For comparison data for the longest ORF in lncRNA_noribo is also shown (except for fruit fly due to insufficient data). (B) Box-plots of TE distribution in primary ORF, 5′UTR, and 3′UTR regions. The area within the box-plot comprises 50% of the data, and the line represents the median value. The analysis considered all transcripts with 5′UTR and 3′UTR longer than 30 nucleotides and >0.2 FPKM in all three regions. The number of transcripts was 1956 codRNA and 159 lncRNA_ribo in mouse, 3558 codRNA and 139 lncRNA_ribo in human, 5216 codRNA and 252 lncRNA_ribo in zebrafish, and 2019 codRNA and 33 lncRNA_ribo in Arabidopsis. (C) Box-plots of TE distribution in primary ORFs, rest of ORFs with ribosome profiling reads and non-ORF regions (interORF). The analysis considered all transcripts with at least two ORFs and more than 30 nucleotides interORF. The number of transcripts was 3264 codRNA and 204 lncRNA_ribo in mouse, 3104 codRNA and 168 lncRNA_ribo in human, 1646 codRNA and 212 lncRNA_ribo in zebrafish, and 1098 codRNA and 25 lncRNA_ribo in Arabidopsis. Fruit fly and yeast were not included in the last two analyses due to insufficient data (<8 lncRNA_ribo meeting the conditions).
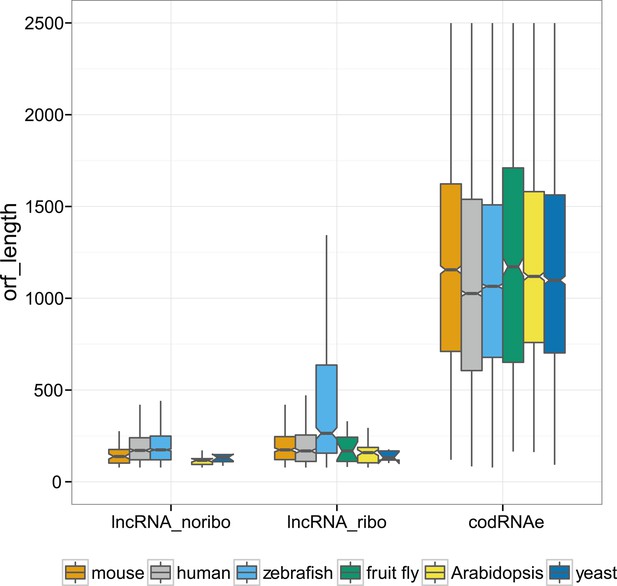
Absolute nucleotide length of ORFs in different kinds of transcripts.
In codRNAs and lncRNA_ribo, we selected the primary ORF (the ORF with the largest number of ribosome profiling reads), whereas in lncRNA_noribo we selected the longest ORF.
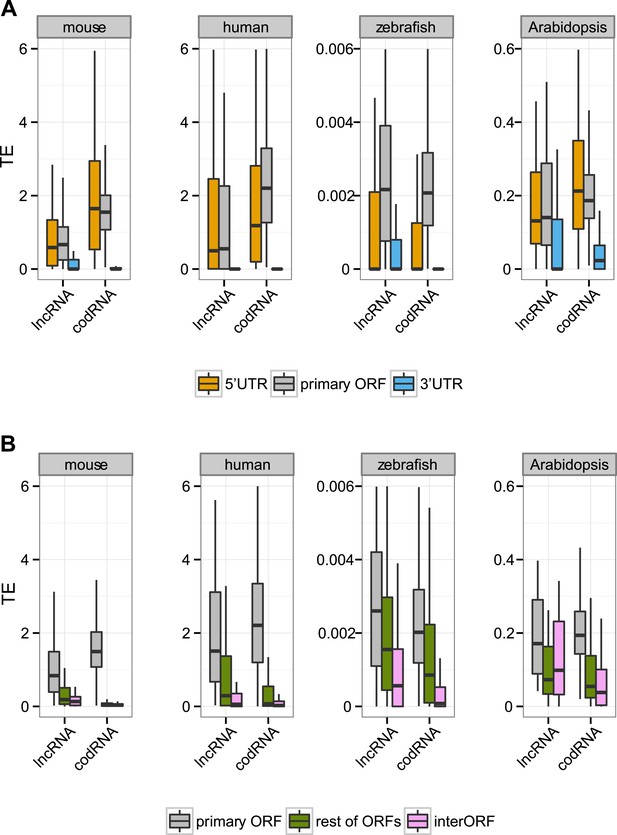
Translational efficiency in single-isoform genes.
(A) Box-plots of TE distribution in primary ORF, 5′UTR, and 3′UTR regions. The analysis considered only genes with one isoform, with UTR and ORF regions expressed at >0.2 FPKM and with 5′UTR and 3′UTR longer than 30 nucleotides. The number of transcripts was 980 codRNA and 97 lncRNA_ribo in mouse, 758 codRNA and 36 lncRNA_ribo in human, 3763 codRNA and 117 lncRNA_ribo in zebrafish, and 1495 codRNA and 32 lncRNA_ribo in Arabidopsis. (B) Box-plots of TE distribution in primary ORFs, other ORFs with ribosome profiling reads and non-ORF regions (interORFs). The analysis only considered genes with one isoform in which these regions were longer than 30 nucleotides and with expression >0.2 FPKM. The number of transcripts was 1691 codRNA and 113 lncRNA_ribo in mouse, 763 codRNA and 54 lncRNA_ribo in human, 1170 codRNA and 108 lncRNA_ribo in zebrafish, and 817 codRNA and 25 lncRNA_ribo in Arabidopsis.
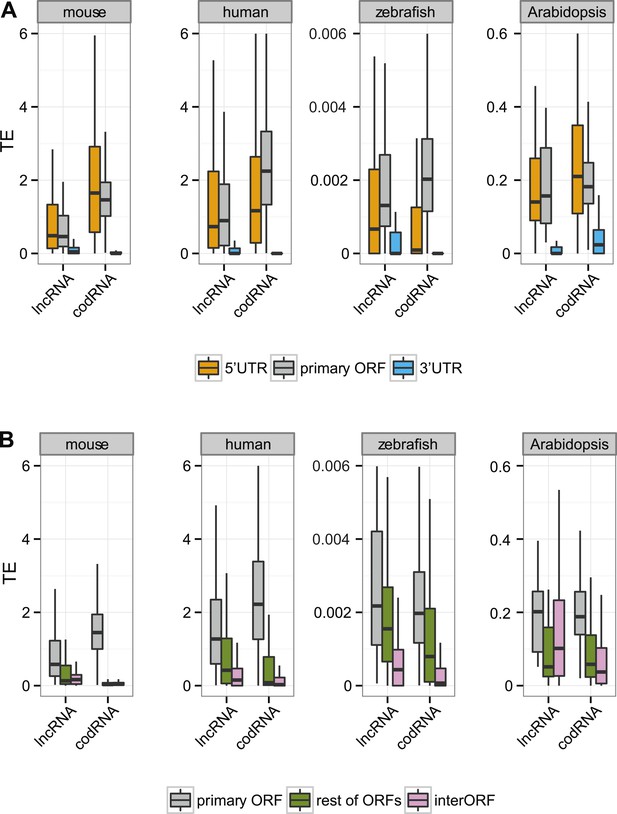
Translational efficiency in annotated transcripts.
(A) Box-plots of TE distribution in primary ORF, 5′UTR, and 3′UTR regions. The analysis considered only annotated transcripts, with UTR and ORF regions expressed at >0.2 FPKM and with 5′UTR and 3′UTR longer than 30 nucleotides. The number of transcripts was 1956 codRNA and 92 lncRNA_ribo in mouse, 3558 codRNA and 138 lncRNA_ribo in human, 5216 codRNA and 54 lncRNA_ribo in zebrafish, and 2019 codRNA and 22 lncRNA_ribo in Arabidopsis. (B) Box-plots of TE distribution in primary ORFs, other ORFs with ribosome profiling reads (rest ORFs) and non-ORF regions (interORF). The analysis only considered annotated transcripts in which these regions were longer than 30 nucleotides and with expression >0.2 FPKM. The number of transcripts was 3264 codRNA and 128 lncRNA_ribo in mouse, 3104 codRNA and 167 lncRNA_ribo in human, 1646 codRNA and 58 lncRNA_ribo in zebrafish, and 1098 codRNA and 18 lncRNA_ribo in Arabidopsis.
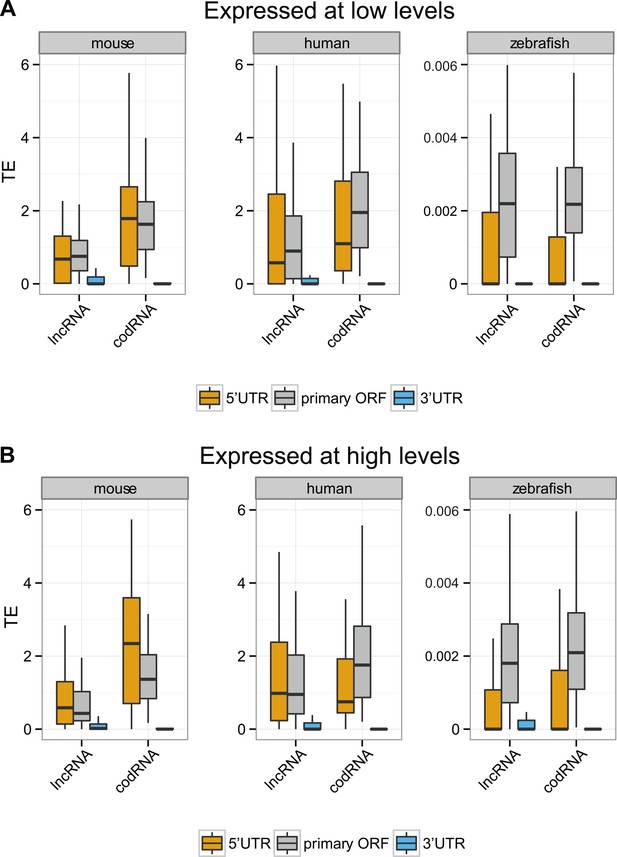
Translational efficiency in transcripts expressed at different levels.
We restricted this analysis to transcripts with ORF and UTR regions expressed at >0.2 FPKM and with 5′UTR and 3′UTR longer than 30 nucleotides. (A) Expressed at low levels: transcripts expressed at 0.5–2 FPKM, (B) expressed at high levels: transcripts expressed at 2–10 FPKM. codRNAs were sampled in such a way as to have the same gene expression distribution as the corresponding lncRNA set. Results for species in which all sets contained at least 20 transcripts are shown.
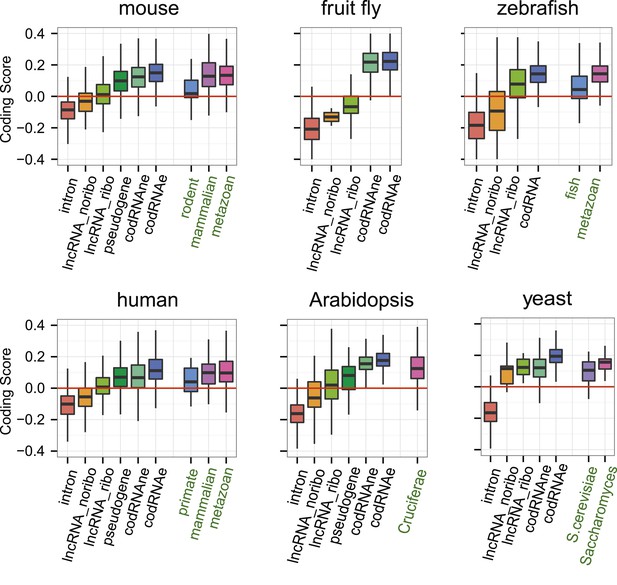
Coding scores in ORFs from different types of transcripts.
Intron: randomly selected intronic regions; lncRNA_noribo: lncRNAs not associated with ribosomes; lncRNA_ribo: lncRNAs associated with ribosomes; pseudogene: pseudogenes associated with ribosomes; codRNAne: coding transcripts encoding non-validated proteins associated with ribosomes; codRNAe: coding transcripts encoding experimentally validated proteins. The coding score was calculated as the log ratio of hexamer frequencies in coding vs intronic sequences. In lncRNA_noribo and introns, we considered the longest ORF and in the rest of transcripts the primary ORF. The Class ‘pseudogene’ was only included in species with more than 20 expressed pseudogenes with mapped ribosome profiling reads. The coding score of the primary ORF in lncRNAs (lncRNA_ribo) was significantly higher than the coding score in ORFs defined in introns (Wilcoxon test, human, mouse, zebrafish, and Arabidopsis p < 10−16; fruit fly and yeast p < 10−4, Wilcoxon test) and in lncRNA_ribo it was significantly higher than in lncRNA_noribo in four species (Wilcoxon test, human, mouse and zebrafish p < 10−5, and Arabidopsis p < 0.05). Transcripts from genes of different evolutionary age were taken from the literature (see manuscript text). The number of transcripts was 68 for rodent, 127/123 for mammalian (mouse/human as reference species), 11,203/13,423/9812 for metazoan (mouse/human/zebrafish), 162 for fish, 208 for Crucifera, 28 for S. cerevisiae and 84 for Saccharomyces. The youngest class of codRNAs displayed similar scores than lncRNA_ribo in mouse, zebrafish, and yeast (classes rodent, fish and S. cerevisiae, respectively), being only significantly higher in human and Arabidopsis (Wilcoxon test, p < 0.005; classes primate and Cruciferae). We did not analyze young genes in fruit fly due to lack of a suitable young set of codRNAs in this species.
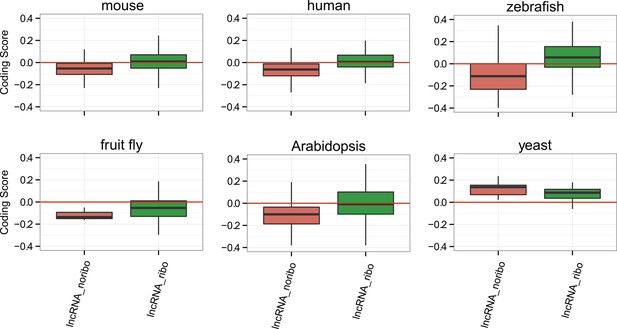
Coding scores for the longest ORF.
Comparison between lncRNAs associated and not associated with ribosomes using the longest ORF in both cases (lncRNA_ribo and lncRNA_noribo, respectively). Differences between lncRNA_ribo and lncRNA_noribo are significant by a Wilcoxon test (p < 10−10 in human, mouse, and zebrafish; p < 0.005 in Arabidopsis).
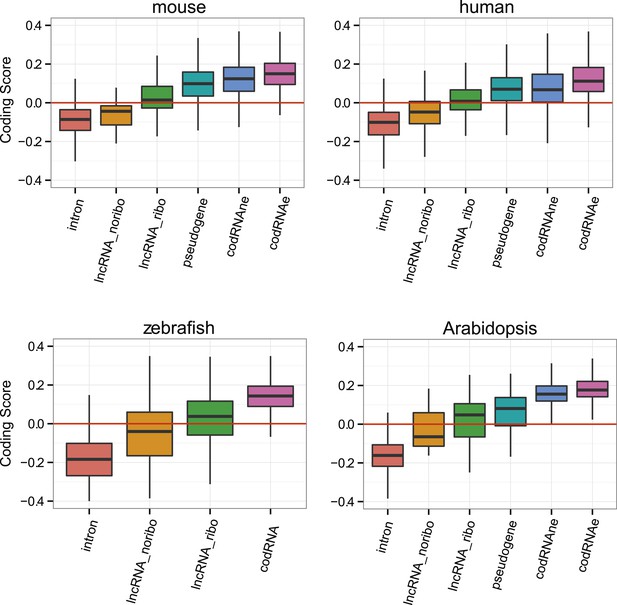
Coding scores in different classes of annotated sequences.
Comparison between different transcript classes using only annotated lncRNAs. Yeast transcriptome is composed of very few annotated lncRNAs, and this analysis could not be performed.
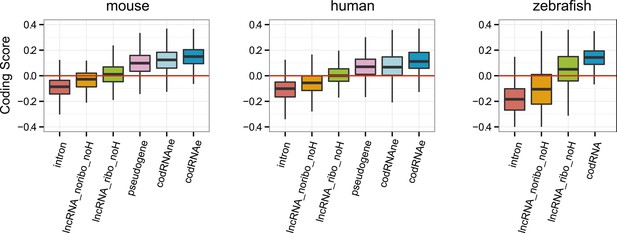
Coding scores in lncRNAs without homologues in other species.
Comparison between different transcript classes using only lncRNA with no homologues (noH) in other species. Only species in which several lncRNA_ribo and lncRNA_noribo had homology matches were considered.
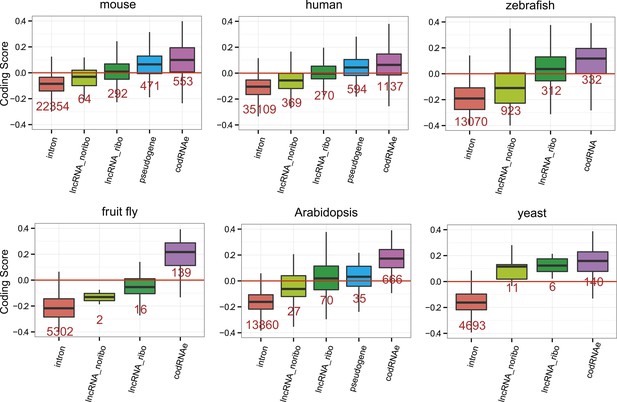
Coding scores in small ORFs from different types of transcripts.
Here we only employed lncRNAs in which the primary ORF was shorter than 100 amino acids. codRNA refers to joined codRNAe and codRNAne sets, since experimentally verified proteins are usually longer than 100 amino acid. The number of transcripts is shown in red.
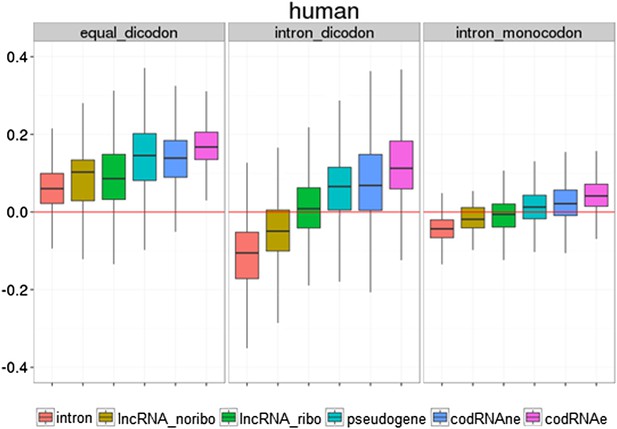
Use of different coding statistics in human transcripts.
Equal dicodon was based on the observed hexamer frequencies in coding sequences vs hexamer equiprobability, intron dicodon was based on the differences between hexamer frequencies in coding vs non-coding sequences and intron_monocodon was based on the observed codon frequences in coding sequences vs codon equiprobability.
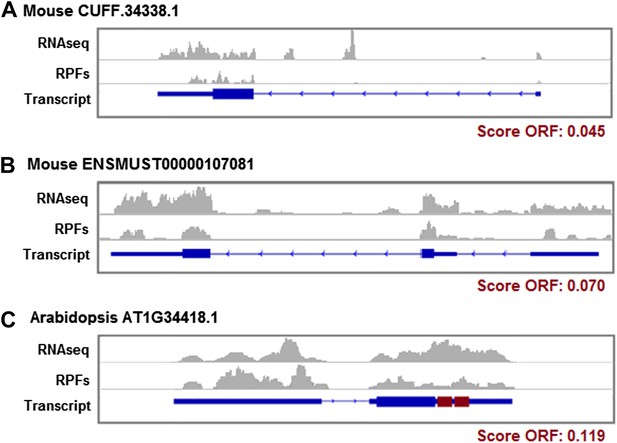
Ribosome protection patterns in transcripts containing short ORFs.
(A) Mouse CUFF.34338.1 (chr5:113183493–113188347) is a novel lncRNA, it contains an ORF encoding a 169 amino acid protein associated with ribosomes and with protein-coding homologues in human, zebrafish, and yeast. (B) ENSMUST00000107081 is an annotated codRNA in mouse which evolved recently since no homologues were found in any other species. It has a small ORF that translates a 55 amino acid protein. (C) AT1G34418.1 is an annotated lncRNA in Arabidopsis showing abundant association with ribosomes in the 5′UTR region, the primary ORF (34 amino acid) and the final region of the transcript, which contains two redundant ORFs (in red) coding the sequence: MGLGFVN(V/F)LLGM. RNAseq: profile of RNAseq reads. RPFs: profile of ribosome profiling reads. Exon-intron transcript structures are represented; the thickest boxes on the exons are the primary ORFs.
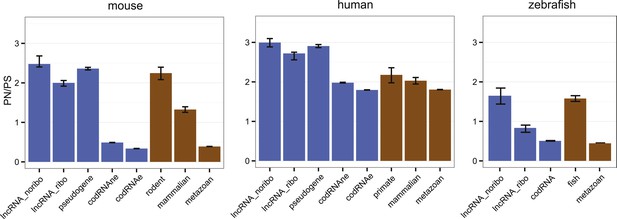
Selective pressure in ORFs from different types of transcripts.
PN/PS: ratio between the number of non-synonymous and synonymous single nucleotide polymorphisms (SNPs) in the complete set of primary ORFs for a given class of transcripts (in lncRNA_noribo the longest ORF was considered). In blue, data for different coding and non-coding transcript classes. In brown, data for different age codRNA classes. The bars represent the 95% confidence interval for the PN/PS value. For the species not shown there was not sufficient data to perform this analysis.
Tables
Data sets used in the study
| Species | GEO Accession | Mapped reads (millions) | Max read length (bp) | Description | Reference | |
|---|---|---|---|---|---|---|
| Mouse M. musculus | RNA-seq | GSE30839 | 226.0 | 43 | ES cells, E14 | Ingolia et al., 2011 |
| Ribosome profiling | GSE30839 | 39.2 | 47 | |||
| Human H. sapiens | RNA-seq | GSE22004 | 29.8 | 36 | HeLa cells | Guo et al., 2010 |
| Ribosome profiling | GSE22004 | 78.3 | 36 | |||
| Zebrafish D. rerio | RNA-seq | GSE32900 | 1382.2 | 2 × 75 | Series of developmental stages | Chew et al., 2013 |
| Ribosome profiling | GSE46512 | 1040.0 | 44 | |||
| Fruit fly D. melanogaster | RNA-seq | GSE49197 | 1317.9 | 50 | 0–2hr embryos, wild type | Dunn et al., 2013 |
| Ribosome profiling | GSE49197 | 105.7 | 50 | |||
| Arabidopsis A. thaliana | RNA-seq | GSE50597 | 79.8 | 51 | No stress conditions, TRAP purification | Juntawong et al., 2014 |
| Ribosome profiling | GSE50597 | 140.3 | 51 | |||
| Yeast S. cerevisiae | RNA-seq | GSE52119 | 20.54 | 50 | GSY83, diploid | McManus et al., 2014 |
| Ribosome profiling | GSE52119 | 6.83 | 50 | |||
Fraction of transcripts associated with ribosomes
| codRNA | lncRNA | |||||
|---|---|---|---|---|---|---|
| Expressed | Associated with ribosomes (RP) | Expressed | Associated with ribosomes (RP) | |||
| Total | Stringent | Total | Stringent | |||
| Mouse | 14,245 | 14,196 (99.7%) | 13,918 (97.7%) | 476 | 390 (81.9%) | 367 (77.1%) |
| Human | 17,011 | 16,630 (97.8%) | 16,617 (97.7%) | 934 | 403 (43.1%) | 343 (36.7%) |
| Zebrafish | 12,595 | 11,643 (92.4%) | 11,637 (92.4%) | 2392 | 726 (30.4%) | 684 (28.6%) |
| Fruit fly | 8041 | 8031 (99.9%) | 7623 (94.8%) | 28 | 22 (78.6%) | 10 (35.7%) |
| Arabidopsis | 19,162 | 18,879 (98.5%) | 10,329 (53.9%) | 139 | 93 (66.9%) | 68 (48.9%) |
| Yeast | 4740 | 4547 (95.9%) | 4335 (91.5%) | 21 | 6 (28.6%) | 6 (28.6%) |
-
Stringent: number of transcripts significant at p < 0.05 using 3′UTRs as a null model (see ‘Materials and methods’ for more details).
Fraction of translated proteins of different size detected in proteomics databases
| Protein size (amino acids) | ||||
|---|---|---|---|---|
| Species | 24–80 | 81–130 | 131–180 | >180 |
| Mouse | 27/58 (46.6%) | 222/286 (77.6%) | 256/330 (77.6%) | 3716/4786 (77.7%) |
| Human | 116/272 (42.6%) | 536/748 (71.7%) | 669/875 (76.5%) | 6757/8964 (75.4%) |
| Yeast | 27/30 (90.0%) | 168/207 (81.1%) | 234/265 (88.3%) | 2934/3224 (91.0%) |
-
Only transcripts encoding experimentally validated proteins (codRNAe) were considered.
Additional files
-
Supplementary file 1
Long non-coding RNAs as a source of new peptides. (A) Details on the number of coding transcripts associated with ribosomes. (B) ORF density and length in different types of transcripts. (C) Details on the number of non-coding transcripts associated with ribosomes. (D) Homology hits for ORFs. (E) GC content (%) in ORFs and complete sequences. (F) PN and PS values for different sequence subsets.
- https://doi.org/10.7554/eLife.03523.024
-
Supplementary file 2
(A) Human ncRNA literature. (B) IncRNA homologies. (C) IncRNA top coding score. (D) Young codRNAe.
- https://doi.org/10.7554/eLife.03523.025





