Favipiravir elicits antiviral mutagenesis during virus replication in vivo
Figures
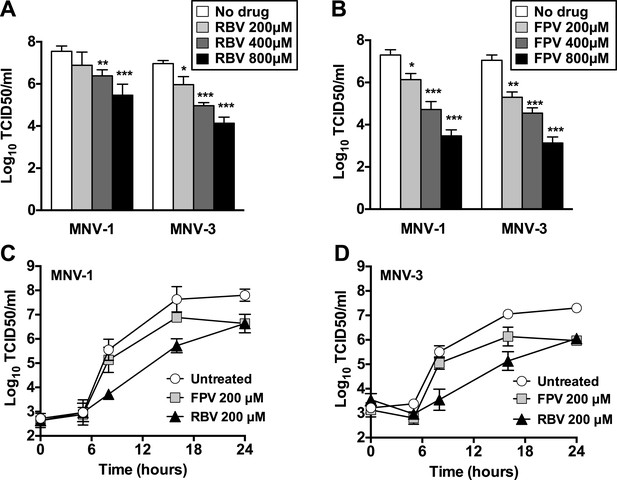
Ribavirin and favipiravir are efficient inhibitors of norovirus replication.
(A) MNV-1 and MNV-3 viral yields obtained after infection of confluent monolayers of RAW264.7 cells in the absence (white bars) or presence of 200 (light grey), 400 (dark grey), or 800 μM ribavirin (black). MNV was inoculated at an MOI of 0.01 TCID50/cell and infections were allowed to proceed for 24 hr when cultures were freeze-thawed for virus release. (B) MNV-1 and MNV-3 viral yields obtained after infection of confluent monolayers of RAW264.7 cells in the absence (white bars) or presence of 200 (light grey), 400 (dark grey), or 800 μM favipiravir (black). MNV was inoculated at an MOI of 0.01 TCID50/cell and infections were allowed to proceed for 24 hr. Statistically significant differences are represented (p < 0.05, *; p < 0.01, **; p < 0.001, ***; 2-way ANOVA test). (C and D) Kinetics of MNV-1 and MNV-3 infection in the presence of ribavirin or favipiravir. Confluent monolayers of RAW264.7 cells were infected with MNV-1 (C) or MNV-3 (D) at an MOI of 5 TCID50/cell. Infected cell cultures were treated with 200 μM ribavirin (RBV) or favipiravir (FPV) as explained in ‘Materials and methods’. Replication kinetics of MNV-1 and MNV-3 in untreated infected cells are shown in parallel (DMEM). Every time point is the average of three biological replicates (±SD).
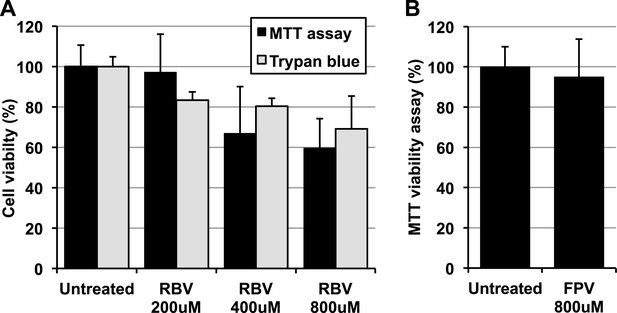
Favipiravir and ribavirin toxicity in RAW264.7 cells.
(A) Ribavirin toxicity upon RAW264.7 cells was scored using both trypan blue, which measures the proportion of dead cells, and CellTiter-Blue Cell Viability Assay (Promega), which accounts for living active cells. (B) Favipiravir toxicity was determined by CellTiter-Blue Cell Viability Assay (Promega).
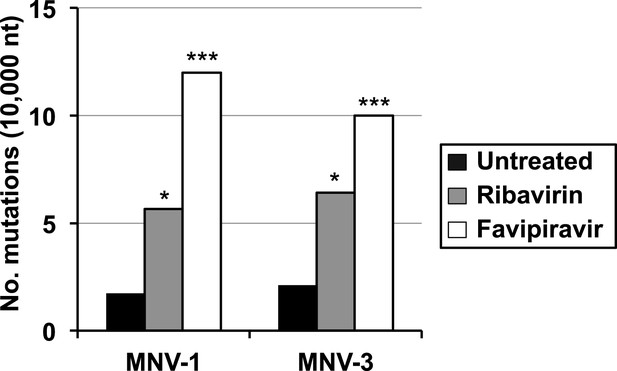
Increased mutation frequencies in virus quasispecies treated with ribavirin or favipiravir.
Mutation frequencies are represented as the average number of mutations found every 10,000 nucleotides sequenced in MNV-1 and MNV-3 populations after 4 passages in RAW264.7 cells in the absence or presence of 200 μM ribavirin (RBV) or favipiravir (FPV) (p < 0.05, *; p < 0.001, ***, Mann–Whitney U test).
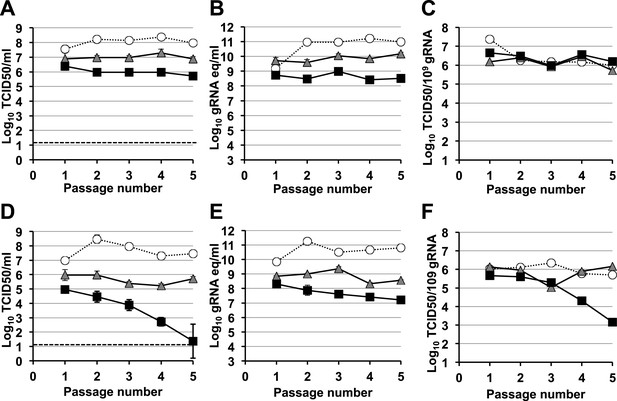
Murine norovirus titres decrease during serial passage in the presence of ribavirin.
MNV-1 (A, B, C) and MNV-3 (D, E, F) were serially passaged in the absence (white circles) or presence of 200 (grey triangles) or 400 μM ribavirin (black squares). Virus was inoculated at an MOI of 0.1 TCID50/cell in passage 1. Subsequent passages were carried out with 200 μl (1/10 vol) of neat virus recovered from the previous passage. The different graphs show the resulting virus titres (A and D), genome copy equivalents (B and E), and the resulting specific infectivity determined for encapsidated genomes (C and F). Specific infectivity values were calculated as the number of infectious viruses (TCID50 units) found in 109 genome copies from data obtained in A, B, D, and E. To calculate the number of genome copy equivalents, non-encapsidated genomes were removed before RNA extraction by micrococcal nuclease treatment.
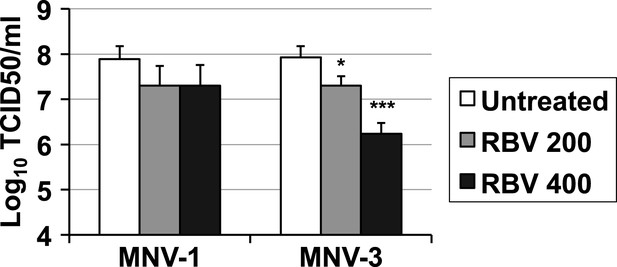
Reduced virus yields in ribavirin-mutagenised populations.
RAW264.7 cells were infected with MNV-1 or MNV-3 recovered after 5 passages in the absence (untreated) or presence of ribavirin (200 or 400 μM) with the exception of 400 μM ribavirin-treated MNV-3 where passage 2 was used due to insufficient virus titres in subsequent passages. Infections were performed at an MOI of 0.01. After adsorption (1 hr at 37°C), virus inoculum was removed and complete media added to infected cultures. Infected cells were then incubated as mentioned in ‘Materials and methods’ for a total of 17 hr before freezing.
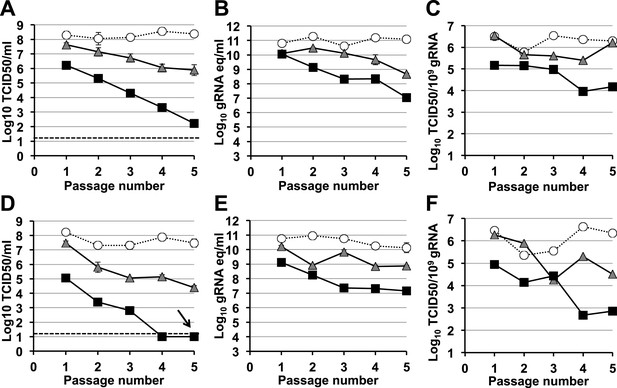
Murine norovirus infectivity decreases during serial passages in the presence of favipiravir.
MNV-1 (A, B, C) and MNV-3 (D, E, F) were serially passaged in the absence (white circles), or presence of 200 (grey triangles) or 400 μM favipiravir (black squares). Passage 1 infections were carried out at an MOI of 0.1 TCID50/cell. Subsequent passages were carried out with 200 μl (1/10 vol) of neat virus recovered from the previous passage. The different graphs show the resulting virus titres (A and D), genome copy equivalents (B and E), and the resulting specific infectivity determined for encapsidated genomes (C and F). Specific infectivity values were calculated as the number of infectious viruses (TCID50 units) found in 109 genome copies from data obtained in A, B, D, and E. To calculate the number of genome copy equivalents, non-encapsidated genomes were removed before RNA extraction by micrococcal nuclease treatment. The arrow in D indicates virus extinction confirmed by the absence of infectious virus and viral RNA (qPCR) after three serial passages in the absence of favipiravir.
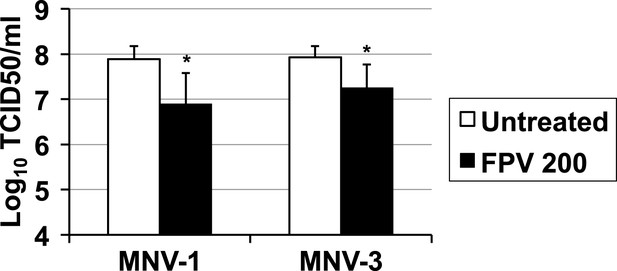
Reduced virus yields in favipiravir-mutagenised populations.
RAW264.7 cells were infected with MNV-1 or MNV-3 recovered after 5 passages in the absence (untreated) or presence of favipiravir (200 μM) Infections were performed at an MOI of 0.01. After adsorption (1 hr at 37°C), virus inoculum was removed and complete media added to infected cultures. Infected cells were then incubated as mentioned in ‘Materials and methods’ for a total of 17 hr before freezing.
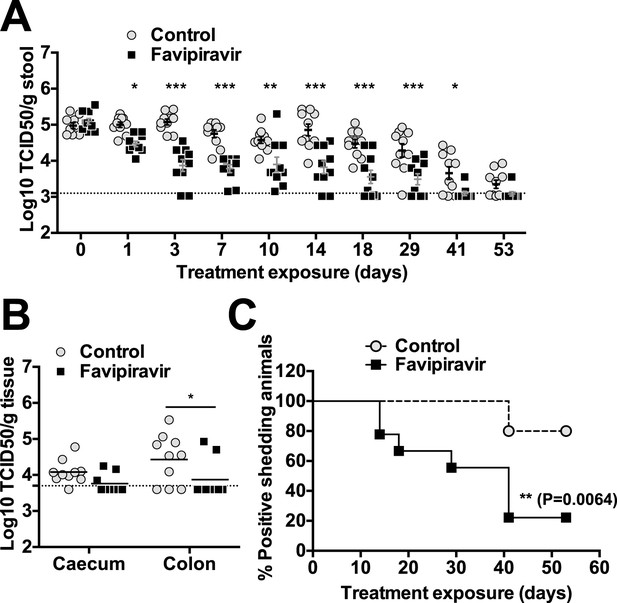
Favipiravir reduces infectious norovirus titres in mice faeces and tissues.
Two groups of ten 4–5-week old C57BL/6 male mice were oral gavage-infected with 104 TCID50 units of MNV-3. 4 weeks after virus inoculation, persistently infected animals were subjected to treatment with either 300 mg/kg animal of favipiravir twice a day (FPV) or with buffer (Control) for 8 weeks. From day 35 onwards, there are nine animals instead of 10 in the favipiravir-treated group (due to the accidental death of one mouse during dosing). (A) Virus titres in faeces of animals untreated or treated with favipiravir. Virus titres were determined by TCID50 assays of faecal samples supernatant previously resuspended at 100 mg/ml in PBS (p < 0.05, *; p < 0.01, **; p < 0.001, ***; 2-way ANOVA test). A dashed line indicates the limit of detection (3.02 Log10 TCID50/g stool). (B) Virus titres in caecum and colon of animals after 53 days of treatment with favipiravir. Virus titres were determined by TCID50 assays of homogenates of caecum and colon resuspended in DMEM at a concentration of 30 mg/ml (p < 0.05, *; 2-way ANOVA test). (C) Reduced positive shedding in animals treated with favipiravir. The percentage of animals shedding detectable virus titre along time, based on A, decreases faster in animals treated with favipiravir than in untreated animals (p = 0.0064, log-rank test).
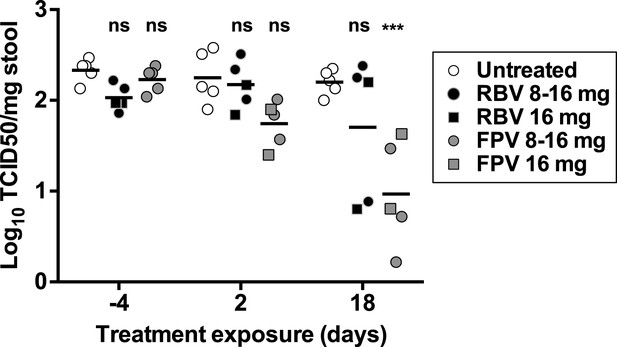
Antiviral activity in vivo of favipiravir and ribavirin.
4–5-week old male C57BL/6 mice were oral gavage-infected with 104 TCID50 units of MNV-3. 4 weeks after infection, animals underwent daily treatment with mutagenic compounds for a total of 18 days (day 0 to day 17), with the exception of days 6 and 13 when the animals were not treated. Animals were dosed by oral gavage with 8 mg/day (3 animals) or 16 mg/day (2 animals) of either ribavirin or favipiravir for 12 days. Afterwards, the animals were treated with 16 mg/day of the same drug for six additional days. Virus titres obtained in faecal samples isolated 4 days before treatment began (−4) and after 2 and 18 days of treatment (***; p < 0.001, 2-way ANOVA test).
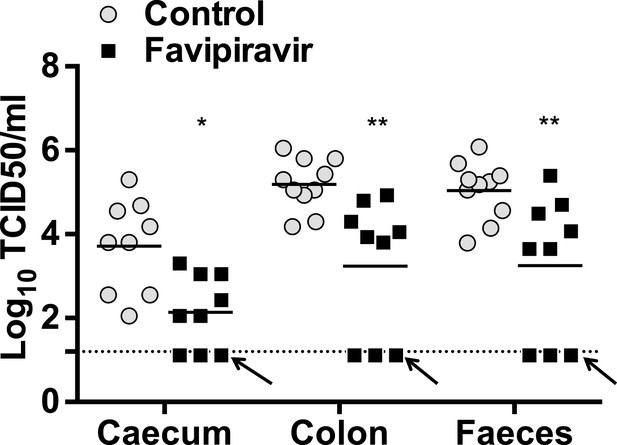
Infectious virus rebound after blind passage of faecal and tissue sample homogenates in RAW264.7 cells.
Virus titres obtained after blind passage infection in RAW264.7 cells of faeces, caecum, and colon homogenates obtained from animals at treatment day 53 (Figure 5A,B). An arrow indicates samples from the same three animals that remained negative after blind passage, indicating extinction. These samples remained negative by TCID50 assay and qPCR after three consecutive blind passages.
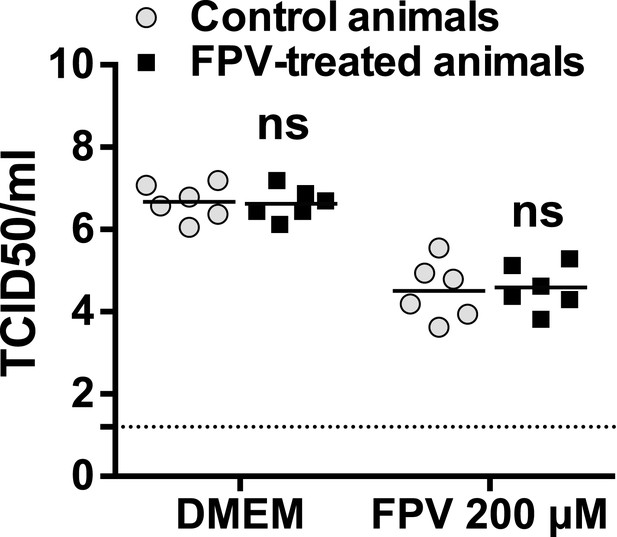
Sensitivity to favipiravir of virus isolated in animal faeces.
Faecal virus samples obtained from untreated or treated mice during 53 days were amplified by blind passage in RAW264.7 cells and treated with favipiravir to examine the possible presence of resistance.
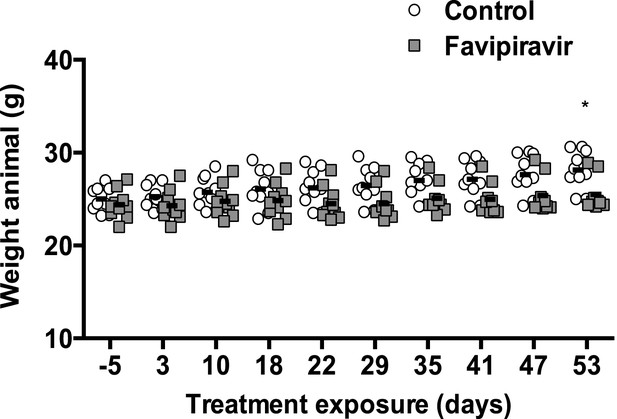
Reduced weigh increase in favipiravir-treated mice.
They are represented animal weights before (−5) and during the treatment with placebo or favipiravir (*; p < 0.05; 2-way ANOVA test).
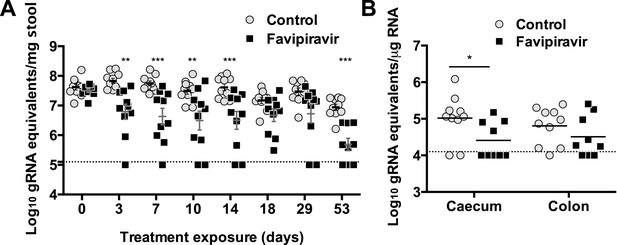
Long exposure to favipiravir results in significantly decreased norovirus RNA levels in animal tissues and faeces.
Two groups of ten male C57BL/6 mice of 4–5-weeks were oral gavage-infected with 104 TCID50 units of MNV-3. 4 weeks after virus inoculation, persistently infected animals were subjected to treatment with either 300 mg/kg animal of favipiravir twice a day (FPV) or with buffer (Control) for 8 weeks. At day 53, there are nine animals instead of ten in favipiravir-treated group due to the accidental death of one mouse during dosing. A dashed line indicates the limit of detection (102 genome copy equivalent per mg of stool). (A) Viral genome copy equivalents isolated in faecal samples (p < 0.05, *; p < 0.01, **; p < 0.001, ***; 2-way ANOVA test). Viral RNA extracted was then reverse transcribed and quantitated as described in ‘Materials and methods’. (B) Viral genome copy equivalents isolated in caecum and colon (p < 0.05, *; 2-way ANOVA test).
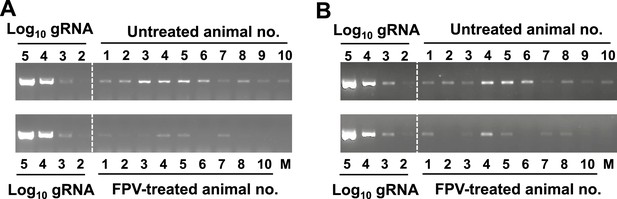
Semi-quantitative analysis of viral RNA in favipiravir-treated and control mice.
Viral RNA was extracted and reverse transcribed as described in ‘Materials and methods’ (Arias et al., 2012a). Standard curve was obtained with known amounts of in vitro transcribed MNV-3 RNA (102 to 105 genome equivalents) that were PCR amplified after reverse transcription. (A) RT-PCR amplification of faecal samples isolated from animals treated during 53 days with favipiravir or placebo. RNA isolated in 50 μg of faeces was RT-PCR amplified and agarose-gel resolved as explained above. (B) RT-PCR amplification of RNA extracted from colon samples isolated from animals treated for 8 weeks with favipiravir or placebo. 50 ng of RNA isolated from different animal colon tissues were RT-PCR amplified as explained above.
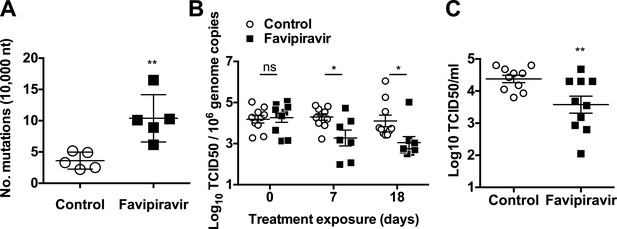
Increased mutation frequencies and decreased infectivity in virus populations isolated from favipiravir-treated animals.
(A) Mutation frequency in virus isolated in faecal samples. Every value in the graph represents the virus mutation frequency in a different animal faecal sample after 18 days of treatment. Mutation frequencies are represented as the average number of nucleotide substitutions found in every 10,000 nucleotides sequenced. (0.001 < p < 0.05, **; t test). (B) Decreased infectivity in viral RNA isolated from favipiravir-treated animals. Viral RNA isolated from placebo and favipiravir-treated animal faecal samples was quantified and 2 × 105 genome copy equivalents were lipofected in semiconfluent BHK-21 cell monolayers. At 24 hr post-transfection, cells were freeze-thawed and the resulting virus yields determined by TCID50 assays in RAW264.7 cells. They are represented as the virus titres obtained per 106 genome copies isolated from the faeces of infected animals before treatment (day 0) and at treatment days 7 and 18. (C) MNV recovered from favipiravir-treated animals shows reduced replication yields. Virus isolated from animal faeces treated with favipiravir were first amplified in RAW264.7 cells allowing virus replication for 24 hr. Recovered viruses were titrated and 0.01 TCID50 units/cell applied to new RAW264.7 cell monolayers. Virus infections were collected at 8 hr post-infection and the cultures freeze-thawed to release infectious virus (**; p < 0.01; t test).
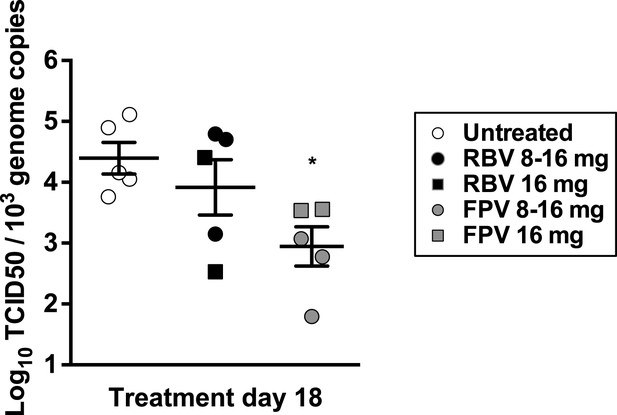
Specific infectivity of viral RNA isolated from treated animals.
4–5-week old male C57BL/6 mice were oral gavage-infected with 104 TCID50 units of MNV-3. 4 weeks after infection, animals underwent daily treatment with mutagenic compounds during a total of 18 days (day 0 to day 17) with the exception of days 6 and 13 when the animals were not treated. Animals were dosed by oral gavage with 8 mg/day (3 animals) or 16 mg/day (2 animals) of either ribavirin or favipiravir for 12 days. Afterwards, the animals were treated with 16 mg/day of the same drug for six additional days. Viral RNA isolated from placebo-, ribavirin-, and favipiravir-treated animal faecal samples were quantified and 5 × 104 genome copy equivalents lipofected in semiconfluent BHK-21 cell monolayers. At 24 hr post-transfection cells were freeze-thawed and resulting virus yields determined by TCID50 assays in RAW264.7 cells. They are represented the infectivity values for RNA samples isolated from faeces collected at day 18.
Tables
Mutation type distribution in MNV populations treated with ribavirin and favipiravir
| Untreated | Ribavirin | Favipiravir | ||||
|---|---|---|---|---|---|---|
| MNV-1 | MNV-3 | MNV-1 | MNV-3 | MNV-1 | MNV-3 | |
| A → G | 4 | 4 | 1 | 3 | 9 | 5 |
| U → C | 1 | 3 | 1 | 2 | 3 | 7 |
| G → A | 0 | 2 | 5 | 2 | 2 | 0 |
| C → U | 4 | 1 | 4 | 4 | 0 | 4 |
| Transversions | 1 | 3 | 1 | 2 | 1 | 0 |
| Deletions | 1 | 0 | 1 | 0 | 0 | 0 |
| Total nucleotides sequenced* | 63,395 | 65,822 | 23,358 | 20,243 | 12,488 | 16,003 |
-
*
Total number of nucleotides sequenced in each different untreated or treated population analysed.
-
Proportion of different types of mutations observed in untreated, or ribavirin- or favipiravir-treated MNV populations.





