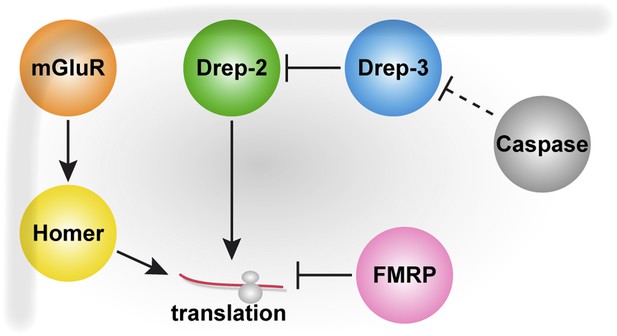
Drep-2 is a novel synaptic protein important for learning and memory
Figures
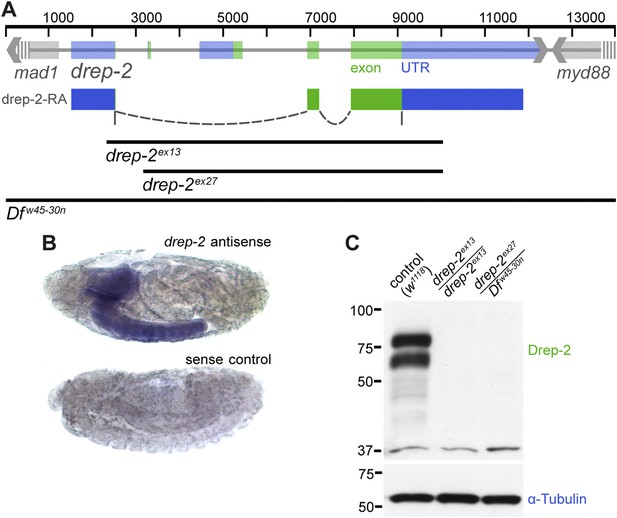
Expression and mutants of drep-2.
(A) Genetic scheme of the drep-2 locus on chromosome IIR. The neighboring genes mad-1 and myd88 extend beyond the sequence displayed. The cDNA labeled drep-2-RA was used for rescue experiments. Blue: untranslated regions; green: exons; black lines: deleted regions in the mutants. (B) In situ hybridization of drep-2 reveals a neuronal expression pattern (stage 17). (C) Western blot of adult fly head extracts using the anti-Drep-2C-Term antibody. Drep-2 isoforms are predicted to run at 52 and 58 kDa. The signal is absent in both the drep-2ex13 and the drep-2ex27/Dfw45-30n mutant.

Drep protein alignment.
Sequence alignment of all four Drosophila Dff proteins, as well as human (HS) and murine (MM) Dff40. Drep-4 has the strongest similarity to Dff40, yet Drep-2 also shows conserved motifs in addition to the CIDE-N domain. The alignment was created using Geneious v5.3.6. (http://www.geneious.com)
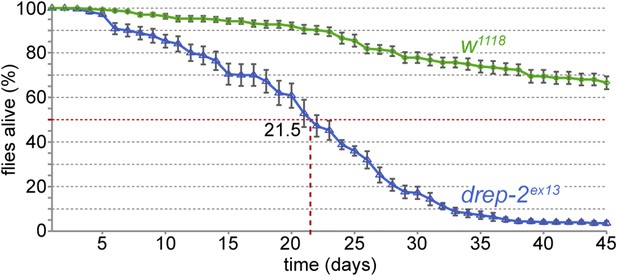
Reduced lifespan of drep-2ex13 mutants.
Comparison to isogenic w1118 control flies: 50% of mutant flies were dead after 21.5 days. Mutant: n = 10 vials (each containing 25 flies), control: n = 11.
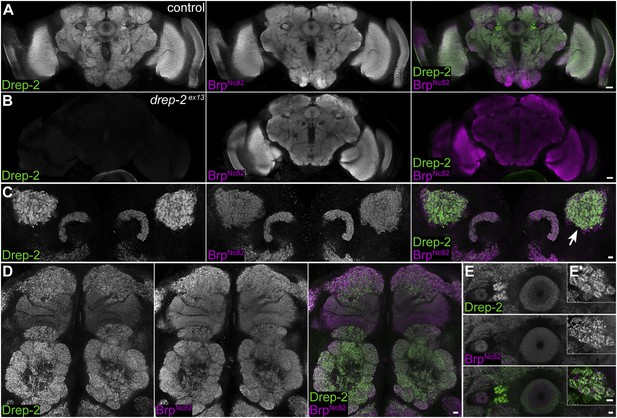
Synaptic Drep-2 staining in the CNS.
(A–B) Confocal frontal sections of adult Drosophila brains. Anti-Drep-2C-Term and BrpNc82 immunostaining; the latter marks all synaptic active zones. Synaptic Drep-2C-Term signal is visible throughout the brain of wild-type flies (A). Complete loss of the anti-Drep-2C-Term staining can be observed in drep-2ex13 mutants (B). Scale bars: 20 µm. (C–E) Frontal sections of wild-type brains, anti-Drep-2C-Term, and BrpNc82 staining. Scale bars: 5 µm. (C) Posterior–dorsal detail showing strong Drep-2 staining in MB calyces (arrow). (D) Anterior frontal section with antennal lobes and MB lobes. (E) Ellipsoid body in the central complex and bulbs (lateral triangles) (E′: magnification of strong Drep-2 staining in bulbs).
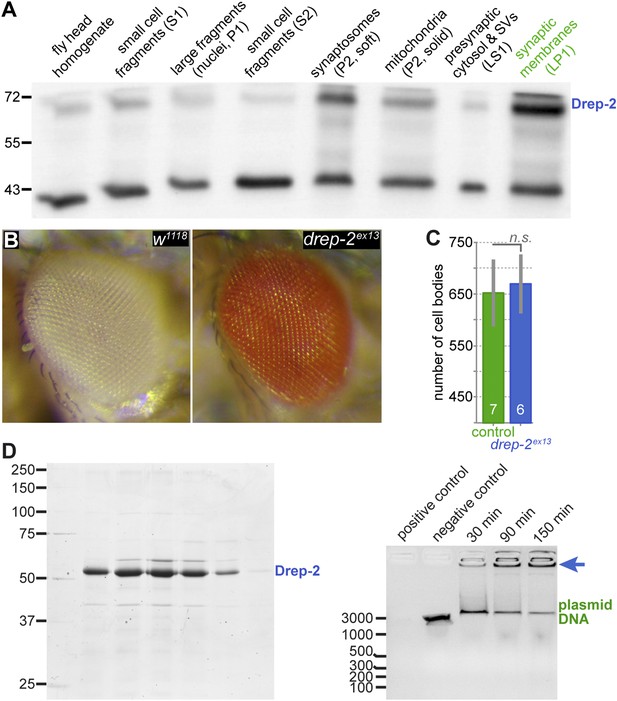
No evidence for a role of Drep-2 in regulation of apoptosis.
(A) Synaptosome-like preparation of adult wild-type head extracts (Depner et al., 2014), probed with Drep-2C-Term. Drep-2 is concentrated in fractions containing synaptic membranes. S = supernatant, P = pellet, L = (after) lysis. Please see the protocol by Depner et al. (2014) for a more detailed explanation of the fractionation procedure. (B) Mutants (drep-2ex13) did not show a rough eye phenotype. The facet eyes of flies, highly ordered structures, are often affected in apoptosis mutants. By contrast, the eyes of drep-2 mutants appeared normal. (C) The number of mb247-positive KCs does not differ between drep-2ex13 mutants and controls. GFP was expressed using the MB KC driver mb247-Gal4. GFP-positive cell bodies were counted and compared between genotypes. No significant difference was found between mean cell body counts (Mann–Whitney U test, p = 0.886). Average cell body counts were in the expected range: control = 651, mutant = 669, published = 700 (Schwaerzel et al., 2002). (D) Purified Drep-2 does not degrade linearized plasmid DNA. Left: SDS-PAGE of the final elusion profile of purified Drep-2, loaded onto a HighLoad Superdex S200 16/60 column. Right: Nuclease activity assay of purified Drep-2 analyzed by 1% (wt/vol) agarose gel. Drep-2 was incubated in a time course experiment with linearized plasmid DNA. No nuclease activity could be detected. Instead, Drep-2 seemed to precipitate DNA, as evidenced by high-molecular DNA not entering into the agarose gel when incubated with Drep-2 (arrow).
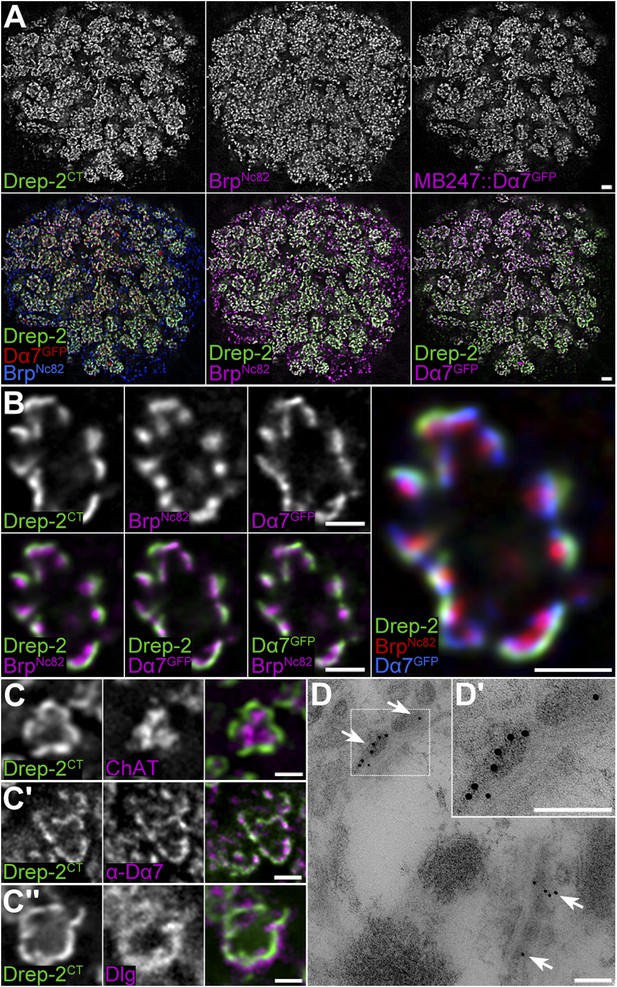
Drep-2 is enriched at KC postsynapses.
(A–B) Drep-2C-Term and BrpNc82 staining in animals expressing the construct mb247::Dα7GFP that marks acetylcholine receptors in MB KCs. (A) Detailed image of the MB calyx. Scale bar: 2 µm. (B) Detail of a single microglomerulus in the calyx. Drep-2C-Term overlaps with postsynaptic mb247::Dα7GFP and not with presynaptic Brp. Scale bars: 1 µm. (C) Localization of Drep-2 relative to choline acetyltransferase (ChAT, presynaptic cytosol, C), the postsynaptic ACh receptor subunit Dα7 (antibody staining, C′), and the postsynaptic scaffolding protein Discs large (Dlg, C″). Drep-2 colocalizes with postsynaptic markers. Scale bars: 1 µm. (D) Post-embedding immunoelectron microscopy of Drep-2C-Term in the calyx. Arrows: Clusters of postsynaptic Drep-2C-Term. Scale bars: 100 nm.
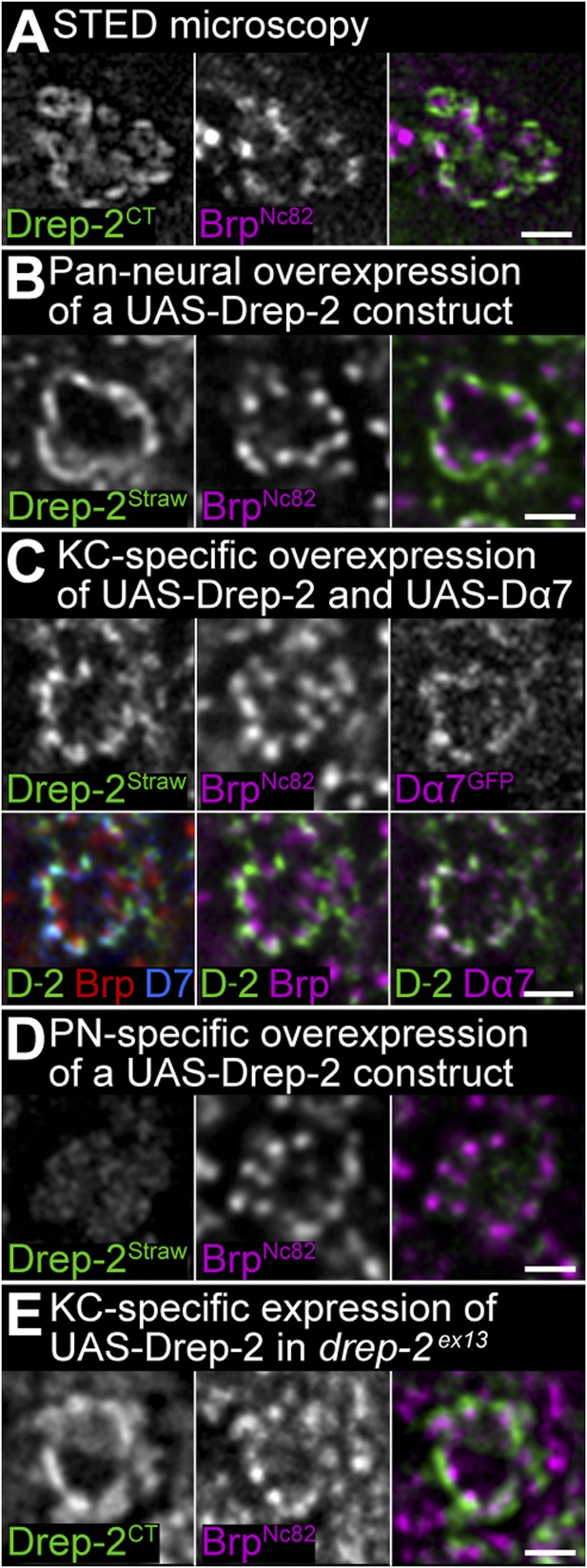
Drep-2 localizes to postsynaptic membranes of KCs in the calyx.
(A) STED microscopy superresolution recording of Drep-2C-Term; the BrpNc82 channel is in normal confocal mode. The Drep-2 signal does not overlap with presynaptic Brp. Scale bar: 1 µm. (B–E) Expression of drep-2 constructs in KCs yields a label resembling the Drep-2 antibody staining. Comparison to BrpNc82; all scale bars: 1 µm. (B) Pan-neural overexpression. Elavc155-Gal4 and UAS-Drep-2mStrawberry; mStrawberry signal is shown. (C) KC-specific overexpression. C305a-Gal4, UAS-Drep-2mStrawberry, and UAS-Dα7GFP; mStrawberry and GFP signals are shown. D-2 = Drep-2mStrawberry, D7 = Dα7GFP. (D) PN-specific overexpression. Gh146-Gal4 and UAS-Drep-2mStrawberry; diffuse mStrawberry is shown. (E) KC-specific expression of UAS-Drep-2 in the drep-2ex13-mutant background. Mb247-Gal4 and untagged UAS-Drep-2; Drep-2C-Term staining is shown.
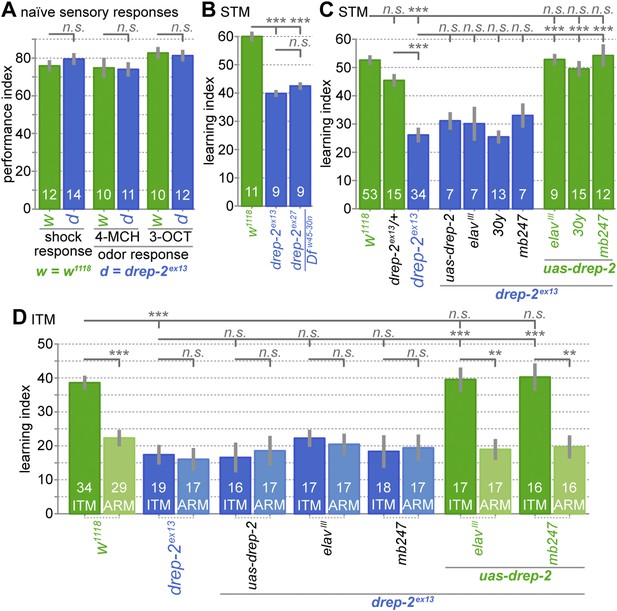
Drep-2 is required in KCs for olfactory short- and intermediate-term memory.
(A) Flies mutant for drep-2 sense electric shock and the odors 4-methyl-cyclohexanol (4-MCH) and 3-octanol (3-OCT) normally; there is no difference in mean performance indices between mutants and isogenic w1118 control flies (Mann–Whitney U tests (MWU)). Sample sizes n are indicated with white numbers; grey bars show SEMs. (B) Both mutants drep-2ex13 and drep-2ex27/Dfw45-30n are deficient in aversive olfactory conditioning, 3 min STM in a T-maze. The graph shows mean learning indices and SEMs. Mutants performed significantly worse than isogenic controls (MWU: p = 0.00001 for both comparisons, Bonferroni-corrected significance level α = 0.0167, 3 tests). (C) Re-expression of drep-2 cDNA with elavIII-Gal4 (pan-neural), 30y-Gal4 (MB KCs), or mb247-Gal4 (MB KCs) restores the deficit to normal levels. Heterozygous drep-2ex13 mutants do not display a significant deficit. MWU for individual comparisons showed a significant difference between these groups (α = 0.0042, 12 tests): w1118 and drep-2ex13 (p < 0.00001), drep-2ex13/drep-2ex13 and drep-2ex13/+ (p < 0.00001), drep-2ex13 and drep-2ex13;uas-drep-2/elavIII-gal4 (p < 0.00001), drep-2ex13 and drep-2ex13;uas-drep-2/30y-gal4 (p < 0.00001), drep-2ex13 and drep-2ex13;uas-drep-2/mb247-gal4 (p < 0.00001). None of the differences indicated as not significant had a p < 0.12, except for w1118 and drep-2ex13/+ (p = 0.03851; not significant in the case of α = 0.0042). (D) Intermediate-term memory (ITM = ASM + ARM) performance. Mutants (drep-2ex13) are defective in ASM, but not in ARM. The defect can be restored with elavIII-Gal4 or mb247-Gal4 (30y-Gal4 was not used here). Statistical tests were run separately for ITM and ARM. For ITM, MWU for individual comparisons showed a significant difference between these groups (α = 0.00625, 8 tests): w1118 and drep-2ex13 (p < 0.0001), drep-2ex13 and drep-2ex13;uas-drep-2/elavIII-gal4 (p < 0.0001), drep-2ex13 and drep-2ex13;uas-drep-2/mb247-gal4 (p < 0.0001). For assessing differences in ARM, ITM and ARM performances of each genotype were compared with MWU. The following genotypes showed a significant difference between ITM and ARM (α = 0.0071, 7 tests): w1118 (p < 0.0001), drep-2ex13;uas-drep-2/elavIII-gal4 (p = 0.0002), drep-2ex13;uas-drep-2/mb247-gal4 (p = 0.0006). None of the differences indicated as not significant had a p < 0.11.
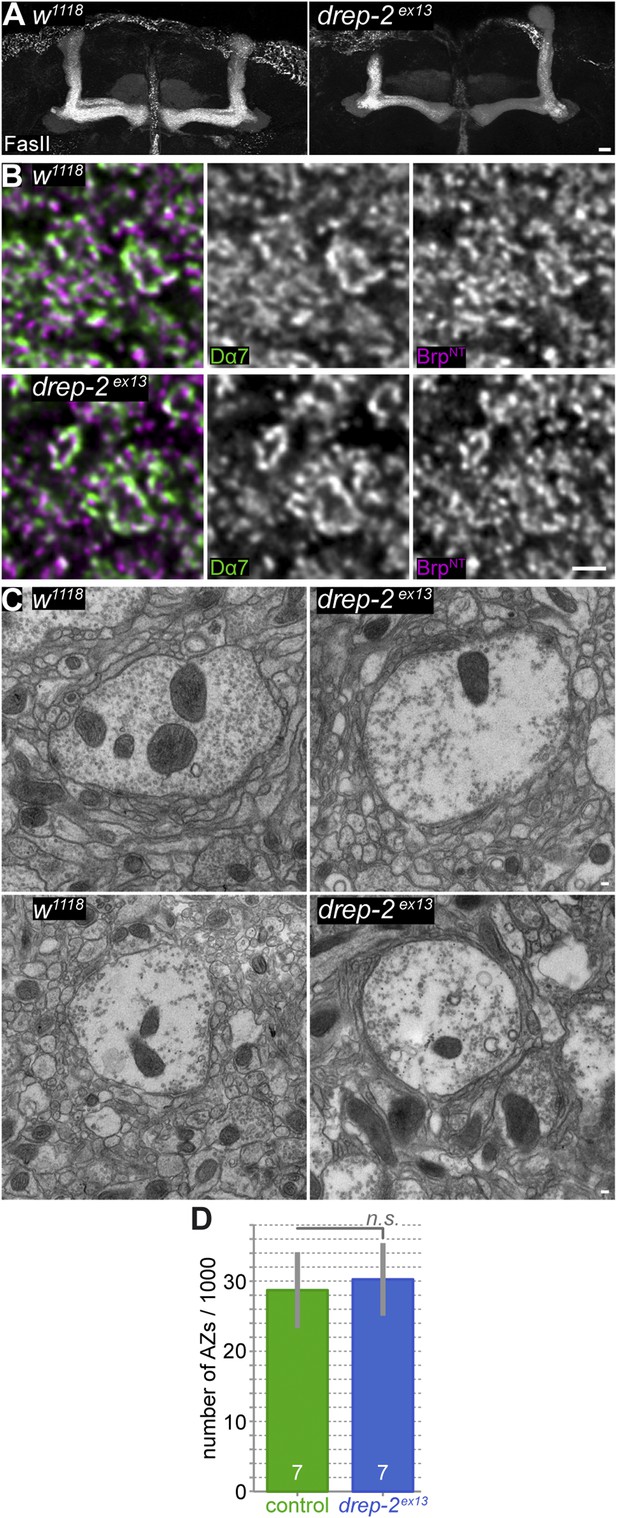
PN-KC synapses appear morphologically normal in drep-2 mutants.
(A) Absence of major neuroanatomical defects in drep-2ex13 mutant brains. MB lobes, Fasciclin II (FasII) staining, maximum intensity projections. Scale bar: 10 μm. (B) Antibody staining of w1118 control and drep-2ex13 mutant brains, using antibodies against the postsynaptic ACh receptor subunit Dα7 and presynaptic BrpN−Term. Focus on microglomeruli of PN-KC synapses in the MB calyx. Microglomeruli of mutants appear structurally normal. Scale bar: 1 µm. (C) Electron microscopy of w1118 control and drep-2ex13 mutant brains. Microglomeruli and postsynaptic KC profiles of mutants appear structurally normal. Scale bars: 100 nm. (D) The number of synapses (active zones) in the MB calyx does not significantly differ between drep-2ex13 mutants and w1118 controls. Syd-1-positive spots were counted and compared between genotypes as described (Kremer et al., 2010). No significant difference was found between the number of spots (MWU, p = 0.62). Average synapse counts were in the range expected (28,000–30,000 [Kremer et al., 2010]).
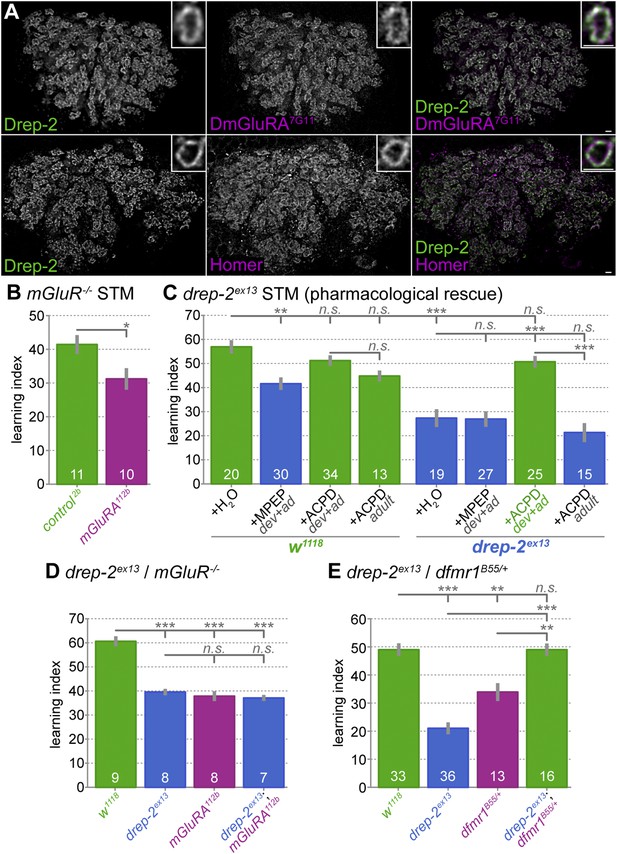
Functional overlap between Drep-2 and mGluR in olfactory conditioning.
(A) Wildtype adult MB calyces stained with Drep-2C-Term and DmGluRA7G11 (first row) or with Drep-2C-Term and anti-Homer (second row). Drep-2 colocalizes tightly with both proteins. The insets show single microglomeruli. Scale bars: 2 µm. (B) Flies carrying the mutation dmGluRA112b are deficient in aversive olfactory conditioning STM when compared to isogenic dmGluRA2b controls that do express DmGluRA; MWU: p = 0.043, α = 0.05. The graph shows mean learning indices and SEMs; sample sizes n are indicated with white numbers. (C) The drep-2ex13 phenotype in olfactory STM can be rescued by raising animals on food containing the DmGluRA agonist 1S,3R-ACPD (ACPD). Food was supplemented throughout development and adulthood with either the DmGluRA receptor antagonist MPEP (9.7 µM) or the agonist ACPD (72.2 µM) diluted in H2O (label: dev+ad). Control animals received only H2O. One group of animals was transferred to food supplemented by ACPD only after eclosion and not during development; the corresponding experiments are indicated by the label +ACPD adult. MPEP lowered the w1118 performance significantly (MWU p = 0.0003). MPEP did not alter drep-2ex13 indices (p = 0.8772) and ACPD did not change w1118 performance (p = 0.1145). ACPD rescued the drep-2 mutant phenotype to control levels if fed during both development and adulthood (comparison of drep-2ex13 +ACPD dev+ad to untreated drep-2ex13: p < 0.00001; comparison to untreated w1118: p = 0.0945). ACPD did not rescue the mutant phenotype if fed only during adulthood (+ACPD adult, no significant difference to untreated drep-2ex13 (p = 0.2281), significant difference to mutants treated with ACPD during both development and adulthood (p < 0.00001)). The difference between untreated w1118 and drep-2ex13 flies was also significant (p < 0.00001). Significance level α = 0.005 (10 tests). (D) Phenotypes of drep-2ex13; dmGluRA112b double mutants were non-additive. Both drep-2ex13 and dmGluRA112b single mutants showed significantly lower olfactory STM than isogenic controls (MWU, p = 0.00008 for both comparisons). Double mutants showed similar learning indices (comparison to w1118: p = 0.00018). The two single mutants and the double mutant did not significantly differ from each other (p > 0.178). α = 0.0083 (6 tests). (E) Loss of drep-2 antagonizes dfmr1 phenotypes in olfactory conditioning STM. Both homozygous drep-2ex13 mutants and heterozygous dfmr1B55/+ mutants are deficient in olfactory learning STM, but double mutants carrying both alleles do learn. The graph shows mean learning indices and SEMs. MWU for individual comparisons (α = 0.01, 5 tests): w1118 and drep-2ex13 p < 0.00001, w1118 and dfmr1B55/+ p = 0.00069, w1118 and drep-2ex13; dfmr1B55/+ p = 0.83751, drep-2ex13 and drep-2ex13; dfmr1B55/+ p < 0.00001, dfmr1B55/+ and drep-2ex13; dfmr1B55/+ p = 0.00071.
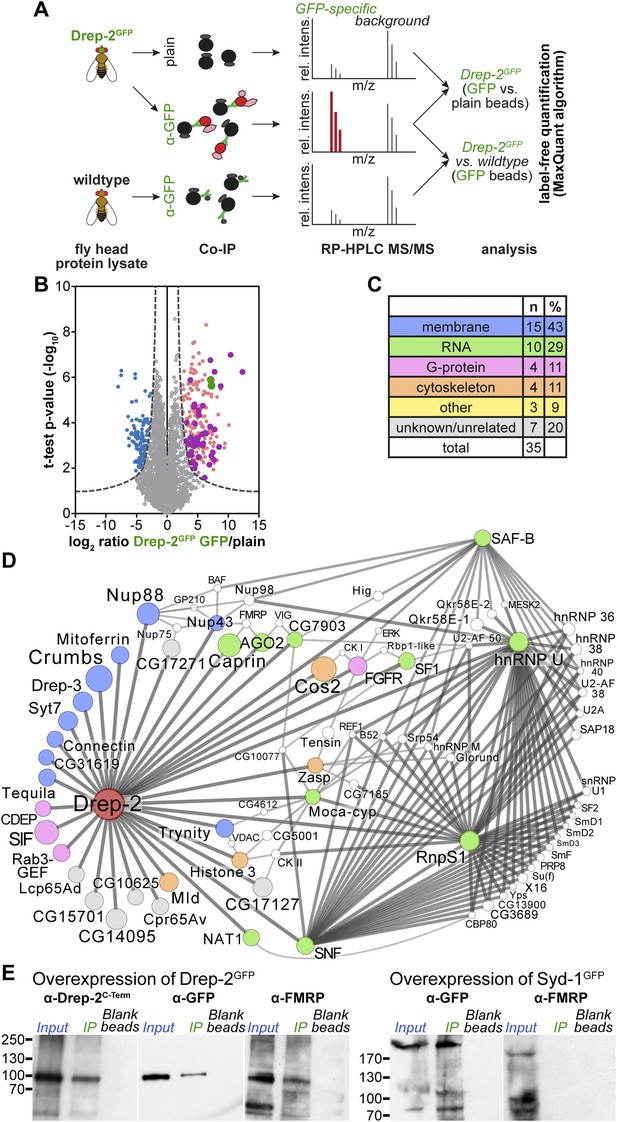
Quantitative mass spectrometry: Drep-2 and FMRP were found in a common protein complex.
(A) Strategy for the identification of Drep-2 interactors by quantitative mass spectrometry. UAS-Drep-2GFP was overexpressed using the pan-neural driver line elavc155-Gal4. (B) Volcano plot showing proteins from Drep-2GFP flies binding to anti-GFP and/or plain control beads. A hyperbolic curve (set at an FDR of 1%) separates GFP-enriched proteins (light pink) from background (grey). Proteins enriched in the control are shown in blue. Proteins that were significantly enriched, both in Drep-2GFP flies and in independent control experiments with wild-type flies, are colored magenta (n = 35). Drep-2 and GFP are shown as green dots. (C) Classification of the 35 core network proteins; multiple counts were allowed. (D) Network of the 35 proteins that were significantly and reproducibly enriched in GFP pulldown experiments (at an FDR of 1%, magenta-colored dots in B). Additional putative interactors of the core network (FDR set at 10%) are shown in white (Supplementary file 2). The circle (node) and font size correspond to the rank within the results (indicated in Supplementary files 1 and 2). The line (edge) width and shade correspond to the number of interactions each of the significantly enriched proteins has with others. The line/edge length is arbitrary. (E) Anti-FMRP probing confirmed the specific presence of FMRP in Drep-2GFP complexes. Head extracts of flies expressing Drep-2GFP or the presynaptic protein Syd-1GFP were processed in parallel. FMRP was only enriched in preparations of Drep-2GFP extracts. Immunoprecipitations were performed using either GFP-Trap-A beads (lanes labeled IP) or blocked agarose beads as binding control (labeled Blank beads).
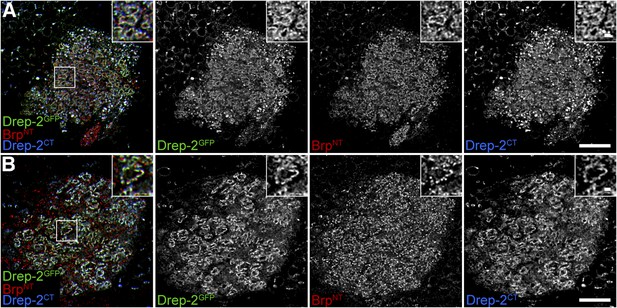
Drep-2GFP colocalizes with endogenous Drep-2.
(A) Pan-neural overexpression of UAS-drep-2GFP by elavc155-Gal4. MB calyx stained with anti-GFP, BrpN-Term and Drep-2C-Term. The Drep-2GFP label does not differ from the Drep-2C-Term antibody staining, compare also to Figure 4 and Figure 4—figure supplement 1. Scale bars: 10 µm and 1 µm (insets). (B) KC-specific expression of UAS-drep-2GFP by mb247-Gal4 in drep-2ex13 mutants. Staining and scale bars as in (A).
Additional files
-
Supplementary file 1
Mass spectrometry: core proteins enriched over both controls. The table shows proteins enriched at an FDR of 1%. Ranks are based on the ratios of Drep-2GFP IPs, GFP beads vs. plain beads (compare Figure 7A–B). Putative functions were derived from Flybase. GFP (#6) was removed from this list.
- https://doi.org/10.7554/eLife.03895.015
-
Supplementary file 2
Mass spectrometry: all proteins enriched at an FDR of 10%. The table shows two ratios for all proteins: first, Drep-2GFP animals, GFP beads vs. plain beads and, second, GFP beads, Drep-2GFP vs. wild-type (wt) animals (compare Figure 7A). “+” symbols indicate whether a protein was significantly enriched at a given FDR. Proteins are sorted according to the first ratio. Proteins were labeled in cyan if significantly enriched for both ratios at an FDR of 1%; these proteins constitute the core interactors. Protein names were labeled in red if the protein was positively enriched for the first ratio but negatively enriched for the second one, and thus probably constitutes a false positive. Protein names were labeled in shades of green if the protein was included as an additional interactor in the network of interacting proteins (Figure 7D): dark green = FDR 1%, medium green = FDR 5%, light green = FDR 10%.
- https://doi.org/10.7554/eLife.03895.016