Thermodynamic evidence for a dual transport mechanism in a POT peptide transporter
Figures
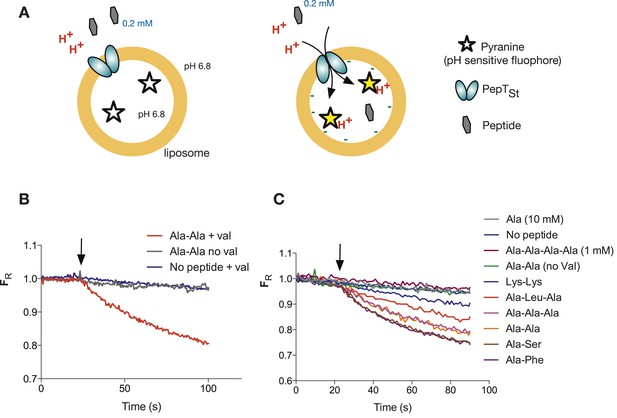
Monitoring peptide-coupled proton transport using the pH sensitive dye, pyranine.
(A) Experimental setup to monitor proton flux. PepTSt is reconstituted into liposomes loaded with pyranine and a high concentration of potassium ions (120 mM), the external solution contains peptide and a low potassium concentration (1.2 mM). On addition of valinomycin, the membrane becomes highly potassium permeable, generating a hyperpolarised membrane potential (negative inside) this drives the uptake of peptide with protons, protonating the pyranine dye and altering its fluorescent properties. (B) Representative pyranine fluorescence traces produced from the set up described in (A) indicating that acidification of the liposomal lumen only occurs in the presence of a valinomycin (black arrow)-induced membrane potential with peptide. The Y axis indicates the fluorescence ratio as stated in the methods. (C) PepTSt can only transport di- and tri-peptides. To initiate transport, valinomycin was added to all experiments at the time indicated by the black arrow and the external substrate concentration was 0.2 mM. Data were normalised to the first time point for ease of comparison.
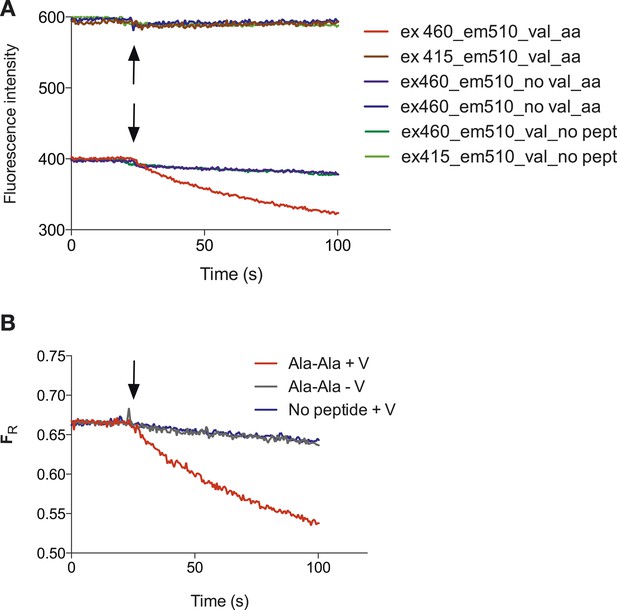
Representative raw data of the pyranine fluorescence traces.
Raw data produced from the set up described in (Figure 1A) indicating that acidification of the liposomal lumen only occurs in the presence of a valinomycin (black arrow)-induced membrane potential with peptide, this data were used to make Figure 1B. (B) Shows the data from (A) but shown as a ratio of the spectra obtained at 460/415.
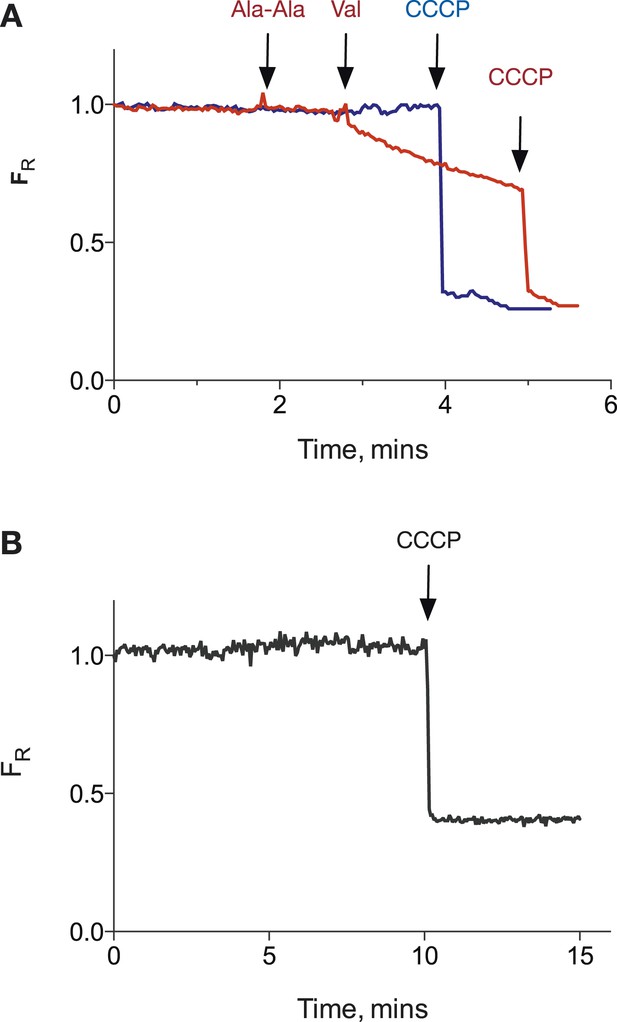
PepTSt POPE:POPG proteoliposomes can hold a pH gradient of 1 unit.
(A) Potential proton leakage was monitored using pyranine in a system with internal pH at 6.8 and external pH at 5.8. The proton gradient was collapsed by the addition of CCCP. (B) as (A) but internal pH was 7.5 and external pH was 6.5.
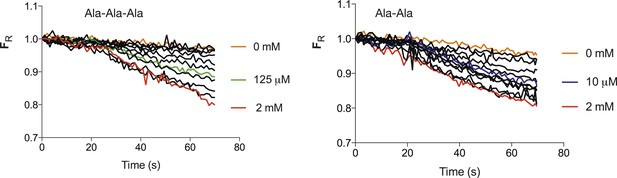
Transport strength of Ala–Ala vs Ala-Ala-Ala.
Increasing concentrations of peptide both Ala-Ala-Ala (Left) and Ala-Ala (right) lead to an increase in the fluorescence change until a maximum level is reached. The mid point of the fluorescence change between maximum decrease and that with no peptide for Ala-Ala-Ala is ∼125 µM and 10 µM for Ala–Ala, an approximate 10-fold difference. (Concentrations used for tri-ala were 0, 0.1, 1, 5, 10, 25, 50, 125, 250, 500, 1000, 2000 µM and di-ala 0, 0.01, 0.1, 1, 5, 10, 25, 50, 100, 250, 1000, 2000 µM).
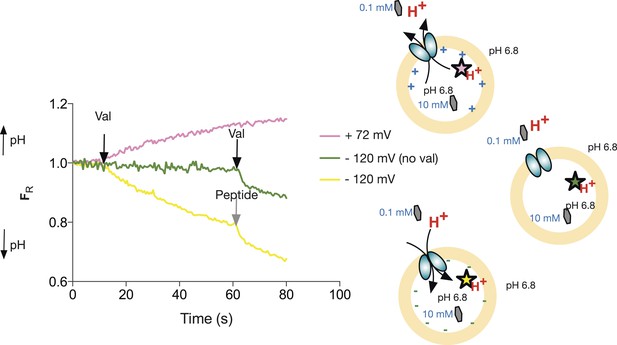
Peptide transport depends on the imposed voltage.
Proteoliposomes in the presence of a 100-fold peptide gradient (0.1 mM outside and 10 mM inside) and no pH gradient (pH 6.8 both inside and outside). No proton flux occurs until the transmembrane potential is shunted by addition of valinomycin (green trace, val added at black arrow). PepTSt can drive transport of peptides (and protons) against this gradient into the interior of the liposome using the proton electrochemical gradient when a negative voltage is imposed by valinomycin addition. (negative inside—yellow line). Further addition of peptide (0.5 mM) results in additional uptake (grey arrow). When a large (inside) positive voltage is applied (same peptide and pH gradients), peptides (and protons) exit the liposome (pink line).
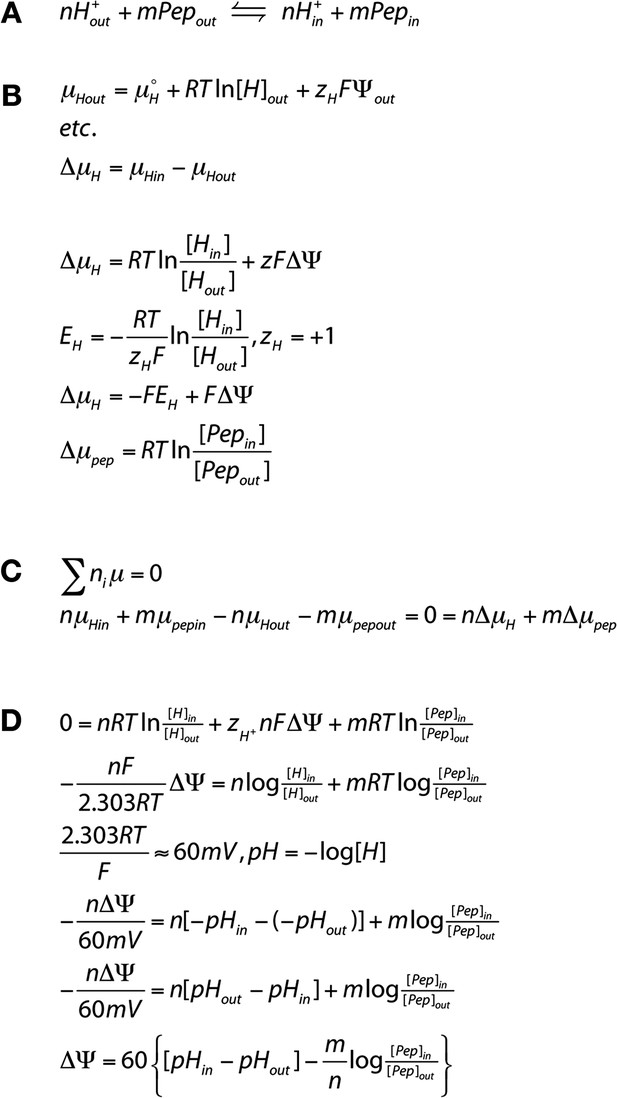

Derivation of the transport equation for a proton peptide transporter.
We assume that transport occurs with a fixed stoichiometry as reflected in (A). Pep indicates the neutral peptide and the subscripts, ‘out’ and ‘in’ refer to peptide or protons outside or inside the liposome membrane, ‘n’ is the number of protons transported per cycle and ‘m’ is the number of peptides. We seek to determine n/m the stoichiometric ratio of protons:peptide. For the reaction shown in (B), μ is the chemical potential of the species, R is the universal gas constant, T is the temperature in (K), F is the Faraday constant, ZH is the proton charge, ΔΨ is the voltage difference across the membrane, where ΔΨ = Ψin − Ψout. This is equivalent to the sign convention that the outside of the liposome is defined as ground (Ψout = 0). At equilibrium (C), the equation can be rearranged (D) to yield the equilibrium potential, ΔΨ, in terms of the values of pH and [peptide] and the stoichiometric ratio.
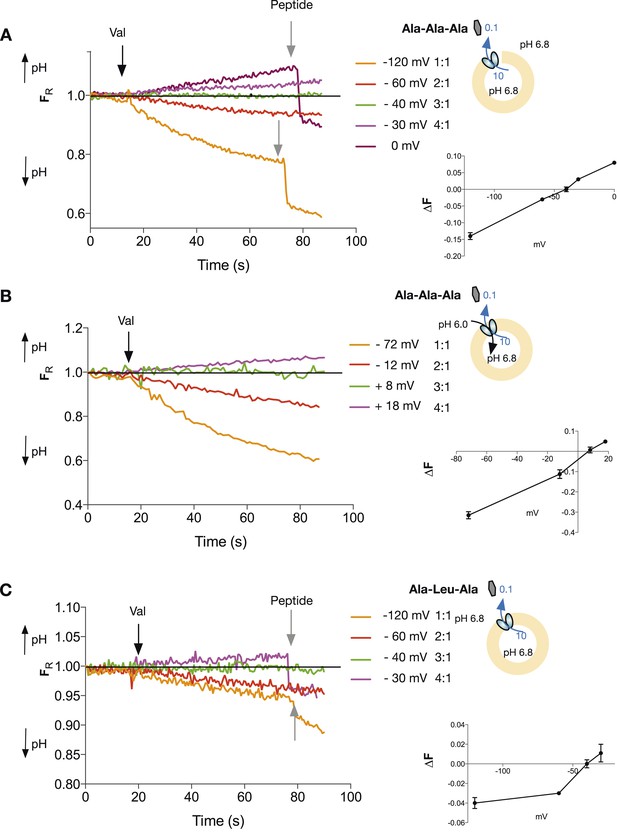
Tri-peptides are co-transported with three protons.
The potassium gradient across the liposomes was varied in order to set the desired voltages (on valinomycin addition, black arrow) to achieve no net proton movement (indicated by the black line at FR of 1.0). The number next to the voltages are the proton:peptide stoichiometry that would reverse at that voltage. The internal peptide concentration (10 mM) (Tri-ala for A and B, and Ala-Leu-Ala for C) was 100-fold above that of the external concentration and the pH was inside 6.8, outside 6.8 (for A and C) and 6.0 (for B). Representative traces are shown for each experiment, which were repeated at least three independent times. The line graph for each experiment represents the mean change in fluorescence at time point 60 s and S.E.M. is indicated. Grey arrows indicate the addition of more peptide, to show that the system can be further manipulated.
-
Figure 3—source data 1
Table showing the conditions used to calculate the stoichiometry of transport.
- https://doi.org/10.7554/eLife.04273.010
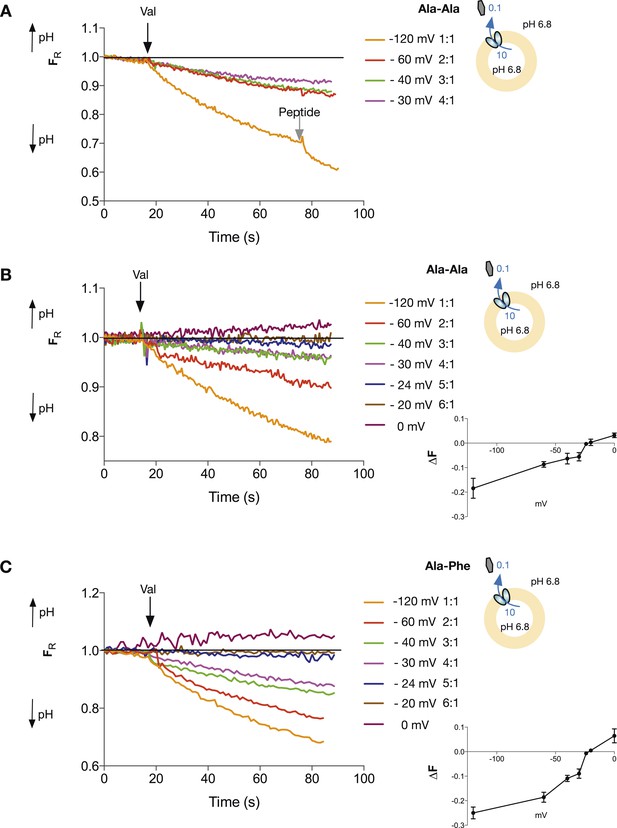
Di-peptide transport requires more protons than tri-peptide.
The potassium gradient across the liposomes was varied in order to set the desired voltages (on addition of valinomycin, black arrow) to achieve no net proton movement (indicated by the black line at FR of 1.0). Only voltages that indicate a stoichiometry of proton:peptide of greater than 5:1 showed either no net movement of protons or reversal for both Ala–Ala (A, B) and Ala–Phe (C) di-peptides. Representative traces are shown for each experiment, which was repeated at least three independent times. The line graph for each experiment represents the mean change in fluorescence at time point 60 s and S.E.M. is indicated. Grey arrows indicate the addition of more peptide, to show that the system can be further manipulated.
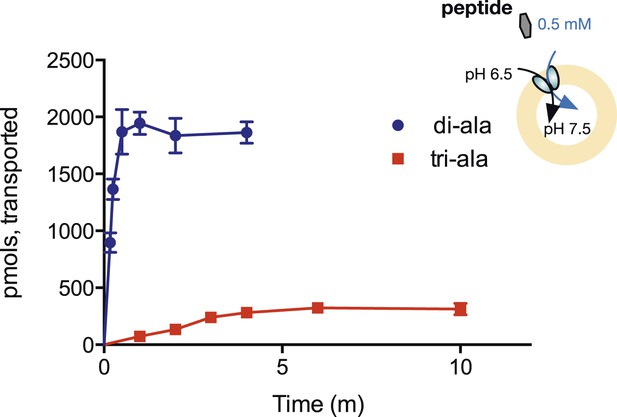
Steady-state accumulation of di- vs tri-alanine.
Peptide transport was driven by an inwardly directed proton gradient in saturating amounts of peptide. Uptake was measured via scintillation counting using radiolabeled peptides (3H for di-alanine and 14C for tri-alanine) and converted to pmols peptide transported per unit time.
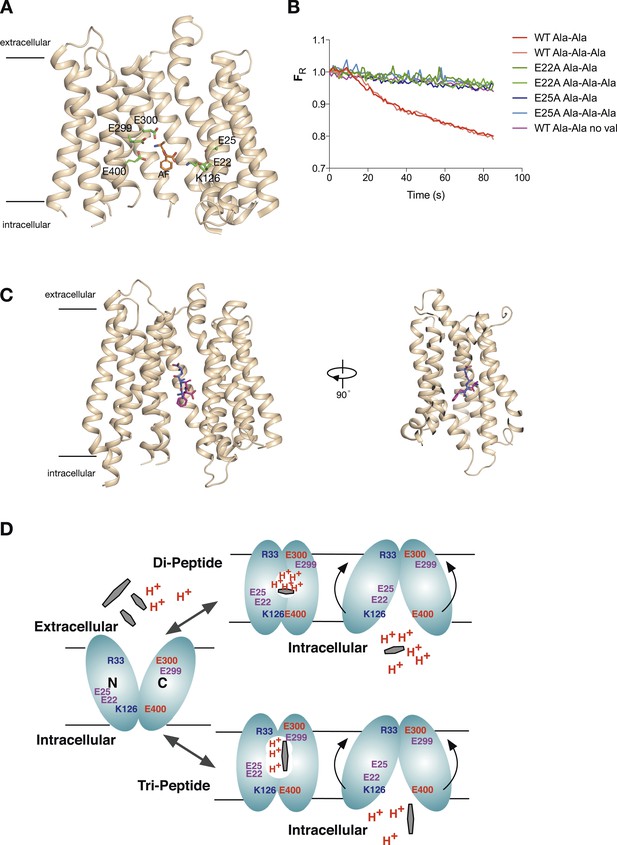
Model for proton:peptide symport.
(A) Crystal structure of PepTSt with bound Ala–Phe peptide (orange) (PBD 4D2C) showing the protonatable side chains within the binding site (green). Helices TM5 and TM8 have been removed for clarity. (B) E22A and E25A variants of PepTSt are unable to couple proton movement to the transport either di-alanine or tri-alanine. (C) Ala–Phe and Ala-Ala-Ala adopt different orientations within the binding site of PepTSt. Two views of PepTSt shown in the plane of the membrane and rotated 90°, with Ala-Ala-Ala (blue) and Ala–Phe (magenta) shown as sticks. Helices TM5 and TM8 have been removed for clarity. (D) Model for proton:peptide symport in PepTSt. Di-peptides transport requires at least four protons, whereas tri-peptides require only three, suggesting this is the lowest number of protons required to drive the conformational changes required for alternating access transport. Essential residues are indicated; residues involved in the potential stabilising salt bridges are labelled in blue and red, whereas protonatable side chains are labelled in purple.
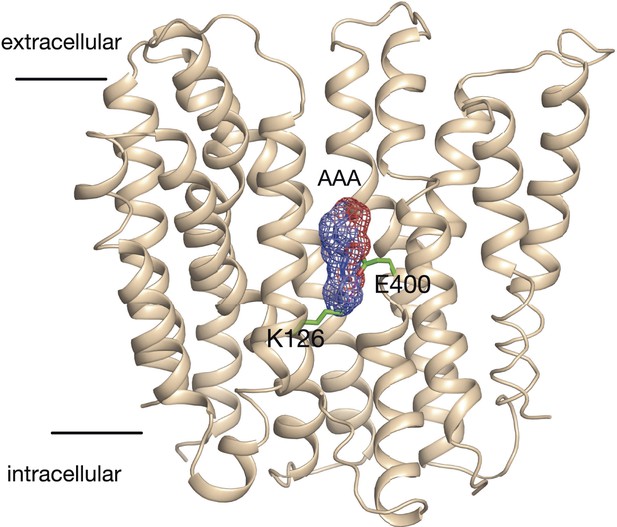
Model of the outward facing state of PepTSt with bound Ala-Ala-Ala.
The position of the tri-alanine peptide (mesh) would obstruct the formation of the predicted salt bridge between residue Lys126 and Glu400.





