Response to Nodal morphogen gradient is determined by the kinetics of target gene induction
Figures
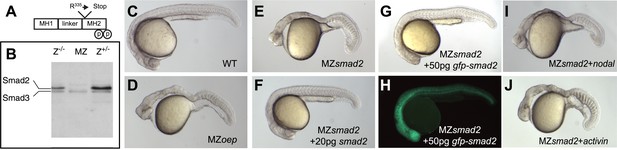
Maternal Smad2 is necessary for mesendoderm specification by Nodal signaling.
(A) Illustration of the Smad2 protein showing the position of the ENU-induced non-sense mutation. (B) Western blot against Smad2/3 on 24 hpf embryos of different genotypes for smad2. MZ, maternal-zygotic homozygotes, Z−/−, zygotic homozygotes, Z+/−, zygotic heterozygotes. The pool of maternally contributed Smad2 protein persists for at least 24 hr in zygotic homozygous embryos while it is depleted in MZsmad2 mutants. (C–J) Phenotypic analysis of 36 hpf zebrafish embryos. (C) Wild-type embryo. (D) MZoep embryo: maternal-zygotic mutant for one-eyed pinhead (oep), a cell surface protein required for Nodal signaling (Gritsman et al., 1999). (E) MZsmad2 embryo. Msmad2 mutants display a very similar phenotype (not shown). (F) MZsmad2 embryo rescued with 20 pg of smad2 mRNA. (G–H) MZsmad2 embryo rescued with 50 pg of gfp-smad2 mRNA (brightfield (G), epifluorescence (H)). smad2 mRNA appears to be more effective in rescuing the prechordal plate defects in MZsmad2 mutants as compared to gfp-smad2 mRNA. (I) MZsmad2 embryo injected with 5 pg mRNA for the zebrafish Nodal homolog squint. (J) MZsmad2 embryo injected with 5 pg mRNA for activin. Note that while Activin can activate the Nodal pathway in the absence of oep (Gritsman et al., 1999; Cheng et al., 2003), neither Squint nor Activin can activate the pathway in the absence of Smad2.
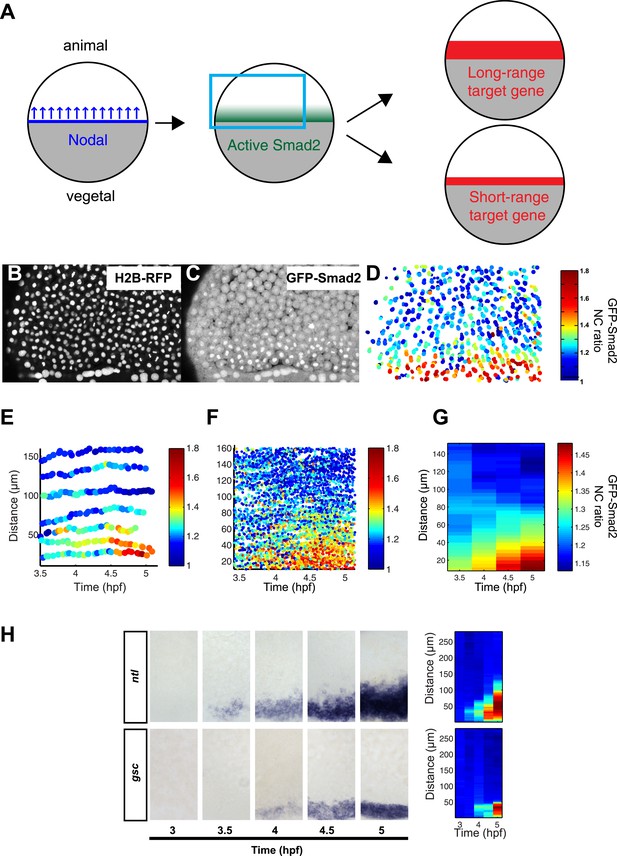
Dynamics of Nodal signaling in vivo.
(A) Illustration of Nodal signaling input–output relationship during blastula stage. Gray = yolk, white = blastoderm. Nodal is produced at the margin, diffuses and forms a gradient along the vegetal–animal axis. Nodal signaling induces a gradient of activated Smad2, which induces long- and short-range target gene expression. (B and C) Maximal intensity projection of a confocal stack of a Histone 2B-RFP (B), GFP-Smad2 (C) double transgenic embryo at 50% epiboly (blue box in (A)). GFP-Smad2 strongly accumulates in the nuclei of cells close to the margin, the source of Nodal signals. (D) Heatmap of the nucleo-cytoplasmic (NC) ratio of GFP intensity from the embryo in (B and C). Each dot represents the position of a cell (overlay of five consecutive frames, 3-min intervals per frame). Each cell is color-coded according to its GFP NC ratio (see Figure 2—figure supplement 2 for movement of cells). (E) Examples of single cell tracks at different locations along the vegetal–animal axis, showing changes in GFP-Smad2 NC ratio over time. The position of most cells relative to the margin remains constant during blastula stage. Cells close to the margin activate Nodal signaling earlier and at higher levels than cells at a distance from the margin. The short bursts observed in some cell tracks are caused by transient nuclear accumulation of GFP-Smad2 at the onset of nuclear envelope breakdown and are observed even in the absence of Nodal signaling. (F) NC ratio dynamics of tracked cells along the vegetal–animal axis. (G) Mean NC ratio values from (F) in 30 min bins. Note that the range and amplitude of the Smad2 activity gradient increase over the course of 90 min. Basal NC ratio is higher in younger embryos (see Figure 2G, 3.5 hpf). Since this phenomenon is also observed in the absence of Nodal signaling (MZoep mutants), the higher NC ratio is unlikely to reflect early Smad2 activation, but a higher nuclear import/export ratio of GFP-Smad2 during early development. (H) Time course of ntl (upper panel) and gsc (bottom panel) expression detected by RNA in situ hybridization. ntl begins to be induced as early as 3.5 hpf and its domain of expression expands over time to 100–120 µm from the margin; gsc begins to be induced 30 min later than ntl and its domain of expression expands to 50 µm from the margin. Close-up views of dorsal side, animal pole to the top. Right panel, heatmap for the grayscale intensity of in situ hybridization signals along the vegetal–animal axis showing the increase in range and intensity of ntl and gsc expression over time. See Figure 2—figure supplement 3 for comparison of probes and Figure 2—figure supplement 4 for independent validation of gsc and ntl expression domains using Seurat.
-
Figure 2—source data 1
Individual cell tracks and NC ratio.
See Supplementary file 1 for description.
- https://doi.org/10.7554/eLife.05042.006
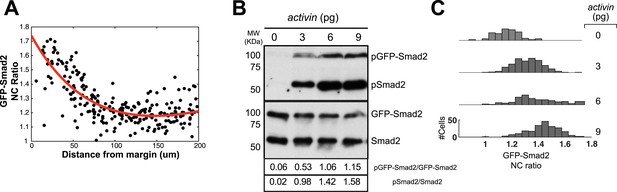
GFP-Smad2 as a sensor of Nodal activity in vivo.
(A) GFP-Smad2 NC ratio as a function of distance from the margin. Black dots represent individual cells and the thick red line shows a polynomial fit. (B) Dose response analysis of Smad2 and GFP-Smad2 phosphorylation by Western blot following increasing amounts of activin mRNA in MZoep mutant. MZoep embryos lack endogenous Nodal activity but ectopic Activin can activate the Nodal pathway in the absence of oep. Numbers indicate amounts of pGFP-Smad2 and pSmad2 relative to GFP-Smad2 and Smad2 signals, respectively. (C) GFP-Smad2 NC ratio distribution following increasing amounts of activin mRNA in MZoep background. Both Smad2/GFP-Smad2 phosphorylation and the GFP-Smad2 NC ratio increase as the Activin levels increase.
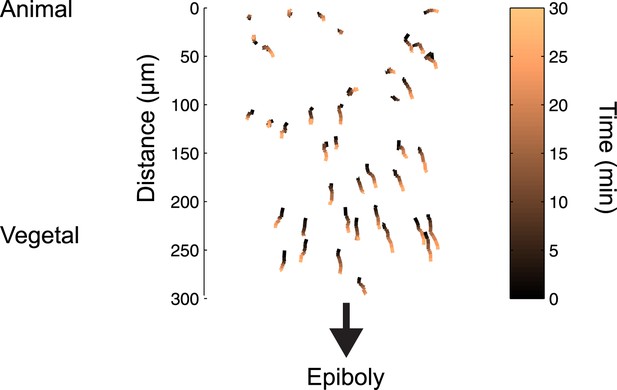
Cell movements during blastula stages.
Tracks of 50 random cells from the embryo shown in Figure 2D over a period of 30 min. Because of epiboly movements, cells move towards the vegetal pole, but their relative position from the margin barely changes (see Figure 2E).
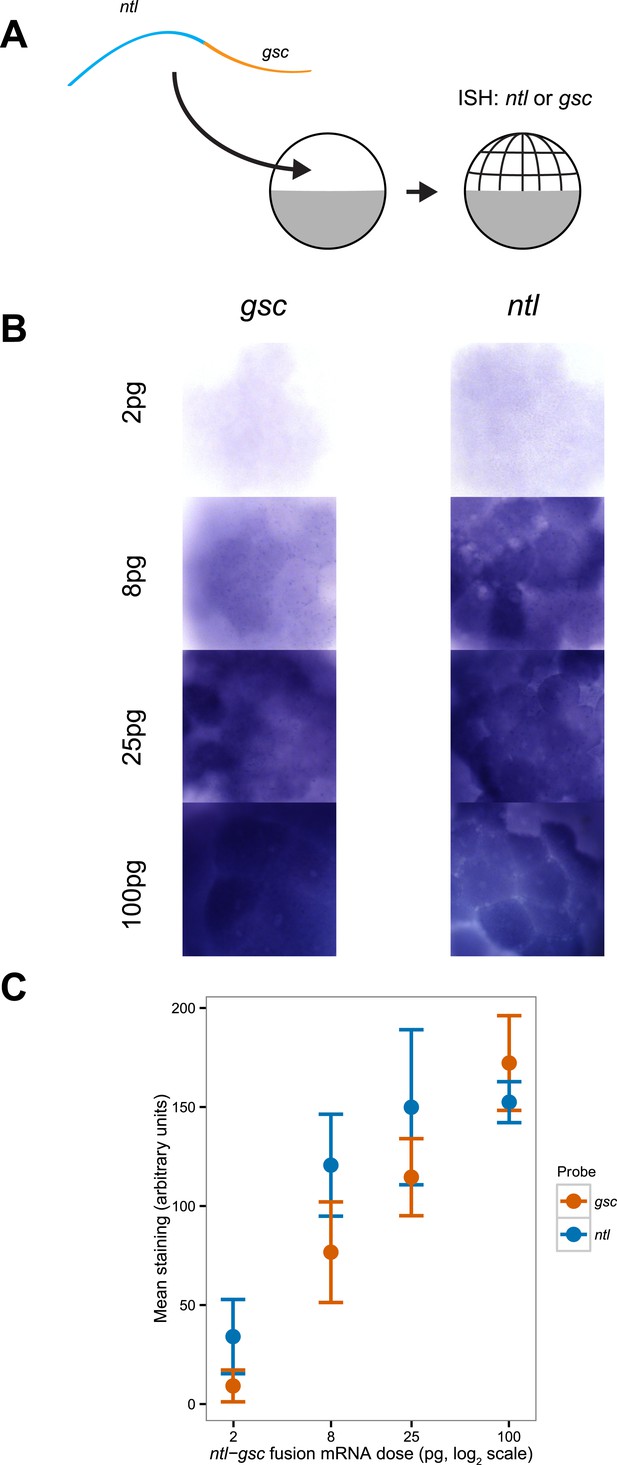
Detection sensitivity of ntl and gsc by in situ hybridization.
(A) Illustration of the experiment: a ntl-gsc fusion mRNA was injected at the one cell stage at four different concentrations (2, 8, 25 and 100 pg). Embryos were then fixed at the 128–256 cell stage and processed for in situ hybridization with either ntl or gsc probes. (B) Representative images of embryos hybridized with gsc (left) or ntl (right) probes at different concentrations. (C) Mean signal intensity and standard deviation as a function of injected fusion mRNA concentration for gsc and ntl (n = 5 embryos). The gsc probe is less sensitive at low concentrations. p = 0.03 at 2 pg, p = 0.03 at 8 pg, p = 0.12 at 25 pg, and p = 0.14 at 100 pg (Two-sample t-test).
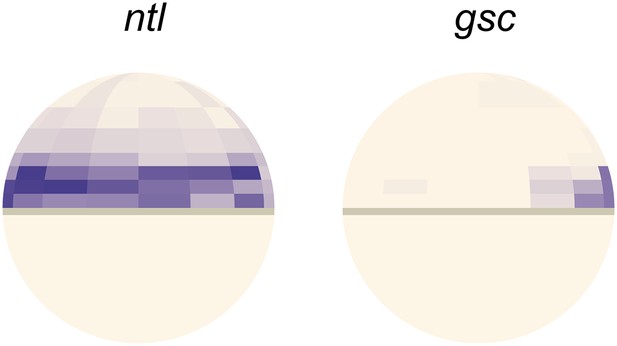
Comparison of ntl and gsc expression pattern from single-cell RNAseq analysis.
Gene expression patterns of ntl (left) and gsc (right) computed from single-cell RNAseq data spatially assigned to a 50% epiboly zebrafish embryo using Seurat (Satija et al., in press). Lateral view, dorsal side to the right.
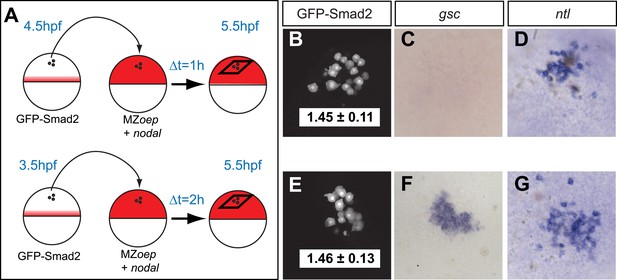
Testing the threshold model.
(A) Schematic of the transplantation experiment. Animal pole cells (black circles) from a GFP-Smad2 transgenic embryo were transplanted into the animal pole of a host embryo that had been injected with mRNA for squint, a zebrafish Nodal gene (red). Host cells were unresponsive to Nodal because they were maternal-zygotic mutants for one-eyed pinhead (MZoep), a cell surface protein required for Nodal signaling. This strategy prevents feedback loops and restricts target gene expression to donor cells. The developmental age of donor cells was matched to host embryos. Black parallelograms indicate imaging plane in subsequent panels. (B–G) Nodal signaling response of donor cells after 1 hr (B–D) or 2 hr (E–G) of exposure to Nodal. (B and E) Projection of confocal stacks of transplanted embryos and associated NC ratio (mean ± std). Activated Smad2 levels are similar in both cases. See Figure 3—figure supplement 1 for time course of GFP-Smad2 N/C ratio. (C and F) RNA in situ hybridization for gsc. (D and G) RNA in situ hybridization for ntl. ntl is expressed after 1 (n = 12/12) or 2 hr (n = 16/16) of Nodal exposure while gsc signal in transplanted cells is only detected after 2 hr of exposure (n = 1/15 at 1 hr, n = 12/14 after 2 hr). Images in B–G are from different embryos. Note that the differences in the duration of Nodal exposure uncouple the activated Smad2 level from target gene expression.
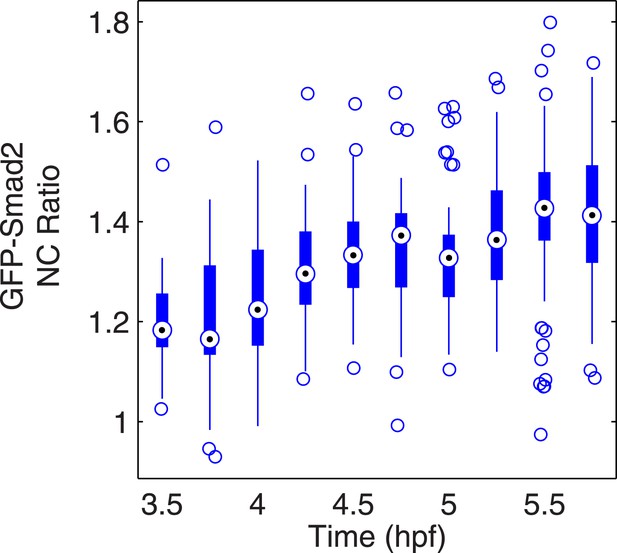
Time course of GFP-Smad2 NC ratio.
Boxplot of the NC ratio of GFP-Smad2 cells over time. GFP-Smad2 cells (n ∼50) were transplanted into a MZoep host embryo injected with 30 pg of squint mRNA (as in Figure 3A, bottom panel), and the NC ratio was determined at different time intervals. Note that the NC ratio is never higher at any time between 3.5 and 5.5 hpf than as compared to 5.5 hpf.
Testing the ratchet model.
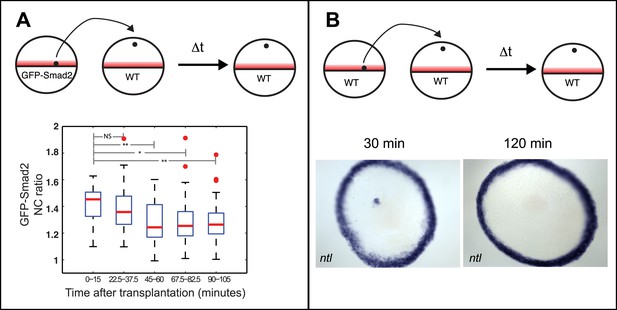
(A) Top: Illustration of transplantation experiments to move cells from a region with Nodal (red) to a region without Nodal. Cells located at the margin of a 30% epiboly GFP-Smad2 transgenic embryo were transplanted to the animal pole of a stage-matched wild-type embryo. The NC ratio of GFP-Smad2 was measured over time by time-lapse microscopy (n = 3 embryos). Bottom: boxplot of the NC ratio distribution at different time intervals after transplantation. Smad2 activity progressively decreases and reaches basal levels after 60 min. Two-sample t-tests p-values are indicated: NS, not significant; *p ≤ 0.05; **p ≤ 0.01. (B) Top: Illustration of the transplantation experiment. Bottom: RNA in situ hybridization for ntl 30 min (left) or 120 min (right) after marginal cells were transplanted to the animal pole. After 30 min, the majority of ectopically transplanted marginal cells express ntl (n = 12/14 transplantations) whereas 2 hr after transplantation, 90% of embryos are devoid of ectopic ntl expression (n = 17/19 transplantations). These results indicate that Nodal pathway activity cannot be maintained for prolonged periods in the absence of Nodal.
A kinetic model for Nodal morphogen interpretation.
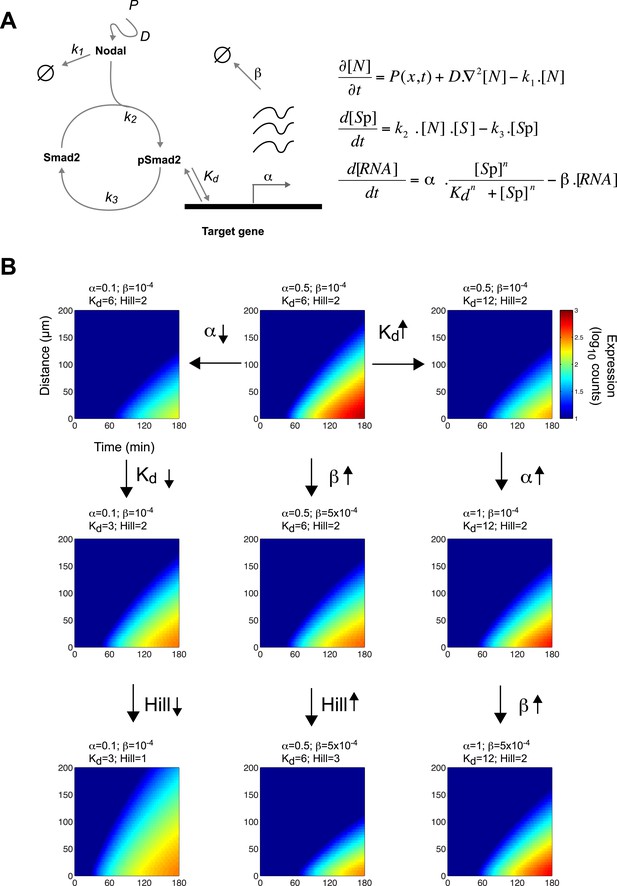
(A) Diagram of the Nodal signaling pathway used for modeling (left) and coupled differential equations describing the changes of Nodal, activated Smad2 and target genes over time (right). The Nodal ligand is locally produced, diffuses and via kinase receptors phosphorylates Smad2. Phosphorylated Smad2 acts as a transcription regulator and binds to target genes to induce transcription. (B) Spatiotemporal gene expression patterns were simulated over 3 hr using the kinetic model. Each panel depicts the expression pattern resulting from a unique combination of the four free parameters involved in mRNA production (transcription rate α, degradation rate β, dissociation constant Kd and Hill coefficient) while other parameters are held constant. Note how changes in these parameters change the range of target gene expression. See Figure 4—figure supplement 1 for more extensive simulations.
Screening for parameters regulating range and onset of target gene expression.
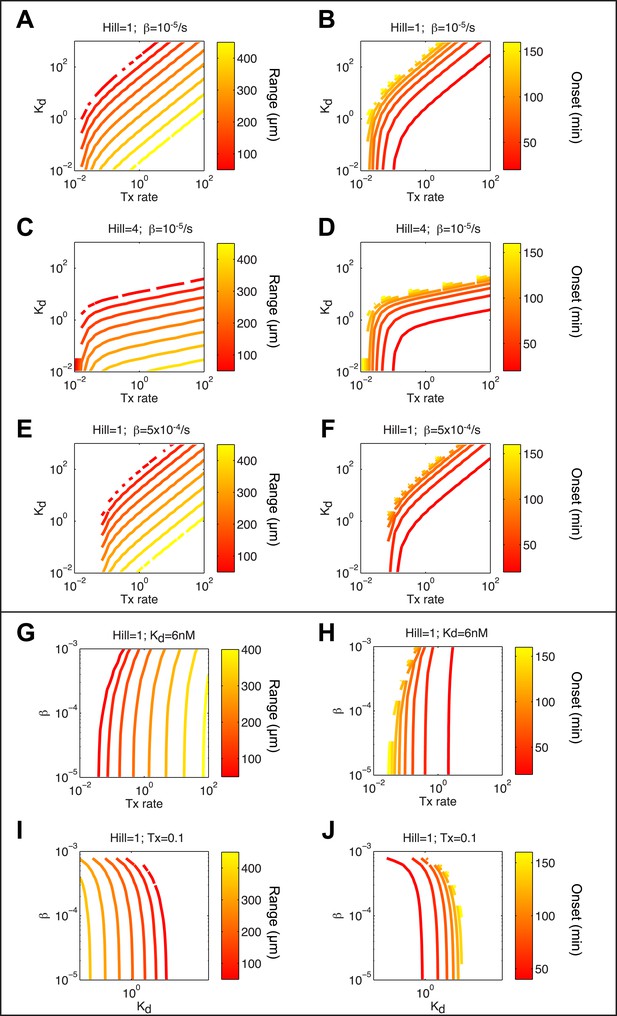
(A, C, E, G, I) Contour plots of maximum range of expression after 3 hr of signaling. (B, D, F, H, J) Contour plots of onset of expression. (A and B) Combined effect of Tx rate (x axis) and Kd (y axis) with a Hill coefficient of 1 and a degradation rate of 10−5 s−1. Tx rate and Kd contribute equally to the range of expression, except at low Kd and low Tx rate, where the range is more sensitive to changes in Tx rate. (C and D) Combined effect of Tx rate (x axis) and Kd (y axis) with a Hill coefficient of 4 and a degradation rate of 10−5 s−1. The range becomes more sensitive to changes in Kd and less sensitive to changes in Tx rate. (E and F) Combined effect of Tx rate (x axis) and Kd (y axis) with a Hill coefficient of 1 and a degradation rate of 5 × 10−4 s−1 (G and H) Combined effect of Tx rate (x axis) and degradation rate (y axis) with a Hill coefficient of 1 and Kd of 6 nM. Range and onset of expression are only sensitive to changes in degradation rates when RNA half-life is very short (half life <30 min). (I and J) Combined effect of Kd (x axis) and degradation rate (y axis) for a Hill coefficient of 1 and Tx rate of 0.1 count/s. Range and onset of expression are only sensitive to changes in degradation rates when RNA half-life is very short (half life <30 min).
Constraining the kinetic model through in vivo measurements.
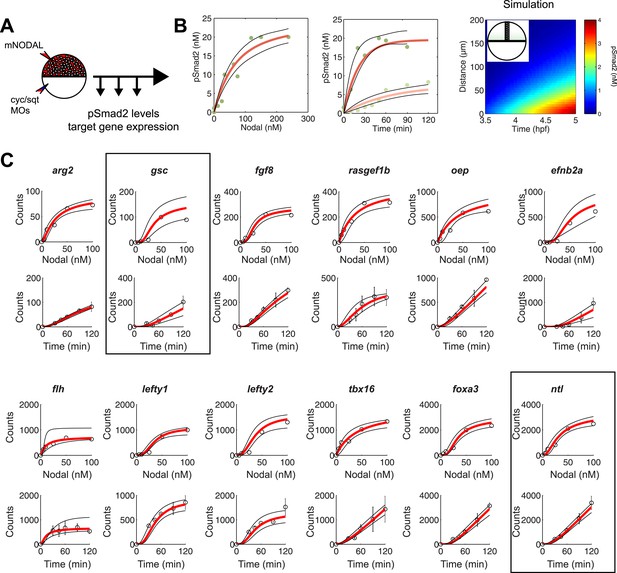
(A) Experimental design: Wild-type embryos were injected at the one-cell stage with squint and cyclops MOs to knock down endogenous Nodal signaling. Morphant embryos were further injected either at 3.5 or 4.5 hpf with recombinant mouse Nodal protein at different concentrations in the extracelluar space. They were then incubated for different periods of time and processed for Western blot to determine pSmad2 levels or for NanoString to assess mRNA levels. (B) Dose-response (left panel) and time course (middle panel) of Smad2 activation at high (100 nM, dark green) and low (10 nM, light green) Nodal concentrations. Dots represent experimental data points and orange lines show model simulations with k2 = 3.13 × 10−6 nM−1s−1 and k3 = 1.8 × 10−4 s−1. Black lines represent the 95% confidence intervals of data predictions. (Right panel) Simulated spatial distribution of Smad2 activation in a one-dimensional column of cells from 3.5 to 5 hpf in response to Nodal production from a source that extends from L = 0 to 25 µm. (C) Dose-response (top) and time course (bottom) data of 12 direct Nodal targets (black dots). Given a specific set of parameters for each gene, the model (red line) recapitulates the dynamics of gene expression. Black lines represent fits encompassing the 95% prediction confidence intervals. gsc and ntl dynamics are highlighted within black boxes.
-
Figure 5—source data 1
NanoString Probeset.
Listed are all the genes and target sequences included in the probeset.
- https://doi.org/10.7554/eLife.05042.017
-
Figure 5—source data 2
Smad2 associated peaks after Nodal injection.
- https://doi.org/10.7554/eLife.05042.018
-
Figure 5—source data 3
Smad2 associated peaks after Nodal signaling inhibition.
- https://doi.org/10.7554/eLife.05042.019
-
Figure 5—source data 4
FoxH1 associated peaks after Nodal injection.
- https://doi.org/10.7554/eLife.05042.020
-
Figure 5—source data 5
FoxH1 associated peaks after Nodal signaling inhibition.
- https://doi.org/10.7554/eLife.05042.021
-
Figure 5—source data 6
NanoString counts of Nodal target genes.
Nanostring counts of the 61 direct and indirect Nodal target genes. See Source code 2 for details.
- https://doi.org/10.7554/eLife.05042.022
-
Figure 5—source data 7
Nodal target genes identified in the NanoString codeset and their associated characteristics.
(A) Simulated transcription rate associated with the best fit. (B) Range of transcription rates encompassing the 95% confidence intervals. (C) Simulated degradation rate associated with the best fit. (D) Range of degradation rates encompassing the 95% confidence intervals. (E) Simulated Kd associated with the best fit. (F) Range of Kd encompassing the 95% confidence intervals. (G) Simulated Hill coefficient associated with the best fit. (H) Range of Hill coefficient values encompassing the 95% confidence intervals. (I) Coefficient of determination value of the best fit. (J) Maximum mRNA levels obtained in NanoString experiments (counts). (K) Maximum range of simulated spatial expression. The predicted spatial gene expression ranges only build on the simulated response to Nodal signaling kinetics. Regulation by other pathways is not taken into account and might change the in vivo expression patterns of these genes. (L) Degree of insensitivity to cycloheximide treatment: Strong (+), mild (±) or no (−) induction by Nodal after cycloheximide. (M and N) Amplitude of FoxH1 and Smad2 associated peaks: ++high, +medium, ±low, -background levels. See text and methods for details.
- https://doi.org/10.7554/eLife.05042.023
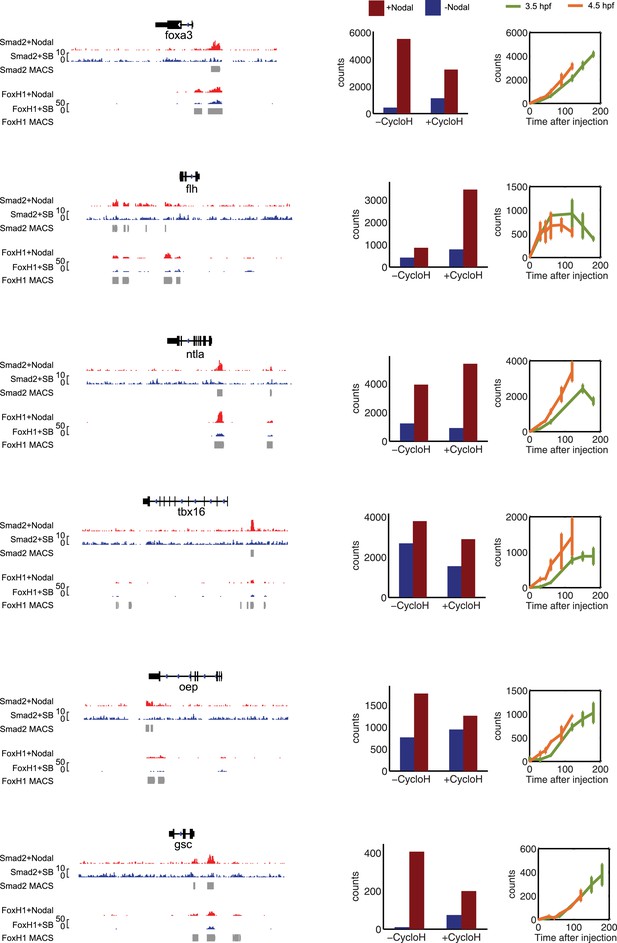
Characterization of direct Nodal target genes.
Left panel: binding peaks of Smad2 and its associated transcription factor FoxH1 at dome stage after injection of zebrafish Nodal Squint mRNA (red) or after treatment with the Nodal signaling inhibitor SB505124 (blue). Peaks called by the MACS algorithm are indicated (gray blocks). Middle panel: NanoString count levels after injection of recombinant mouse Nodal protein (red) or after SB505124 treatment (blue) in the absence (−cycloH) or in the presence (+cycloH) of the translation inhibitor cycloheximide. Right panel: time course induction of target genes after Nodal injection at 3.5 hpf (green) or 4.5 hpf (orange). In each case, (1) a specific Smad2 peak associated with a FoxH1 peak appears in the vicinity of the TSS upon Squint injection, (2) the Nodal-induced expression is not abolished by cycloheximide treatment, and (3) induction kinetics have similar trajectories independently of the stage of Nodal exposure, suggesting that these genes are direct targets of Nodal signaling.
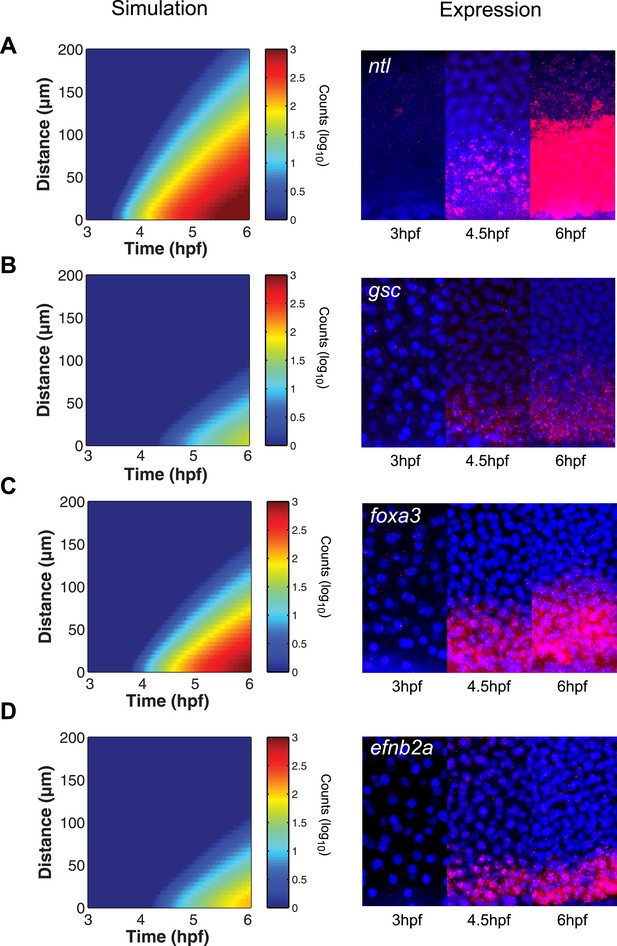
The kinetic model predicts gene expression patterns.
Comparison of kinetic model simulations and RNA fluorescent in situ hybridization for ntl (A), gsc (B), foxa3 (C), efnb2a (D). Left panels: simulations of spatiotemporal expression patterns over 3 hr along a 200 µm-high column of cells using gene-specific parameters identified in the parameter screen. Right panels: RNA fluorescent in situ hybridization at 3, 4.5 and 6 hpf. The size of the embryonic field is 100 µm wide and 200 µm high. Animal pole to the top.
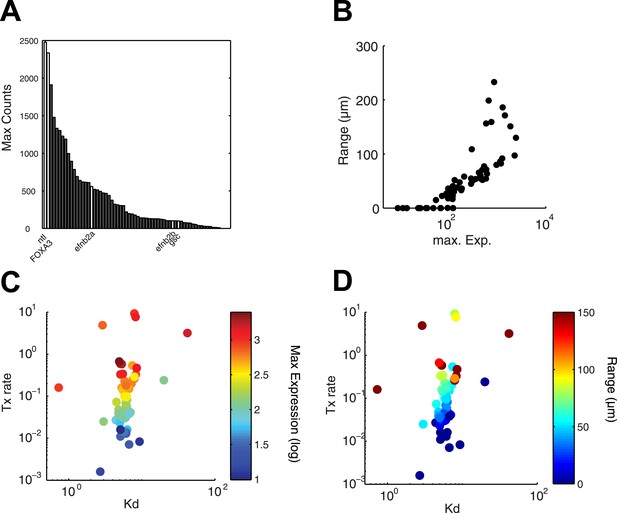
Range of expression correlates with maximal transcription rate.
(A) Bar graph showing the number of counts detected 90 min after injection of 100 nM of recombinant Nodal protein for the 61 Nodal-responsive genes (direct and indirect) identified in the NanoString codeset. Some of the genes used in this study are highlighted. (B) Scatter plot comparing maximal expression and simulated range. Highly expressed genes tend to have a longer range of expression. (C and D) Scatter plots comparing fitted Kd and maximal transcription rate (Tx rate) in relation to maximal expression (C) and in relation to simulated spatial range of expression (D). Most Kd values remain in a narrow range while transcription rates spread over several orders of magnitude.
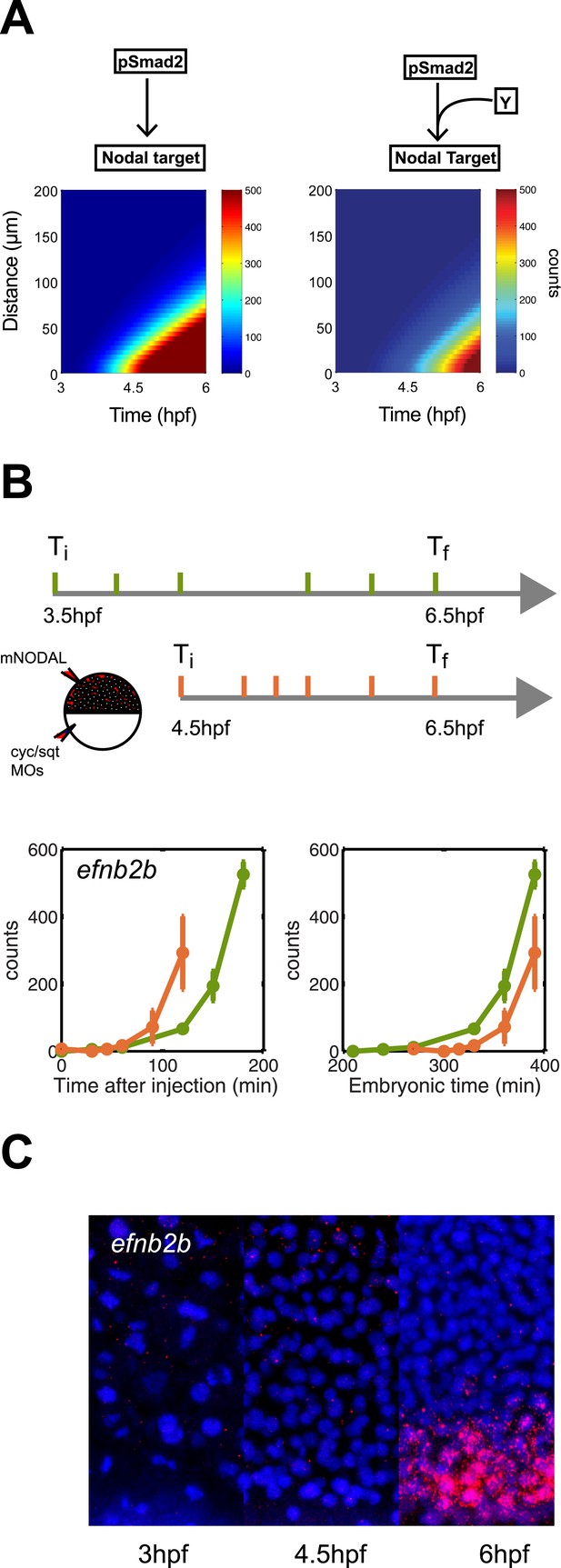
Delayed onset of transcription restricts expression range.
(A) Simulation of efnb2b expression using the kinetic model without (left) or with (right) a co-transcriptional activator Y. The dependence on Y delays the onset of efnb2b expression and reduces its range. (B) Top: Experimental design. Bottom: Time-course induction of efnb2b after injecting recombinant Nodal protein at 3.5 hpf (green) and 4.5 hpf (orange). The induction kinetics of this gene are very slow, but the later Nodal is injected, the faster its induction. Note that counts for the expression of late target genes are higher after early injection compared to later injections. This effect might be due to the fact that after early injections phospho-Smad2 levels are high for a longer period before a gene becomes competent to respond as compared to late injections, when there is a shorter time window of high phospho-Smad2 levels. There might be a priming mechanism in which longer exposure to activated Smad2 increases gene expression when competence is reached. (C) RNA fluorescent in situ hybridization for efnb2b at 3, 4.5 and 6 hpf. Expression of efnb2b is only detected at 6 hpf, although Nodal signaling and the expression of most other Nodal targets commences much earlier.
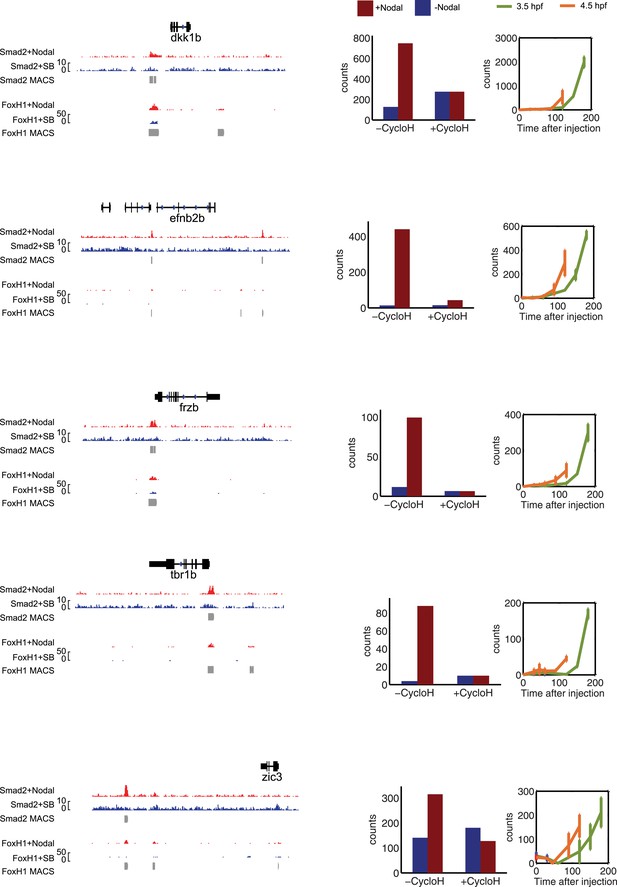
Characterization of co-regulated Nodal target genes.
Left panel: binding peaks of Smad2 and the associated transcription factor FoxH1 at dome stage after injection of Squint mRNA (red) or after treatment with the Nodal signaling inhibitor SB505124 (blue). Peaks called by the MACS algorithm are indicated (gray blocks). Middle panel: NanoString count levels after mNodal injection (red) or after SB treatment (blue) in the absence (−cycloH) or the presence (+cycloH) of the translation inhibitor cycloheximide. Right panel: time course induction of target genes after Nodal injection at 3.5 hpf (green) or 4.5 hpf (orange). Although these genes have Smad2/FoxH1 binding sites, their induction is abolished in the presence of cycloheximide, suggesting that an additional transcriptional co-regulator controls their transcription. Moreover, these genes are delayed in their onset of expression. The length of the delay depends on the stage at which Nodal is applied.
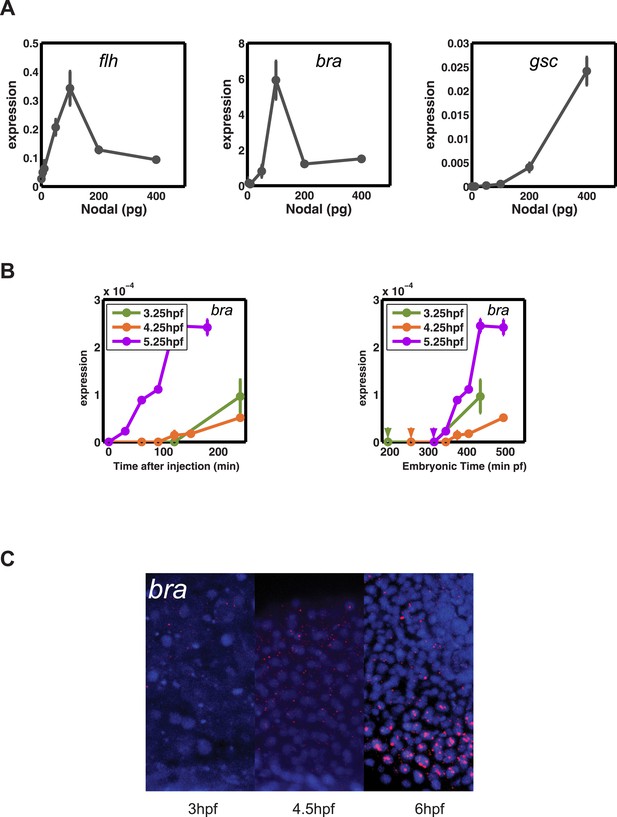
Transcriptional competence regulates the onset and range of bra expression.
(A) Concentration-dependent induction of flh (left), bra (middle) and gsc (right). bra and flh can be induced at low Nodal concentrations. (B) Induction dynamics of bra after injecting Nodal at 3.25, 4.25 and 5.25 hpf as a function of time after injection (left) or as a function of absolute embryonic time (right). Arrowheads in the right panel indicate the time of Nodal injection. bra can only be induced when the embryo has reached a specific embryonic stage. bra expression is detected by RT-qPCR. (C) Fluorescent RNA in situ hybridization with bra probe at 3, 4.5, 6 hpf. bra is only detected at 6 hpf in 5 cell tiers.
Tables
β-actin::GFP-Smad2 transgene rescues smad2/smad2 adult lethality
| smad2/+ X smad2/+; Tg(GFP-Smad2)/+ | ||
|---|---|---|
| Genotype | +/+ | Tg(gfp-smad2)/+ |
| +/+ | 11 (37%) | 4 (15%) |
| smad2/+ | 19 (63%) | 17 (60%) |
| smad2/smad2 | 0 (0%) | 7 (25%) |
-
smad2/+ fish were crossed to smad2/+; Tg(GFP-Smad2)/+ fish and their progeny was raised to adulthood and genotyped for smad2 and for Tg(GFP-Smad2). The only recovered adult progeny homozygous for smad2 contains a copy of the GFP-Smad2 transgene.
Additional files
-
Supplementary file 1
Scripts used for tracking cells and measuring the nucleo-cytoplasmic ratio.
- https://doi.org/10.7554/eLife.05042.030
-
Source code 1
(runPdeSysTestSourceNoLefty.m): Simulation of the spatial distribution of Nodal, acitve Smad2 and Nodal target gene.
- https://doi.org/10.7554/eLife.05042.031
-
Source code 2
(runOdeSysplotNanoM1.m): Simulation of the time- and Nodal concentration- dependent induction of target genes.
- https://doi.org/10.7554/eLife.05042.032





