In vitro generation of human pluripotent stem cell derived lung organoids
Figures
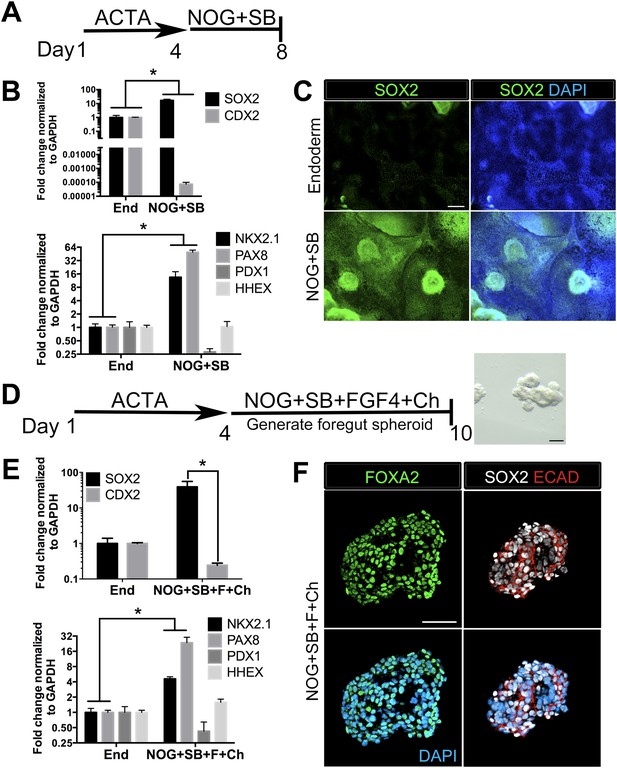
Generation of three-dimensional ventral anterior foregut spheroids from endoderm monolayers.
(A) hESCs were differentiated into foregut endoderm by treating cells with 4 days of Activin A (ACTA) followed by 4 days of NOG+SB. (B) Foregut endoderm (NOG+SB) had high expression of the foregut marker SOX2 while the hindgut marker CDX2 was significantly reduced compared to untreated endoderm controls (End). NOG+SB monolayers had high expression of ventral anterior foregut genes NKX2.1 and PAX8 while the posterior foregut marker PDX1 was reduced. The foregut marker HHEX is expressed in the developing liver, biliary system, and thyroid and remained unchanged. (C) The majority of cells in NOG+SB treated cultures were SOX2 positive (green) compared to the control, in which only scattered clusters of cells were SOX2 positive. The scale bar represents 200 µm. (D) hESCs were differentiated into foregut spheroids by treating cells with 4 days of ACTA and then additional 4–6 days of NOG+SB+FGF4+Ch. Representative images of a spheroid in a matrigel droplet are shown as a whole mount image. Scale bar represents 100 µm. (E) Foregut spheroids (NOG+SB+FGF4+Ch) had high expression of the foregut marker SOX2 while the hindgut marker CDX2 was significantly reduced compared to untreated endoderm control (End) (top panel). Spheroids had high expression of anterior foregut genes NKX2.1 and PAX8 while the posterior foregut marker PDX1 was reduced and HHEX was unchanged (bottom panel). *p < 0.05, error bars represent SEM. (F) The majority of cells in foregut spheroids are FOXA2+ (green, left panel) and SOX2+ (white, right panel) and ECAD+ (red, right panel). Scale bar represent 50 µm.
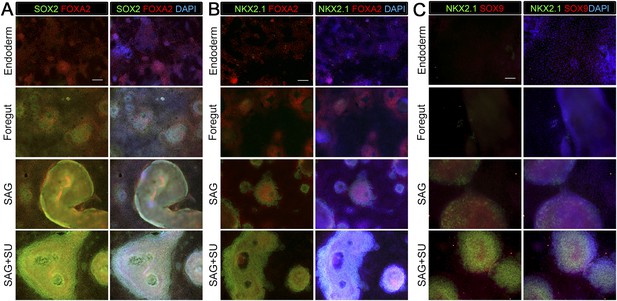
Monolayer cultures express lung specific markers.
Immunohistochemistry for markers expressed in endoderm, ventral foregut or lung epithelium were assessed (SOX2, FOXA2, NKX2.1, SOX9) in endoderm controls, foregut controls or foregut cultures treated with SAG or SAG+SU. (A) All conditions express endoderm marker FOXA2 (red), but the foregut (NOG+SB) control, SAG and SAG+SU treated cultures have co-expression of FOXA2 (red) and SOX2 (green) in the majority of cells. (B) All conditions expressed endoderm marker FOXA2 (red), but only foregut endoderm treated with SAG and SAG+SU have robust NKX2.1+ cells (green) that also express FOXA2 (red). (A–B) Scale bars represent 200 µm and apply to all images. (C) Only foregut endoderm treated with SAG and SAG+SU have robust NKX2.1+ cells (green) with the majority of cell co-expressing with lung epithelial marker SOX9 (red). Scale bar represents 100 µm.
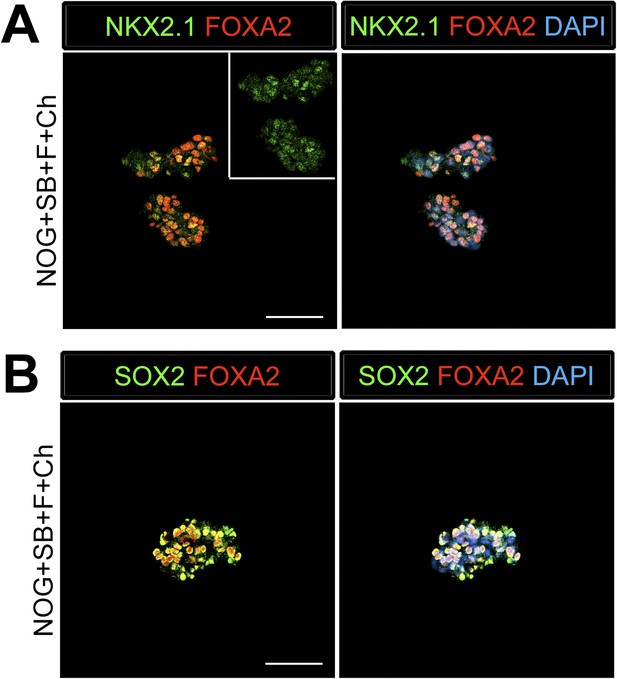
Foregut spheroids co-express endoderm and lung specific markers.
(A) NOG/SB/FGF4/Ch spheroids have weak NKX2.1 (green) expression which co-expresses with endoderm marker FOXA2 (red). (B) The majority of cells in the spheroid express SOX2 (green) and co-stain with FOXA2 (red). Scale bars represent 50 µM.
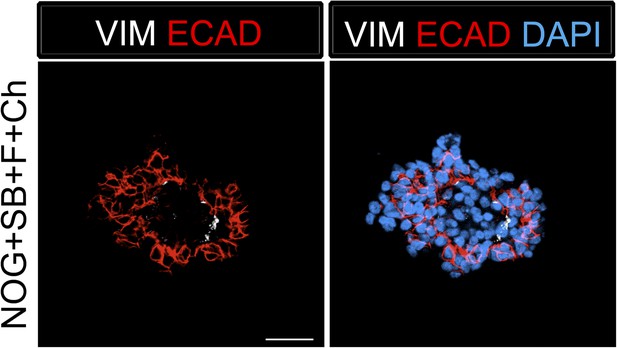
Foregut spheroids consist of both epithelial and mesenchymal cells.
NOG/SB/FGF4/Ch spheroids have a minor population of Vimentin (VIM, white) positive mesenchymal cells, while the majority of cells are epithelial and express ECAD (red). Scale bar represents 50 µM.
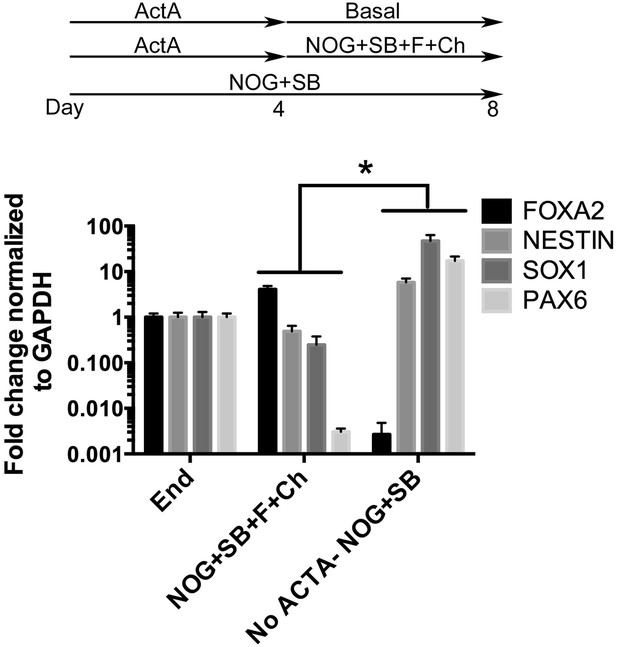
NOG+SB+FGF4+Ch spheroids do not express neural markers.
hESCs were differentiated into endoderm by treating with 4 days of ActivinA (ACTA) and spheroids were generated with an additional 4 days of NOG+SB+FGF4+Ch. Neural cultures were not treated with ACTA, but were treated with NOG+SB for 8 days. Compared to foregut spheroids (NOG+SB+FGF4+Ch), NOG+SB neural cultures had a significant increase in neural markers NESTIN, SOX1, and PAX6 and significant decrease in FOXA2, which is highly expressed in endoderm. *p < 0.05, error bars represent SEM.
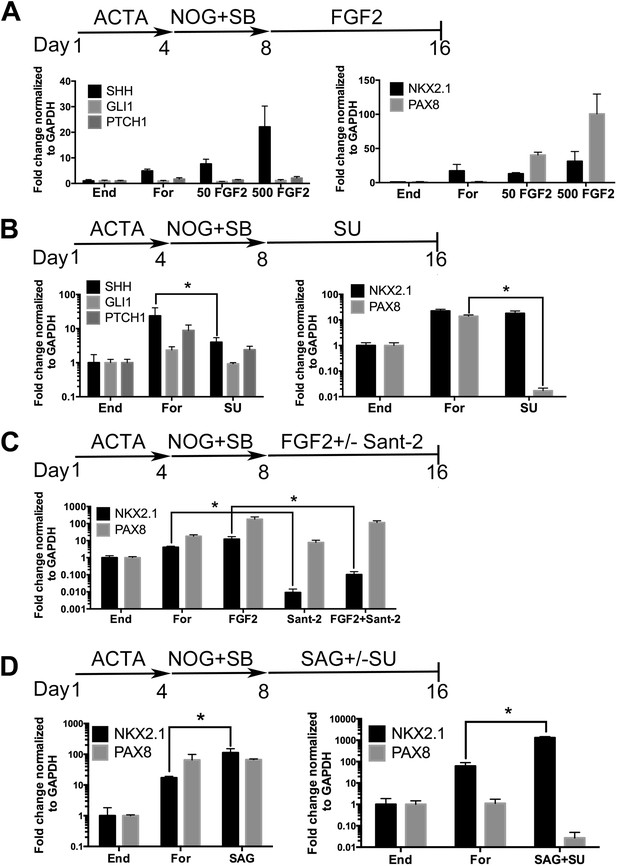
Induction of NKX2.1 in anterior foregut endoderm by modulating FGF and HH signaling.
(A) hESCs were differentiated into endoderm (End) or anterior foregut with NOG+SB (For). Anterior foregut was treated with low (50 ng/ml) and high (500 ng/ml) concentrations of FGF2. FGF2 caused a dose-dependent increase in SHH and PAX8 expression with a modest increase in NKX2.1 expression compared to untreated endoderm controls. Note that NKX2.1 expression is increased by NOG+SB exposure alone (no FGF2). (B) Addition of the FGF inhibitor SU5402 (SU) to NOG+SB foregut cultures (For) caused a significant reduction of SHH and PAX8 expression, but NKX2.1, GLI1, and PTCH1 were not significantly different compared to the foregut controls, in which no growth factors were added after SB+NOG. (C) Addition of the HH inhibitor Sant-2 caused a significant reduction in NKX2.1 compared to foregut control. Similarly when FGF2 (500 ng/ml) and Sant-2 were added simultaneously, the modest NKX2.1 induction caused by FGF2 was significantly reduced whereas PAX8 expression remained unchanged. (D) Foregut endoderm treated with SAG or SAG+SU for 8 days had a 6.5-fold and 21-fold increase of NKX2.1 expression, respectively, compared to untreated foregut controls. PAX8 expression was unchanged in the SAG treated cultures whereas SAG+SU treated cultures demonstrated a 41-fold decrease in PAX8 expression. End = endoderm; For = foregut in all panels. *p < 0.05, error bars represent SEM.
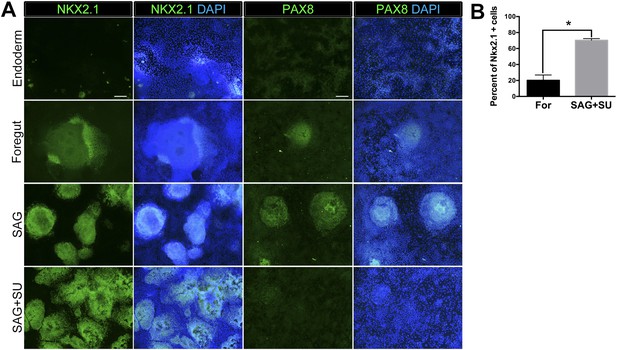
Robust induction of NKX2.1 in foregut endoderm with HH stimulation and FGF inhibition.
(A) Immunohistochemistry of NKX2.1 and PAX8 in endoderm controls, foregut controls or foregut cultures treated with SAG or SAG+SU. Treatment of foregut cultures with SAG or SAG+SU resulted in more NKX2.1+ cells compared to endoderm and foregut controls. Scale bars represent 200 µm and apply to all images. (B) Quantification showed that 20% ± 4% of cells in foregut controls were NKX2.1+, whereas 72% ± 3% cells were positive in SAG+SU treated cultures (*p < 0.05). All error bars represent SEM.
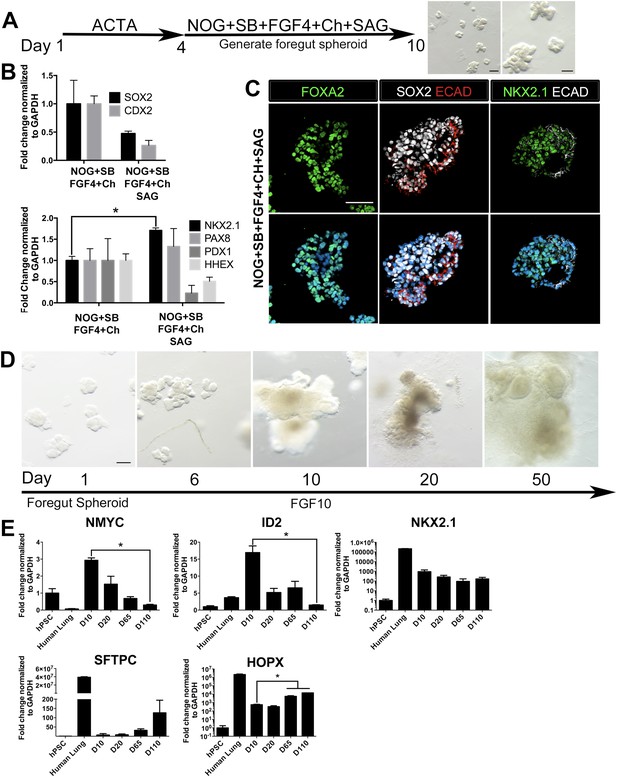
HH-induced ventral foregut spheroids give rise to lung organoids.
(A) hESCs were differentiated into foregut spheroids by treating cells with 4 days of ACTA and then an additional 4–6 days of NOG+SB+FGF4+Ch with the addition of the HH agonist SAG. Representative whole mount images of spheroids in a matrigel droplet are shown at low (left, scale bar 200 µm) and high magnification (right, scale bar 100 µm). (B) The addition of SAG to the NOG+SB+FGF4+Ch spheres caused a reduction in SOX2 and CDX2 transcripts (top panel) and a significant increase of NKX2.1 transcript (bottom panel) compared to NOG+SB+FGF4+Ch spheres (without SAG). Other foregut lineages (PAX8, PDX1, HHEX) were not significantly different when SAG was added. (C) The majority of the cells in NOG+SB+FGF4+Ch+SAG spheres expressed FOXA2, SOX2 and NKX2.1 protein. Scale bars represent 50 µm. (D) Timeline showing NOG+SB+FGF4+Ch+SAG induced foregut spheroids grown and maintained in FGF10. Note that Day 1 is the day spheroids were plated in Matrigel. The scale bar represents 100 µm. (E) Organoids express lung markers in a manner consistent with mouse lung development. All expression is shown relative to undifferentiated pluripotent stem cells (hPSC), and adult human lung is shown as a reference. Lung progenitor markers NMYC and ID2 were very low in adult lung, and were expressed at high levels in early organoid cultures, but were reduced over time (D = Days in culture), whereas NKX2.1 expression remained relatively constant. In contrast, SFTPC is known to be expressed at low levels in distal lung progenitors, but increases and is highly expressed in AECII cells. Consistently, SFTPC is highly expressed in adult human lungs and increases over time in organoid cultures and the AECI marker HOPX is also highly expressed in adult human lung and increases over time in organoids. *p < 0.05. All error bars represent SEM.
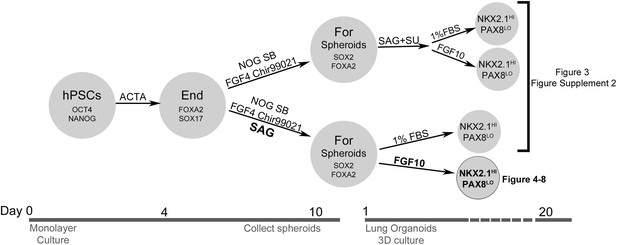
Overview of conditions tested to generate human lung organoids.
hPSCs are OCT4 and NANOG positive. After 4 days of 100 ng/ml Activin A, definitive endoderm (FOXA2 and SOX17 positive) was generated and then treated with two different conditions. In the top branch, NOG+SB+FGF4+Ch spheroids were generated, and different conditions were tested to promote lung organoid differentiation. In the bottom branch, NOG+SB+FGF4+Ch+SAG spheroids were generated, and different conditions were tested to promote lung organoid differentiation. Ultimately, we determined that spheroids generated with NOG+SB+FGF4+Ch+SAG and that were subsequently embedded in Matrigel and expanded in FGF10 gave rise to ‘Human Lung Organoids’ (HLOs).
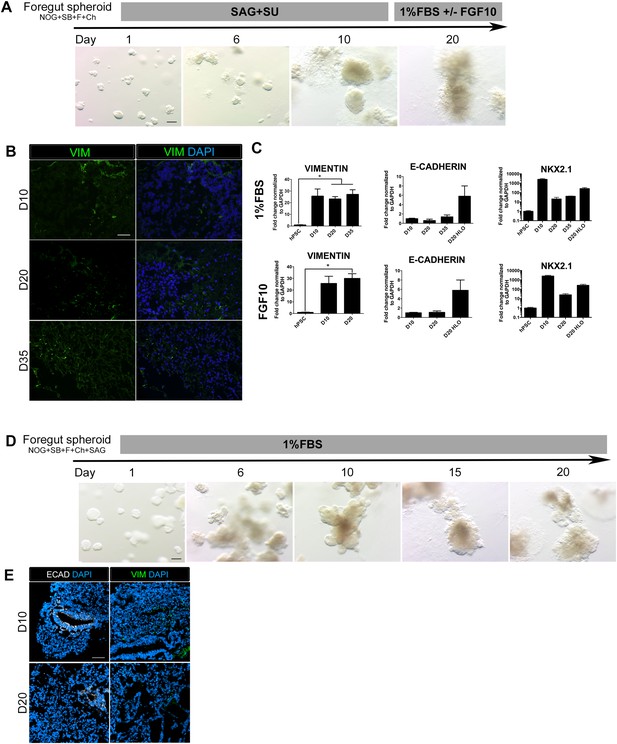
FGF-low culture conditions cause a loss of organoid epithelium over time.
(A) NOG+SB+F+Ch foregut spheroids were generated and then cultured in SAG+SU for 10 days followed by 1% FBS ± FGF10. Timeline images show organoids cultured in 1% FBS. By day 20, 3D structures appeared ‘fuzzy’, which indicates an outgrowth of mesenchymal tissue. Scale bar represents 200 µm. (B) NOG+SB+F+Ch foregut spheroids treated with SAG+SU and maintained in 1% FBS showed an increase in Vimentin (VIM, green) immunofluorescence over time. Scale bar represents 50 µM. (C) NOG+SB+F+Ch foregut spheroids treated with SAG+SU and maintained in 1% FBS (upper panel) or 1% FBS+FGF10 (lower panel) had a significant increase of VIM expression starting at day 20 (D20) compared to hPSCs and showed very weak E-CADHERIN (CDH1) expression compared to D20 HLOS (optimized conditions, as described in Figure 3). Lastly, both conditions appeared to lose NKX2.1 expression over time. (D) NOG+SB+F+Ch+SAG spheroids maintained in 1% FBS (basal media) also appear to lose epithelial structures over time. Scale bar represents 200 µm. (E) By day 20 (D20) the tissue had very few epithelial structures expressing ECAD (white, left panel) and there was robust VIM expression (green, right panel) at both time points. Scale bar represents 50 µm. HLO *p < 0.05. All error bars represent SEM.
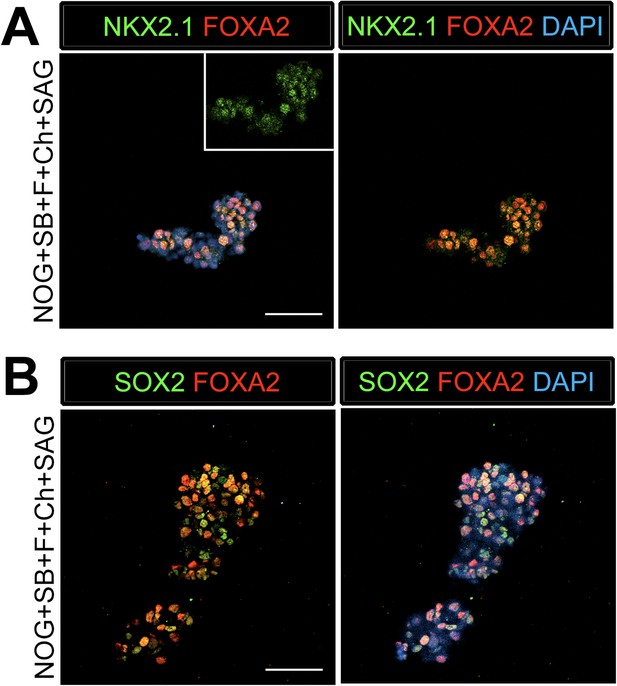
Foregut spheroids express lung and foregut specific markers.
(A) NOG/SB/FGF4/Ch/SAG spheroids coexpress NKX2.1 (green) and the endoderm marker FOXA2 (red). (B) The majority of the cells in the spheroid co-expresses SOX2 (green) and FOXA2 (red). Scale bars represent 50 µM.
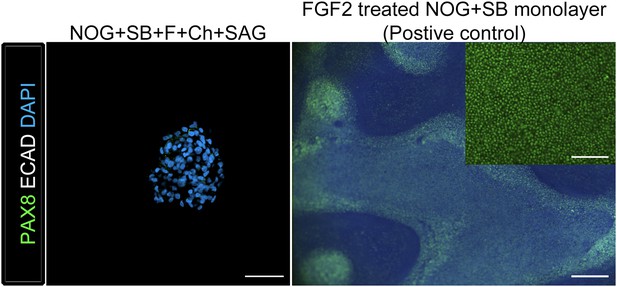
Ventral foregut spheroids do not express appreciable levels of PAX8 protein.
Although NOG+SB+FGF4+Ch+SAG ventral foregut spheroids expressed PAX8 mRNA (Figure 3B), we did not detect PAX8 protein in spheroids using immunofluorescence, whereas PAX8 protein in FGF2 8 day treated foregut monolayers (ACTA followed by NOG/SB) was readily detectable. Left panel: scale bar represents 50 µm. Right panel: scale bar represents 200 µm, inset scale bar represents 100 µm.
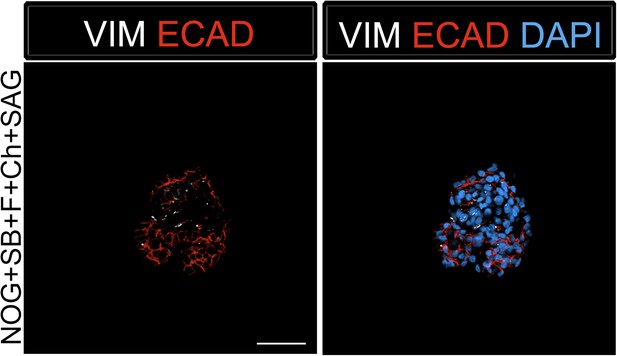
Foregut spheroids consist of both epithelial and mesenchymal cells.
NOG/SB/FGF4/Ch/SAG spheroids have a minor population of Vimentin (VIM, white) positive mesenchymal cells, while the majority of cells are epithelial and express ECAD (red). Scale bar represents 50 µM.
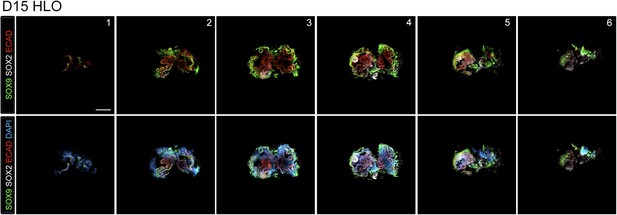
Lung organoids contain both proximal and distal domains.
NOG/SB/FGF4/Ch/SAG spheroids cultured for 15 days with FGF10 express the distal lung epithelium marker SOX9 (green) and proximal marker SOX2 (white) as separate domains in the epithelium labeled by ECAD (red). Z-stack images are shown every 40 µm sections through the HLO. Scale bar represents 200 µm.
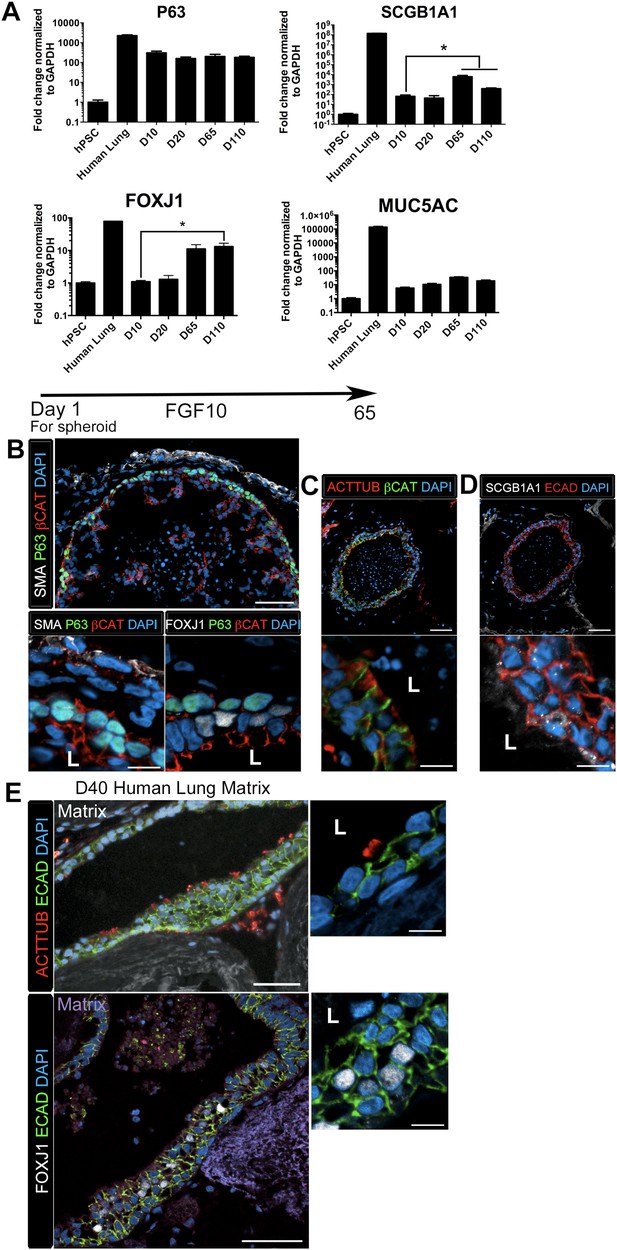
Lung organoids form proximal airway-like structures.
(A) Genes expressed in the proximal airway were examined in organoids across time. The proximal airway cell marker SOX2 decreased over time in HLOs cultures compared to D10 HLOs. Compared to undifferentiated hPSCs, organoids expressed high levels of the basal cell marker P63 at all time points, while expression of the club cell marker SCGB1A1 and ciliated cell marker FOXJ1 increased significantly in prolonged cultures (compared to D10 HLOs). There was an increasing but non-significant trend in goblet cell MUC5AC expression over time in culture. (B) D65 HLOs had structures resembling the proximal airway, in which the epithelium (β-catenin, red) possesses P63+ basal cells (green), and is surrounded by SMA+ (white, upper and lower left panel) mesenchymal tissue. Adjacent to the P63 positive basal cell layer (green, lower, right panel) were FOXJ1 positive cells (white). Scale bars represent 50 µM (top) and 10 µM (bottom). (C) Proximal airway-like epithelium (β-catenin, green) co-stained for ACTTUB on the apical side of the cell (red). Scale bars represent 50 µM (top) and 10 µM (bottom). (D). Proximal airway-like epithelium (E-cadherin, red) also co-stained with Club cell marker CC10 (white, right panel). Scale bars represent 50 µM (top) and 10 µM (bottom). (E) Acellular human lung matrix was seeded with spheroids and cultured for 40 days (D40). Matrices had abundant proximal airway-like structures that had multi-ciliated cells on the apical surfaced labeled by ACTTUB (red, top panel) in low (scale bar 50 µM) and high magnification (scale bar 10 µM). Serial sections showed that cells were also FOXJ1 positive (white, lower panel) with the epithelium outlined in ECAD (green) in low (scale bar 50 µM) and high magnification (scale bar 10 µM). (B–D) ‘L’ in high magnification images indicates the lumen. *p < 0.05. All error bars represent SEM.
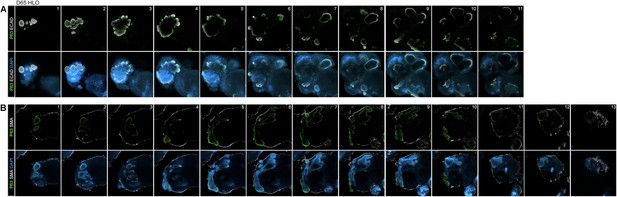
Lung organoids have P63+ epithelium throughout the organoid.
(A) Confocal Z-slices taken at every 40 µm show P63+ (green) and ECAD+ (white) structures through the D65 HLO. (B) Z-slices taken at every 40 µm show SMA (white) surrounding the periphery the HLO with P63 (green) staining within the HLO. Scale bars represent 200 µm.
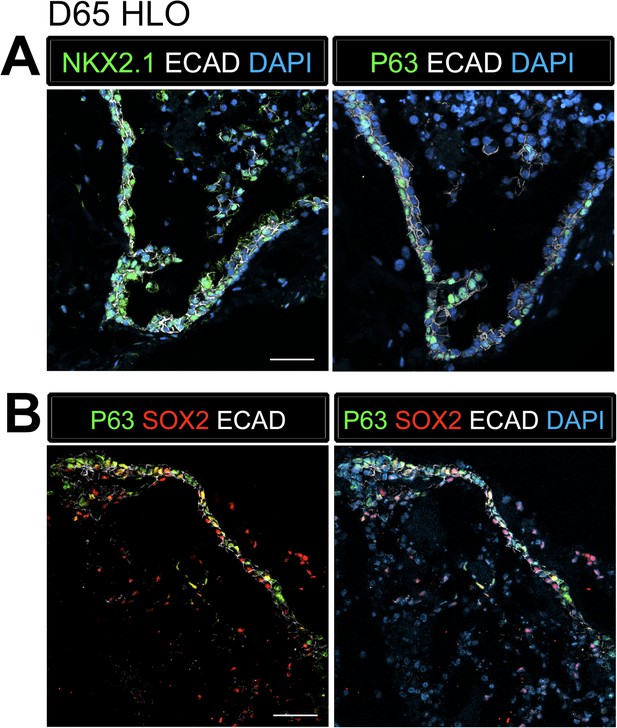
P63+ cells have an NKX2.1+ lung identity.
(A) Serial sections were stained with NKX2.1 and P63 respectively. The adjacent sections expressed ECAD (white) and NKX2.1 (green) in the first section and P63 (green) in the second section. (B) P63+ cells (green) co-expressed the proximal lung marker SOX2 (red) in the epithelium labeled by ECAD (white). Scale bars represent 50 µm.
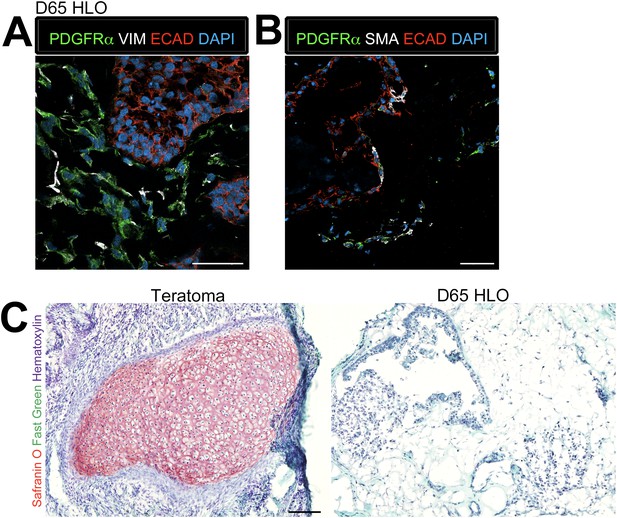
Lung organoids possess multiple types of mesenchymal cells.
(A) D65 HLOs have PDGFRα+ (green) VIM+ (white) double-positive myofibroblasts and PDGFRα−/VIM+ fibroblasts. Scale bar represents 50 µm. (B) D65 HLOs also possesses PDGFRα+ (green) SMA+ (white) double-positive myofibroblasts and PDGFRα−/SMA+ smooth muscle and myofibrblasts. Scale bar represents 50 µm. (C) D65 HLOs do not contain any cartilage whereas positive control iPSC derived teratoma had clear Safranin O staining specific to cartilage. Fast green marks the cytoplasm and hematoxylin the nuclei of both tissues. Scale bar represents 100 µm.
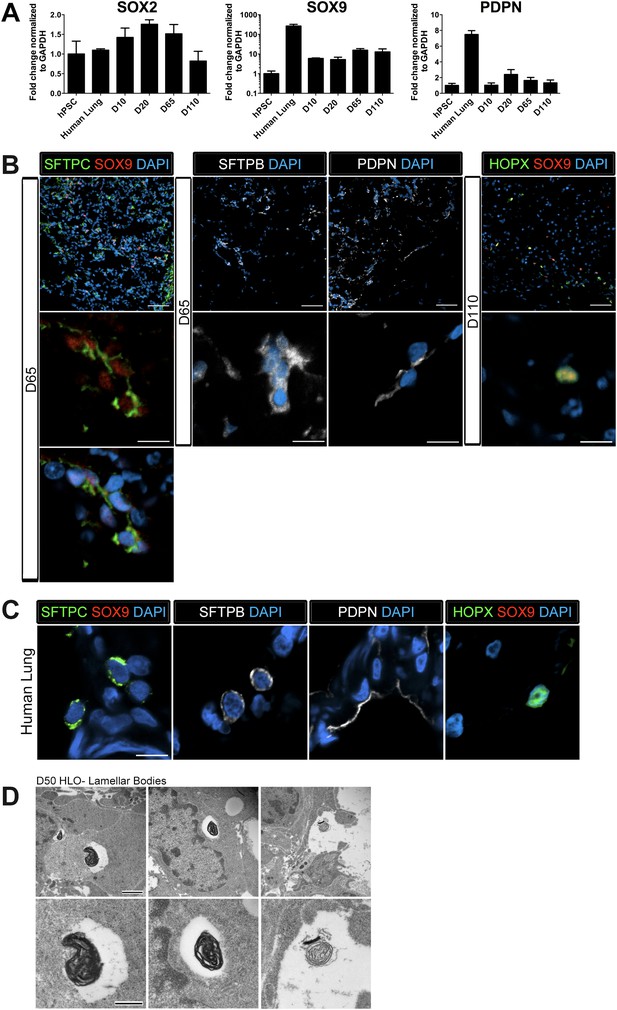
Lung organoids possess abundant distal bipotent progenitor cells.
(A) The expression of the distal progenitor marker SOX9 remained unchanged over time and expression of the AECI marker PDPN was low in HLO cultures. (B) The majority of SFTPC+ cells (green, left panel) co-expressed SOX9 (red). Similarly, many cells expressing the AECI early marker HOPX+ (green, right panel) co-expressed SOX9 (red). Few, scattered cells expressed the late AECII marker SFTPB (white, second panel) or the AECI marker, PDPN (third panel, white). Few PDPN+ cells also showed elongated, squamous morphology seen in the adult lung. (C) Human lung AECII cells labeled with SFTPC (green, left panel) did not co-express SOX9. SFTPB+ cells (white, second panel) in the adult human lung have similar morphology to SFTPB+ cells in HLOs. Human lung AECI cells expressed PDPN (white, third panel), and show characteristic AECI cell shape. Human AECI cells express HOPX (green, right panel), but did not co-express SOX9. (B–C) Scale bar in lower magnification images in B (upper panel) represent 50 µM and the scale bars in higher magnification images in B, C (lower panel) represent 10 µM. (D) D50 HLOs contain lamellar bodies which are organelles specific to AECII cells. Scale bars represent 500 nm.
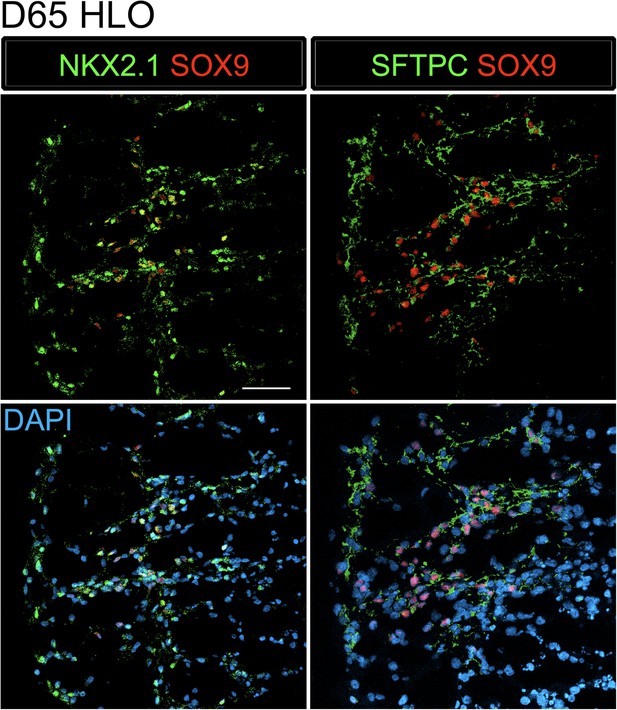
SFTPC+ cells express lung specific markers.
D65 HLOs express lung epithelial markers NKX2.1 (green) and SOX9 (red) and the adjacent section expresses SFTPC (green) and SOX9 (red). Scale bar represents 50 µm.
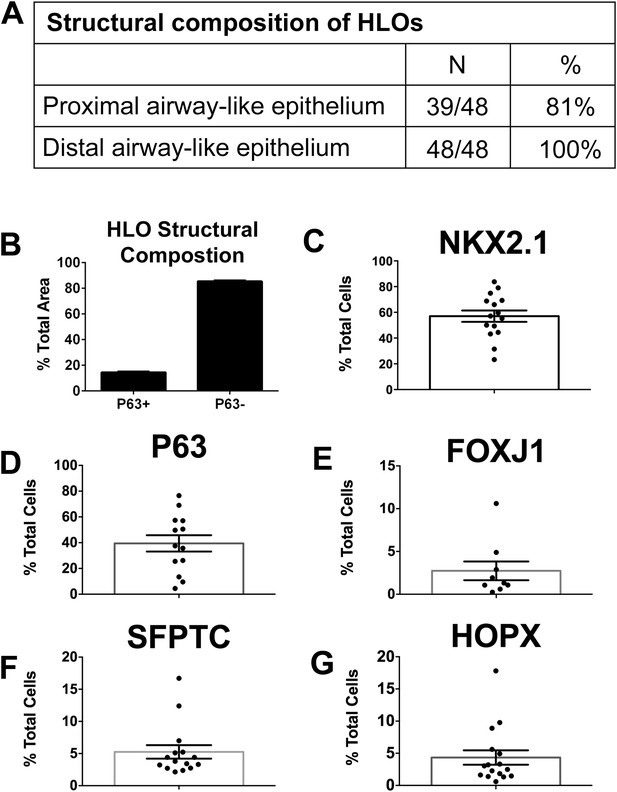
Quantitative assessment of the composition of lung organoids.
(A) HLOs were assessed for proximal airway-like structures (P63+) and distal airway-like structures (P63−/SFTPC+). 81% of HLOs have proximal airway-like epithelium while 100% have distal airway-like epithelium (n = 48 individual HLOs). (B) The average cross-sectional area within an HLO that is comprised of P63+ proximal airway-like and P63−/SFTPC+ distal airway-like epithelium was calculated. Proximal structures comprised 14.5% (±0.6%) of the entire area of the HLO (P63+), whereas 85.5% (±0.6%) of HLO was distal-like epithelium and mesenchyme (P63−). (C–G) The percent of specific cell markers present in an organoid was determined by dividing by the total number of DAPI+ nuclei within the same section (n = 15 individual HLOs). Each point represents the data from an individual HLO while the open bar represents the average percent of cells. (C) On average, 57% of all cells in the HLOs were NKX2.1+, (D) 39% of all cells were P63+, (E) 3% were FOXJ1+, (F) 5% were SFTPC+, (G) 4% of all cells were HOPX+. (B–G) Error bars represent SEM.
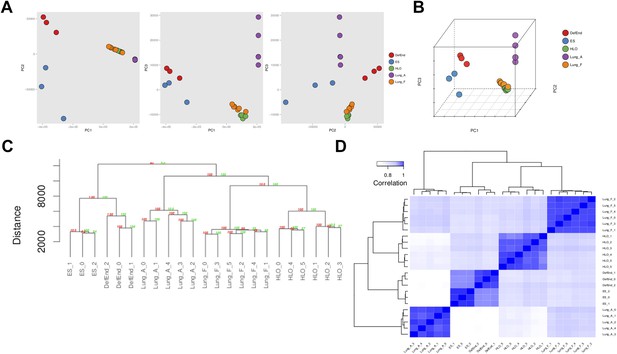
RNA sequencing analysis associates HLOs with fetal lung tissue.
6 HLOs (n = 3 D65 HLOs and n = 3 D110 HLOs) were compared to the undifferentiated H9 stem cells (SC) and definitive endoderm (Def End) and publicly available datasets of adult and fetal human lungs (see Supplementary file 1). (A–B) Plot of the first three principle components generated in the principle component (PC) analysis as pairwise 2-dimensional plots (A) or as an aggregate 3-dimensional projection (B), (C) hierarchical clustering, and (D) Spearman's correlation all demonstrate that HLOs are most closely related to the fetal lung.
Additional files
-
Supplementary file 1
Publicly available RNAseq datasets for human fetal lung representing a range of gestational stages and for adult human lung.
- https://doi.org/10.7554/eLife.05098.025
-
Supplementary file 2
Antibody information and dilutions.
- https://doi.org/10.7554/eLife.05098.026
-
Supplementary file 3
Primer sequences.
- https://doi.org/10.7554/eLife.05098.027





