YcgC represents a new protein deacetylase family in prokaryotes
Figures
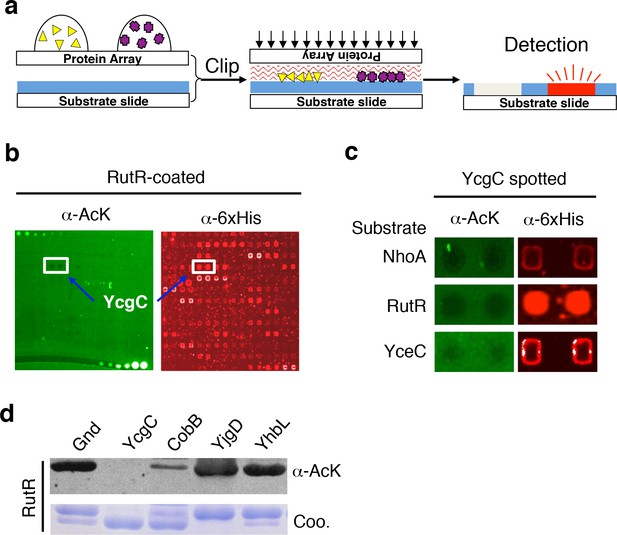
Screening the Escherichia coli proteome to discover new KDACs using the ‘clip-chip’ strategy.
(a) Schematic of the ‘clip-chip’ strategy. (b,c) Identification of YcgC as a potential protein deacetylase. E. coli proteome chips were clipped onto three substrate slides separately coated with acetylated RutR, NhoA, and YceC. After incubation in a protein deacetylase buffer, the reactions were terminated by adding wash buffers, followed by a signal detection step with a pan α-AcK antibody coupled with a Cy3-labeled secondary antibody as detection reagent to visualize the loss of signals (e.g., black holes in (b,c). To determine the identity of proteins that generated the holes, the substrate slide was subsequently probed with an α-6xHis antibody followed by a Cy5-conjugated secondary antibody. (d) Using acetylated RutR proteins purified from E. coli, of the four candidates tested, YcgC showed robust deacetylation activity in vitro. Equal amounts of RutR proteins were used in each reaction and loss of acetylation was detected with the pan α-AcK antibody.
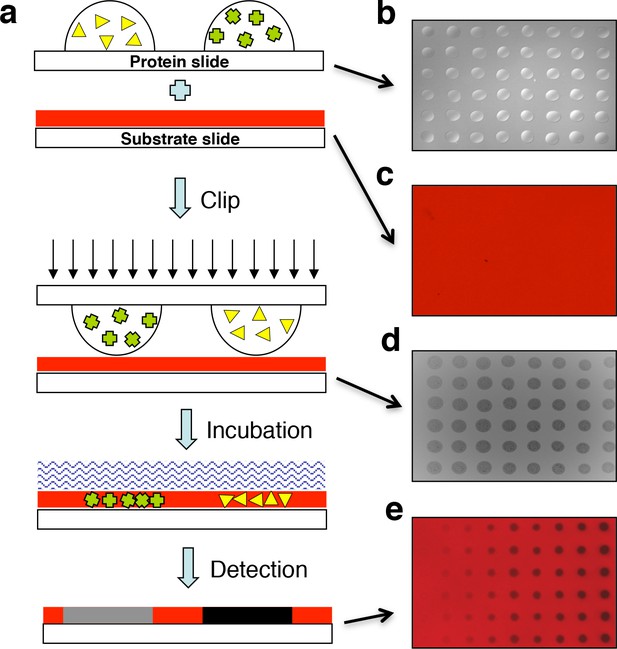
Design of the ‘clip-chip’ strategy.
(a) The ‘clip-chip’ strategy uses two slides, a protein slide that contains proteins of interest printed onto a slide with an appropriate surface, and a second substrate slide on which the enzymatic reactions are carried out. The protein slide containing arrayed protein droplets (b) is first imprinted onto the slide coated with substrate (red) (c) to transfer the proteins of interest from the protein slide onto the substrate slide. (d) Visible and homogenous watermarks indicate that the protein droplets from the protein slide are effectively and evenly transferred to the substrate slide. (e) An appropriate enzymatic reaction buffer is then loaded onto the surface of the imprinted substrate slide and reactions are carried out under appropriate conditions. After a series of stringent washes, the results are recorded with a microarray scanner.
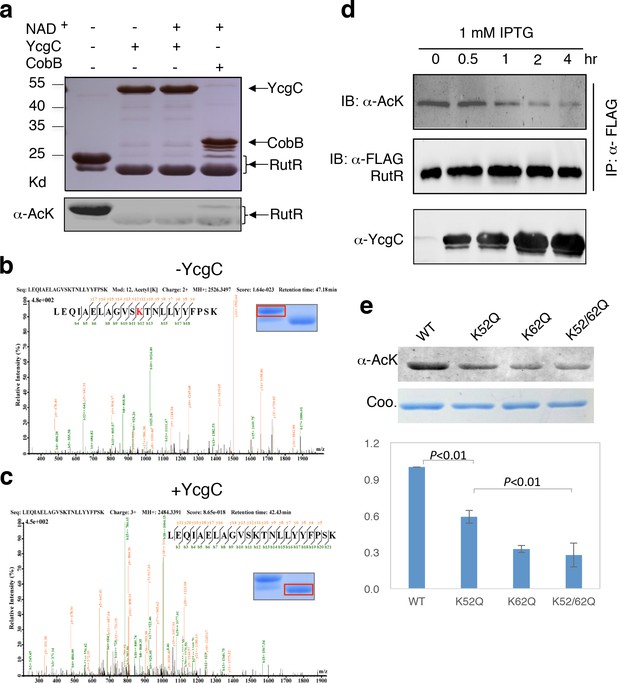
In vitro and in vivo characterization of YcgC’s KDAC activity.
(a) In vitro assays of the KDAC activity of YcgC on RutR demonstrated that its KDAC activity does not require either NAD+ or Zn2+ as cofactors. Incubation with YcgC almost completely abolished the slower migrating acetylated RutR bands (upper panel) as evidenced by immonublotting (lower panel). (b,c) LC-MS/MS analysis to determine the residues of RutR deacetylated by YcgC. RutR was treated with YcgC first and the untreated RutR used as the control. Both these two samples were resolved on a SDS-PAGE gel side by side. The upper band represents the Kac-containing starting materials and the lower band represents the K-containing product, which were then recovered from the gel and subjected for MS/MS analysis (inserts). Lys52 was identified as an acetylated site in RutR protein (b). After incubating with YcgC, acetylation on K52 was no longer detectable (c). (d) RutR is deacetylated by YcgC in Escherichia coli. A 3xFLAG tag was chromosomally inserted at the 3′-end of rutR coding sequence. Acetylation of 3xFLAG-tagged RutR was monitored upon induction of YcgC. While the protein level of RutR was unchanged (middle panel), its acetylation level was dramatically reduced as a function of YcgG induction (upper panel). YcgC’s expression was monitored using a custom-made antibody (lower panel). (e) Mutagenesis of RutR confirmed that K52 and K62 are acetylated in vivo. Two single mutants K52Q and K62Q and one double mutant K52/62Q were constructed. These mutants along with WT RutR were produced and purified in E. coli. Equal amounts of purified proteins were Western blotted with the α-AcK antibody, quantitation of acetylation level of these samples were performed. KDAC: Lysine deacetylase; LC-MS/MS: Liquid chromatography–mass spectrometry; IP: Immunoprecipitation.
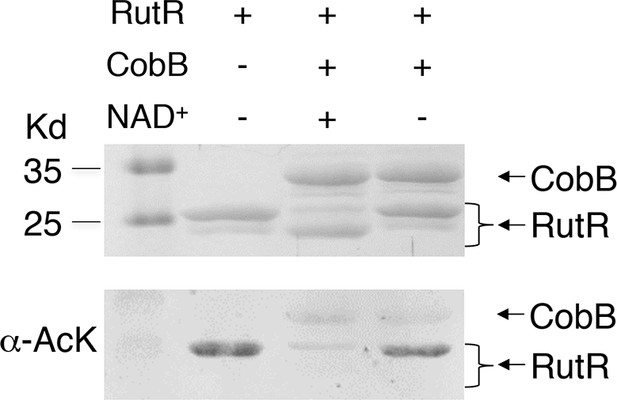
Without NAD+ , CobB could not deacetylate RutR.
In vitro assays of the KDAC activity of CobB on RutR demonstrated that its KDAC activity is dependent on NAD+. Incubation with CobB and NAD+ almost completely abolished the slower migrating acetylated RutR bands (upper panel) as evidenced by immonublotting (lower panel).
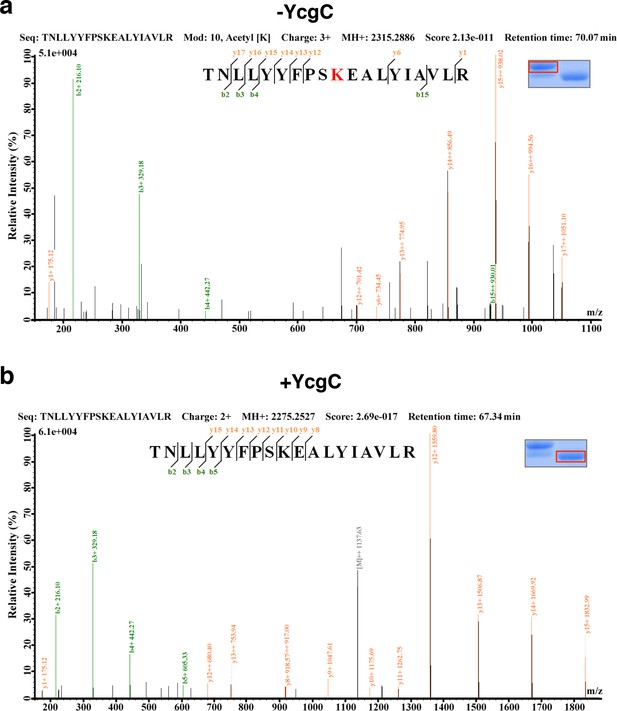
LC-MS/MS analysis to determine the residues of RutR deacetylated by YcgC.
RutR was treated with YcgC first and used the untreated RutR as the control. Both these two samples were resolved on a SDS-PAGE gel side by side. The upper band represents the Kac-containing starting materials and the lower band represents the K-containing product, which was then cut and subjected to MS analysis (the inlet). (a) Lys62 was identified as an acetylated site in RutR. (b) After incubating with YcgC, acetylation on K62 was no longer detectable.
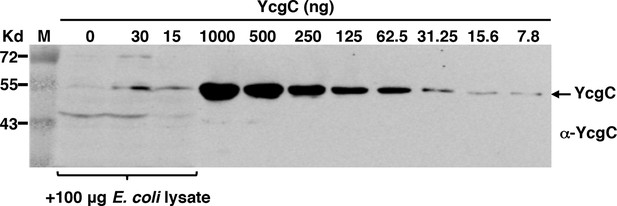

Specificity and sensitivity of the custom-made YcgC monoclonal antibody as assessed by Western blotting.
Specificity was measured by spiking an E. coli total lysate with affinity purified YcgC, and sensitivity by testing serially diluted YcgC.
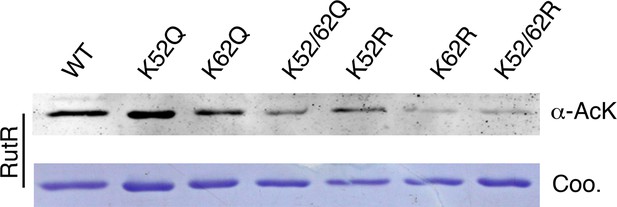
Mutagenesis of RutR confirmed that K52 and K62 are acetylated in vivo.
Four single mutants K52Q, K62Q, K52R, and K62R and two double mutants K52/62Q and K52/62R were constructed. These mutants along with WT RutR were produced and purified in E. coli. Equal amounts of purified proteins were Western blotted with the α-AcK antibody.
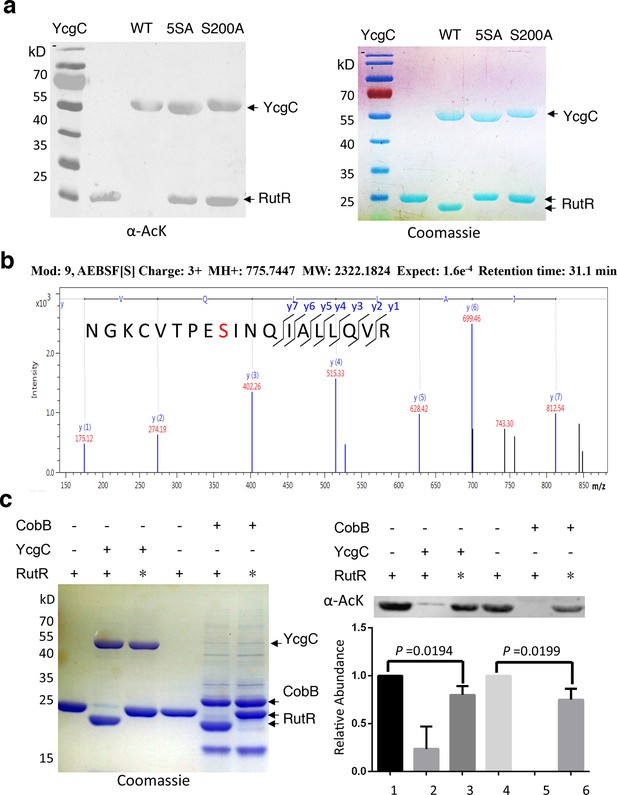
S200 is critical for YcgC’s deacetylation activity.
(a) Two mutants of YcgC, that is, S S8/10/73/77/200A and S200A were constructed through gene synthesis. In vitro assays of the KDAC activity of these two mutants on RutR demonstrated that their KDAC activities were completely abolished. (b) S200 on YcgC was identified as an AEBSF binding site by LC-MS/MS analysis. YcgC was incubated with AEBSF, then trypsin digested and subjected to MS/MS analysis. Upon AEBSF-mediated sulfonation, a 183 Da molecular weight increase is predicted. (c) In vitro assays of the KDAC activity of YcgC on heat-denatured RutR demonstrated that YcgC was still active (right panel), while the downshift band disappeared (left panel). Similar results were observed when heat-denatured RutR was treated with CobB. KDAC: Lysine deacetylase; LC-MS/MS: Liquid chromatography–mass spectrometry; AEBSF: 4-(2-Aminoethyl)benzenesulfonyl fluoride; WT: Wild type.
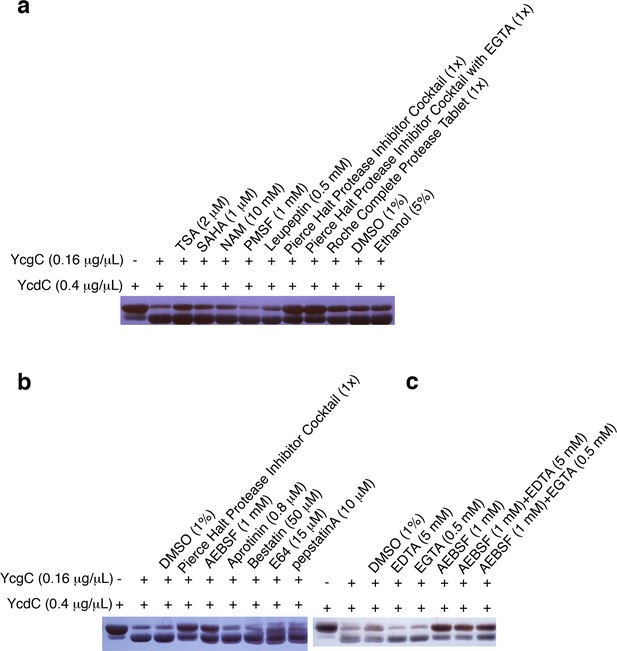
YcgC’s protein deacetylase activity is inhibited by AEBSF.
(a) YcgC’s deacetylase activity on YcdC was monitored when a variety of hydrolase inhibitors were added individually. The solvents of these inhibitors, that is, DMSO and ethanol, were also included as controls. (b) Testing the individual component of the Pierce Halt protease inhibitor cocktail revealed that AEBSF inhibits the protein deacetylase activity of YcgC. (c) YcgC’s protein deacetylase activity is not affected by both EDTA and EGTA.
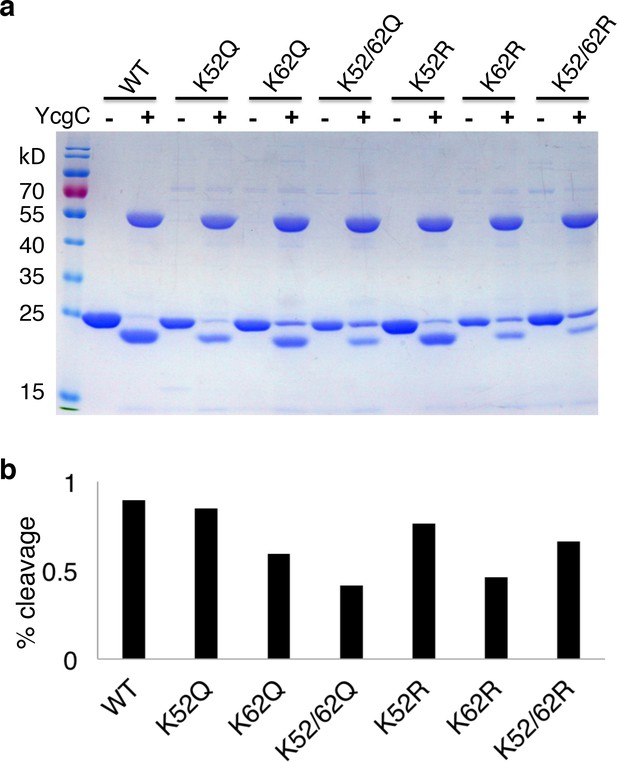
Lysine 62 is critical for RutR proteolysis.
(a) Mutant proteins of K52R, K62R, K52/62R, K52Q, K62Q, and K52/62Q with WT, RutR proteins were treated with YcgC and examined with Coomassie stain. (b) The ratio of cleavaged RutR to intact RutR was also calculated.
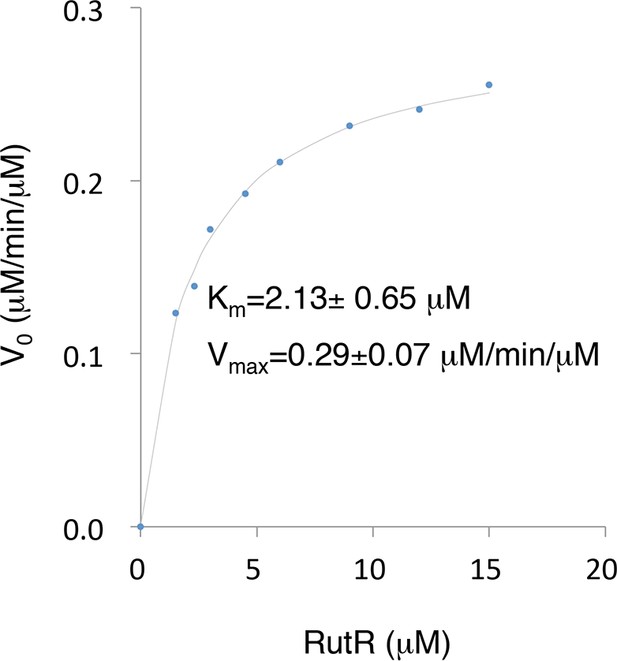
The Km and Vmax values of YcgC were determined using RutR as a substrate.
Because the N-terminal cleavage of RutR is tightly coupled with its deacetylation by YcgC, the downshifted band of RutR in the YcgC deactylation reaction could be conveniently used as a surrogate of YcgC’s activity. YcgC was incubated with serially diluted acetylated RutR. Deacetylation was determined by measuring the intensity of the lower bands on a silver-stained gel.
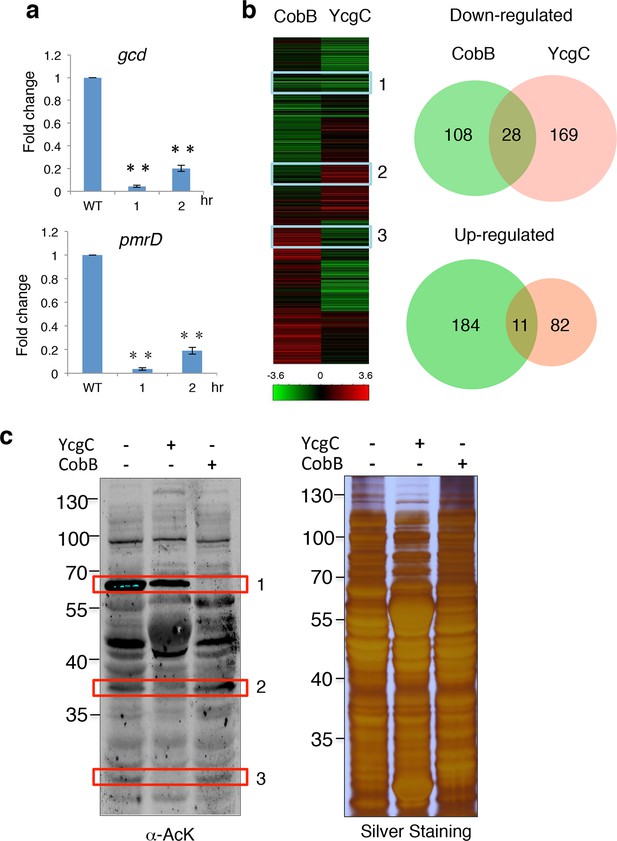
YcgC and CobB target distinct sets of substrates.
(a) YcgC regulates gene expression via deacetylating RutR. Expression of gcd and pmrD is significantly reduced upon RutR induction over a period of 2 hr as measured by quantitative real-time PCR. Double asterisks indicate that the observed fold changes are statistically significant, p<0.01. (b) Global gene expression analysis of ycgC- and cobB-induced cells. Clustering analysis shows clearly that impact on global transcription of induction of ycgC is distinct from that of cobB. Venn diagram showing that there is no significant overlap between genes down- and up-regulated due to CobB and YcgC induction. (c) Overexpression of YcgC affects global protein acetylation levels in E. coli. After ycgC and cobB were separately induced for 1 hr, global acetylation was detected in whole lysates of Escherichia coli using two pan α-AcK antibodies. The WT E. coli strain was also included for comparison. Boxed areas indicate regions that show obviously different staining patterns in ycgC- and cobB-induced cells. PCR: Polymerase chain reaction; WT: Wild type.
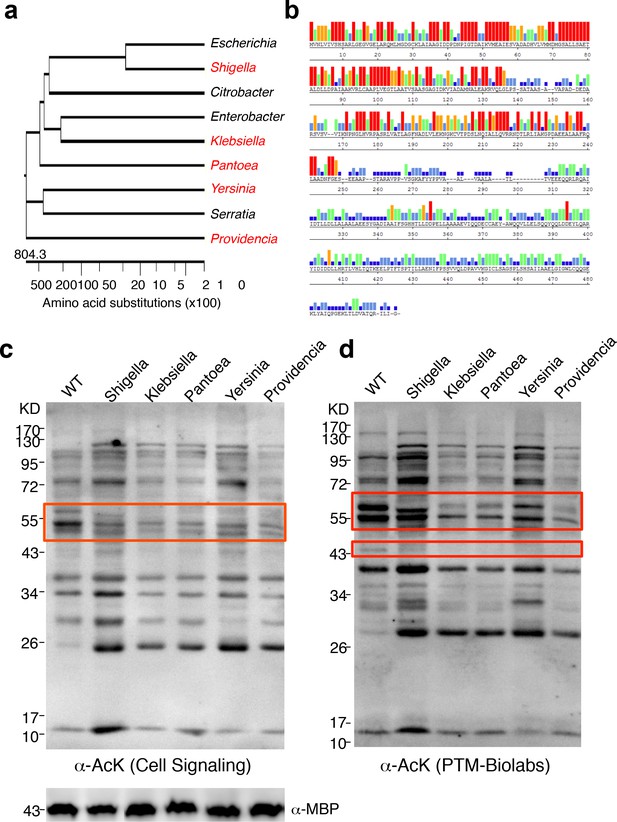
YcgC represents a new family of KDACs.
(a) Five representative YcgC homologs with protein sequence homology ranging from low to high. (b) Amino acid sequence homology analysis between YcgC and five selected YcgC homologs from other bacteria. The consensus strength among the six homologous proteins at each amino acid position of YcgC is indicated with colored bars. Red, orange, green, light blue, dark blue, and blank bars represent 100, 80, 60, 40, 20, and 0% consensus strength, respectively. (c,d) Changes in global E. coli acetylation profiles upon induction of the five YcgC homologs. The five selected YcgC homologs were cloned, transformed into E. coli, and induced to overexpress. Global acetylation profiles of each induced strain were detected with a pan monoclonal antibody (Cell Signaling, #9441) and a pan polyclonal antibody (PTM-Biolabs, PTM-105), as shown in c and d, respectively. WT E. coli cells were also processed in parallel as a comparison. An antibody against myelin basic protein was used as a loading control. WT: Wild type.
Additional files
-
Supplementary file 1
Genes differentially expressed when YcgC was overexpressed.
- https://doi.org/10.7554/eLife.05322.016
-
Supplementary file 2
Genes differentially expressed when CobB was induced.
- https://doi.org/10.7554/eLife.05322.017
-
Supplementary file 3
Clones constructed in this study.
- https://doi.org/10.7554/eLife.05322.018
-
Supplementary file 4
Primers used in this study.
- https://doi.org/10.7554/eLife.05322.019





