A mutant Escherichia coli that attaches peptidoglycan to lipopolysaccharide and displays cell wall on its surface
Figures
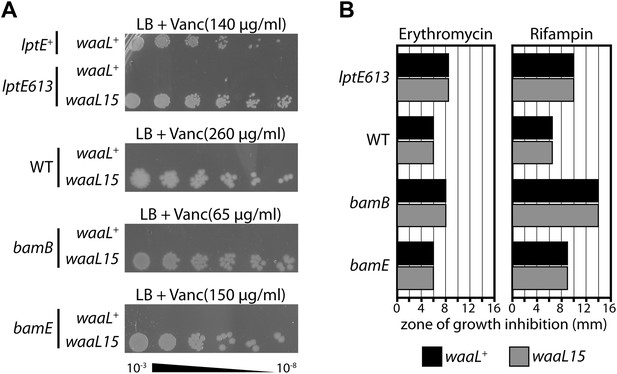
A mutant O-antigen ligase increases vancomycin resistance.
(A) waaL15 provides a strain-independent increase in vancomycin resistance. Isogenic strains, differing by a point mutation in waaL, were plated by serial dilution on LB agar containing indicated amounts of vancomycin. (B) waaL15 does not improve resistance against other antibiotics. Antibiotic discs containing either 15 μg erythromycin or 5 μg rifampin were placed on LB agar overlays inoculated with the indicated strains. Diametric zones of growth inhibition were measured across the disc. The disc diameter was 6 mm and this value represents growth at the disc.
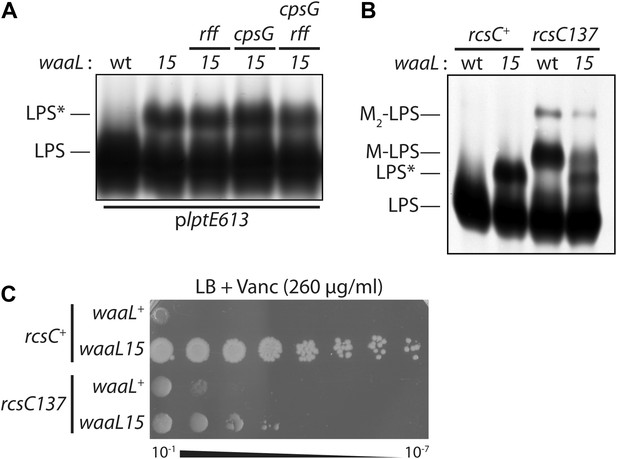
Mutant O-antigen ligase produces a novel form of LPS that is directly responsible for vancomycin resistance.
(A) WaaL15 uses a novel substrate to produce a new LPS glycoform. Isolated LPS was resolved by SDS-PAGE and detected by silver staining. A higher molecular weight glycoform (LPS*) appears in waaL15 strains. Mutations that inactivate biosynthesis of ECA (rff::Tn10-66) or CA (ΔcpsG::kan) do not abrogate LPS* production. (B) Overproduction of CA leads to decreased LPS* abundance. Isogenic strains were constructed to express either wt rcsC+ or the rcsC137 mutant allele that hyper-activates CA biosynthesis. LPS was isolated and visualized as in (A). LPS molecules modified with a one- or two- CA repeat units are labeled M-LPS and M2-LPS, respectively. (C) Reduced LPS* levels correlate with reduced vancomycin resistance. Strains were plated by serial dilution onto LB agar supplemented with vancomycin.
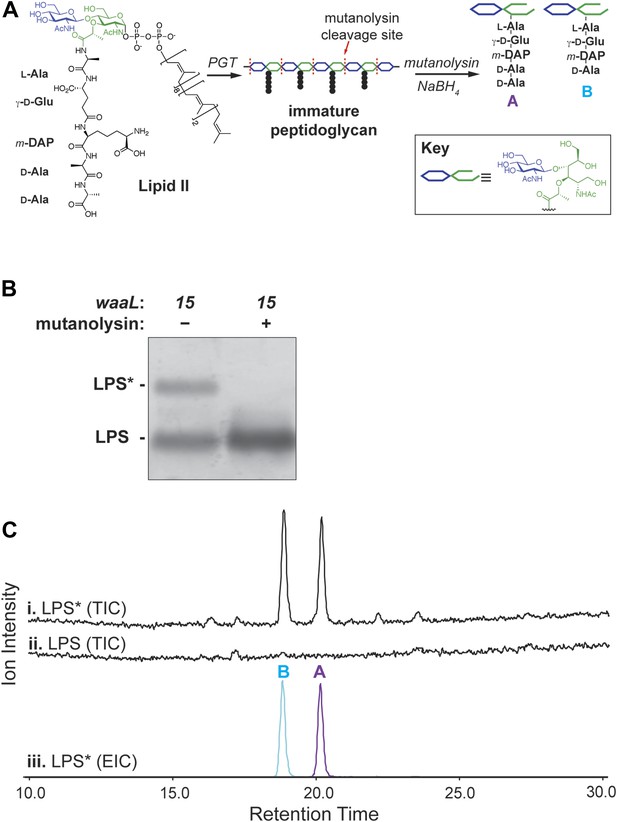
Lipid II is the glycosyl donor for LPS*.
(A) Structure of lipid II and schematic of peptidoglycan cleavage by mutanolysin that releases disaccharide pentapeptide (‘A’) and tetrapeptide (‘B’) species. (B) Treatment of waaL15 isolated LPS with mutanolysin cleaves the LPS* modification. (C) LPS* is glycosylated with equivalent amounts of lipid II-sourced disaccharide pentapeptide and tetrapeptide. Isolated and purified LPS* from waaL15 and LPS from waaL+ were treated with mutanolysin and analyzed by LC-MS. Total ion chromatogram for degradation products (i and ii), and the extracted ion chromatogram for LPS* degradation (iii) are shown. M+H and (M+2H)/2 ions corresponding to each fragment were extracted (A: 1013.3 + 507.2; B: 942.3 + 471.7).
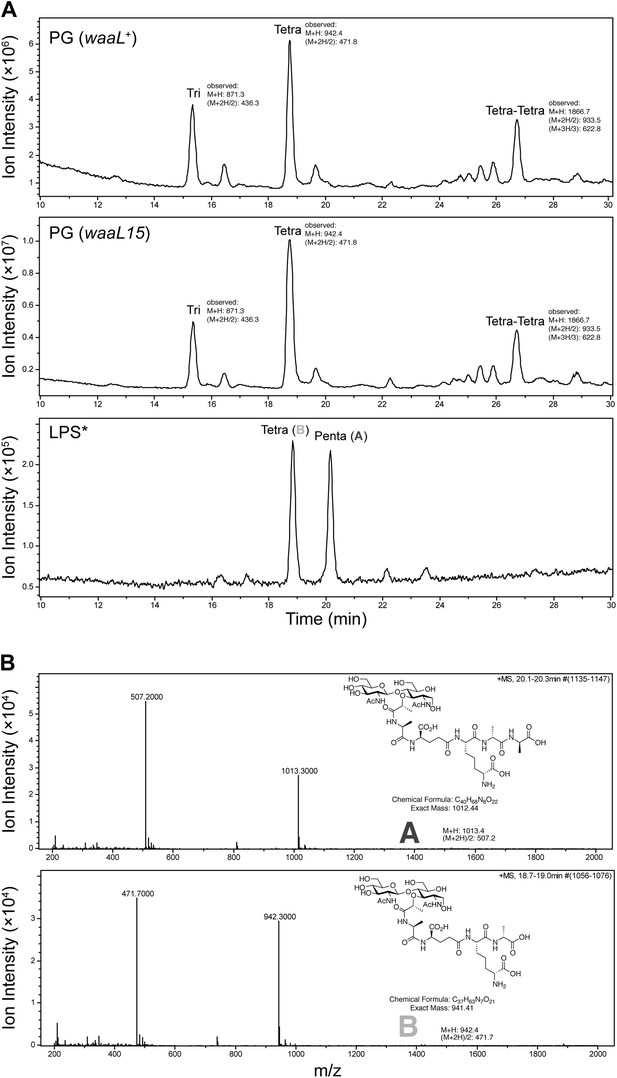
The waaL15 mutation does not affect the PG cell wall.
(A) The total ion chromatograms (TIC) of mutanolysin digested PG cell wall from waaL+ strain MG1210, and waaL15 strains MG1211 are nearly identical. Disaccharide pentapeptide (DPP, fragment A) is present in mutanolysin digested LPS* but not in the PG cell wall samples. (B) Mass spectra and structures for reduced DPP (fragment A) and reduced disaccharide tetrapeptide (fragment B) from mutanolysin digested LPS*.
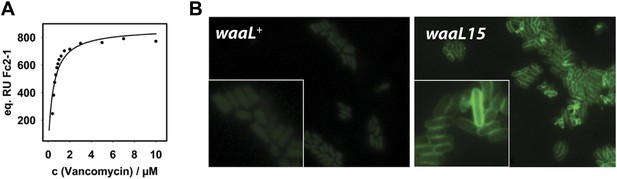
Mutant WaaL attaches peptidoglycan fragments to LPS.
(A) LPS* specifically binds vancomycin. Purified LPS* was immobilized on a CM3 chip and varying concentrations of vancomycin were applied. Binding was measured at 25°C by surface plasmon resonance. Fitting of equilibrium signal yielded a Kd = 0.48 ± 0.08 µM. Standard deviation was measured for 0.6 µM and 1.2 µM and was ±1 RU. (B) Vancomycin binds to LPS* at the cell surface. Live exponential-phase growing cells labeled with 1 μg/ml vancomycin-BODIPY for 10 min. Cells were spotted onto M63 minimal medium agar pads and imaged by fluorescence microscopy.
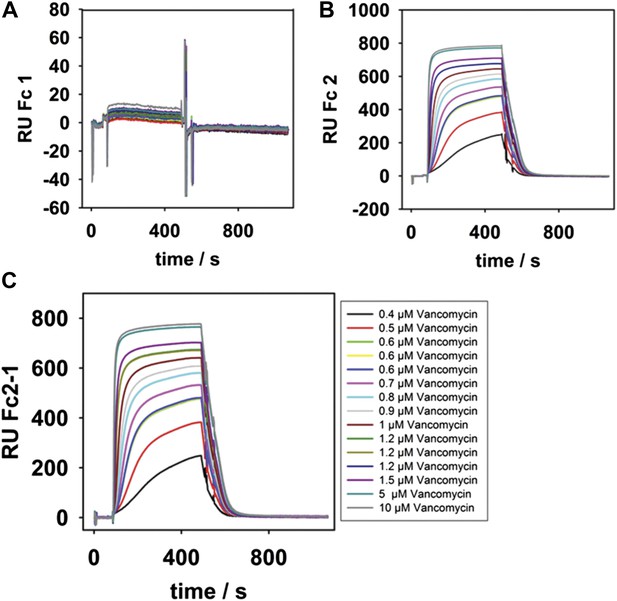
SPR binding kinetics at 25°C.
(A and B), Different concentrations (see legend inset in C) of vancomycin were passed over surfaces of total isolated LPS from waaL+ (strain MG1210) or total isolated LPS from waaL15 (strain MG1211). (C), The reference subtracted kinetics after 400 s of injection. The equilibrium signals were plotted in Figure 4A.
Additional files
-
Supplementary file 1
(A) Synthetic interactions between waaL15 and mutations affecting the elongasome due to limited lipid II availability. (B) Strains used in this study. (C) Plasmids used in this study.
- https://doi.org/10.7554/eLife.05334.009





