Recognition of the small regulatory RNA RydC by the bacterial Hfq protein
Figures
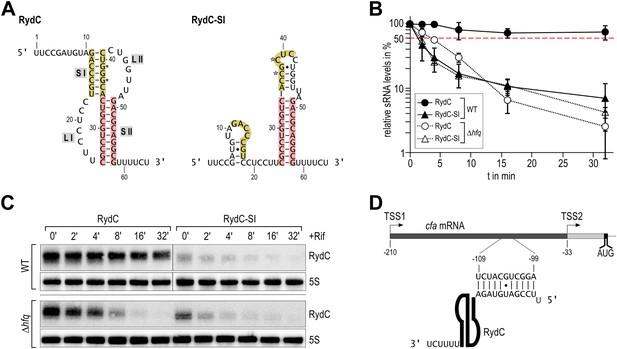
Association of RydC with the chaperone Hfq and its pseudoknot fold are required for sRNA stability.
(A) Secondary structure of the RydC pseudoknot and the predicted alternative structure are formed by the double mutant RydC-SI. Substitutions G37C and G39C are indicated with asterisks. (B, C) Stabilities of RydC and RydC-SI were determined by Northern blot analyses. Total RNA samples were extracted prior to and at indicated time-points after inhibition of transcription by rifampicin in late exponential phase (OD600 of 1) from Salmonella strains ΔrydC (JVS-0291) or Δhfq (JVS-0584), carrying plasmids expressing RydC (pKF42-1) or RydC-SI (pKF60-1) from the constitutive PL promoter. Error bars represent standard deviation calculated from three biological replicates. (D) Predicted duplex formed between the RydC 5′-end (nts 2–11) and the longer isoform cfa mRNA (nts −99 to −109 relative to the translational start site) originating at transcriptional start site 1 (TSS1).
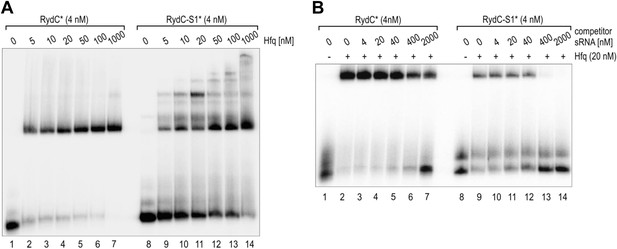
Electrophoretic mobility shift assay (EMSA) of Hfq with RydC or RydC-S1.
(A) Electrophoretic mobility shift assay (EMSA) with in vitro synthesized 5′-end-labelled RydC and RydC-S1 RNAs (RydC* and RydC-S1*, 4 nM) in the presence of increasing concentrations of Hfq protein as indicated. (B) Preformed Hfq/RNA*(RydC* and RydC-S1*, 4 nM) complexes were incubated with increasing concentrations of cold competitor RNA (RydC-S1 for RydC*; RydC for RydC-S1).
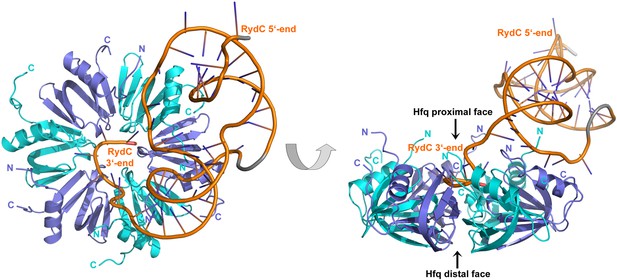
Crystal structure of the RydC/Hfq complex.
The content of the asymmetric unit is shown. The left panel presents a view along the molecular sixfold axis of the Hfq ring core (roughly encompassing residues 3–72), while the right panel is viewed along the perpendicular direction. The 3′-end poly-U tail of RydC is inserted in a groove on the proximal face of the hexamer. The N- and C-terminal regions are indicated with letters (cyan and blue for different Hfq protomers); these are natively unstructured and in multiple conformations, but some portions could be modelled. The RydC phosphate backbone is shown as an orange cartoon, bases are shown as sticks. Nucleotides 20–21 are disordered, but the backbone was modelled to help view the RNA structure and is indicated in grey cartoon style, as is also the last modelled phosphate at the 5′-end of RydC.
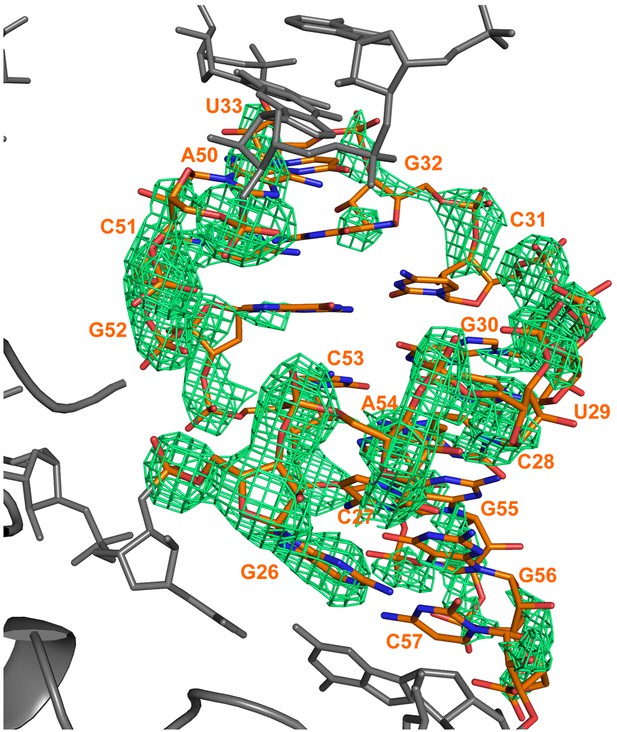
Electron density calculated after refinement of a model in which a portion of a RNA duplex region was omitted (nucleotides 26–33 and 50–57).
The model was refined with REFMAC for 10 cycles and a Fo–Fc map calculated. The difference map is shown in green and is contoured at 3σ. The refined position of the RNA is superimposed on the map. The omitted RydC is shown in orange stick representation, while the rest of RydC and Hfq are coloured grey.
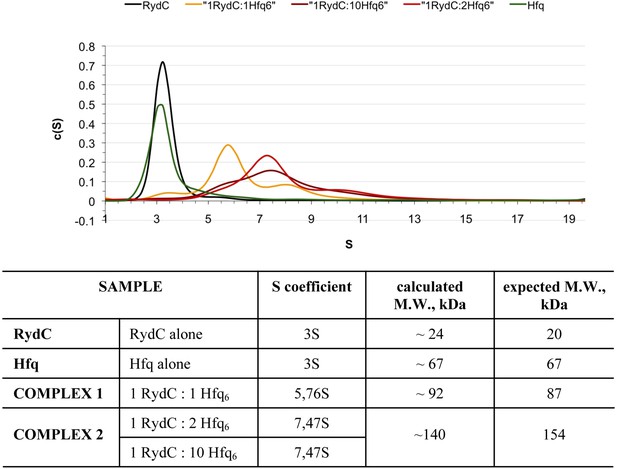
Analytical Ultracentrifugation of Hfq, RydC, and Hfq–RydC mixtures at different RNA:protein ratios.
A 1:1 complex of RydC:Hfq6 can be detected (yellow trace) when the two components are mixed in a 1:1 ratio, but at higher ratios of RydC:Hfq, a 1:2 RydC:Hfq6 can form as well. Protein–RNA mixtures containing different molar ratios (1:1, 1:2, 1:10) of RydC to Hfq hexamer were incubated at room temperature for 30 min in 10 mM Tris–HCl, 20 mM NaCl, 10 mM EDTA, 5 mM DTT. The concentration of RydC in each experiment was 1.5 μM. Hfq alone was also run at a concentration of 173 μM. The sedimentation of the different species was followed both at 260 nm and 275 nm, corresponding respectively to the maximum absorbance of RNA and Hfq. The program Sedfit was used to extrapolate the sedimentation coefficients (S) for RydC, Hfq, and the protein–RNA complexes and their observed mass, after assuming that vHfq = 0.7248, vRydC = 0.5500 and after calculating the vcomplex with the following equation: vcomplex = (MWprotein × vprotein + MWRNA × vRNA)/(MWprotein + MWRNA) = 0.6835.
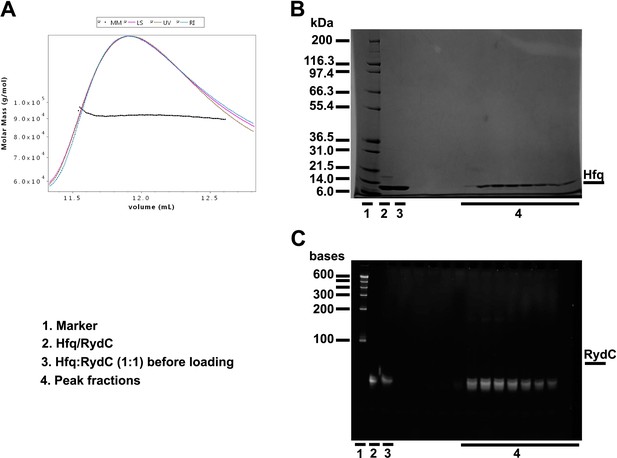
Size exclusion chromatography dynamic light scattering (SEC-MALS).
(A) Elution profile for the RydC:Hfq complex from a Superdex 200 column superimposed on the estimated molecular masses. Fractions over the peak were analysed with a denaturing protein gel (B) and a denaturing RNA gel (C), confirming that both RydC and Hfq are present in the peak. The table in the lower panel summarises the results of the SEC-MALS data evaluations. SEC-MALS measurements were made with a Superdex 200 10/300 column connected to a miniDAWN TREOS multi-angle static Light Scattering and an Optilab T-rEX (refractometer with EXtended range) detector (Wyatt Technology Corporation, Santa Barbara CA, USA). Mixtures of protein and RNA in different molar ratios were run through an S200 10/300 size exclusion chromatography column equilibrated in buffer A (50 mM Tris, pH 7.5, 50 mM NaCl, 50 mM KCl, 5 mM MgCl2, 2 mM DTT). Both RydC and Hfq were diluted in buffer A. The RNA was first annealed (90°C for 2 min, 30 min at room temperature) before mixing it with an equal molar amount of Hfq6. The final concentrations of RydC and Hfq6 in the mixture were equal to 30 μM; Hfq alone was at a molar concentration of 50 μM. BSA (2 mg/ml) was run as positive control. Predicted and observed molecular weights for Hfq and Hfq/RydC complex are listed below.
| Sample | Predicted molecular weight | Observed molecular weight* | Polydispersity |
|---|---|---|---|
| Hfq | 66,996 Da | 64,070 Da (±0.101%) | 1.000 (±0.142%) |
| Hfq:RydC 1:1 | 88,895 Da | 91,650 Da (±0.218%) | 1.000 (±0.308%) |
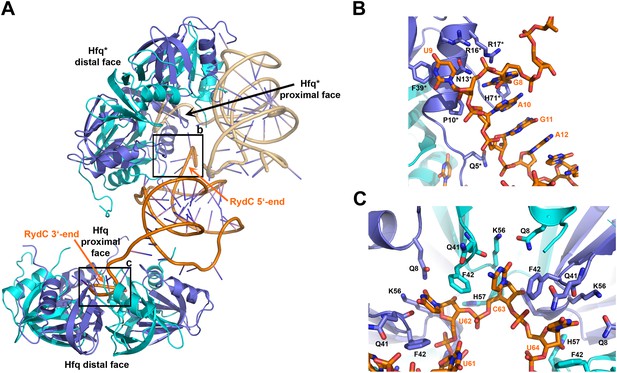
Interactions established by the 3′-end and 5′-end regions of RydC with two adjacent Hfq hexamers in the crystal lattice.
(A) RydC is sandwiched between two Hfq hexamers in the crystal lattice. Two asymmetric units are shown with the RydC in the centre (coloured in orange) bridging two Hfq hexamers. The symmetry-related Hfq molecule is labelled with an asterisk (*). The 3′-end region of RydC in the main asymmetric unit is pointed by an arrow and contacts the proximal face of the principal Hfq hexamer, another arrow indicates the position of the 5′-end of RydC, interacting with a portion of the rim of the Hfq hexamer in the neighbouring asymmetric unit. Two black rectangles define the 5′-end and 3′-end RydC–Hfq contact regions, which are enlarged in panels B and C respectively. (B) Contacts established between the 5′-end seed region of RydC and the rim of a symmetry related Hfq molecule. The main interactions at this interface involve the following amino acid–nucleotide pairs: R17-G8, H71-G8, R16-U9, N13-U9, F39-U9, Q5-G11, P10-A10. Residues belonging to the symmetry related Hfq molecules are labelled with an asterisk (*). (C) Interactions of the poly-U 3′-end of RydC with the recessed groove of the proximal face of Hfq. The uridines and cytosine stack on F42; this and other interactions are similar to those seen in the U6/Hfq crystal structure (Sauer and Weichenrieder, 2011). The side chains of critical residues that contribute to keep RydC in the channel are shown in stick representation. The adjacent protomers of Hfq are coloured in blue and cyan, RydC is shown in orange. Side chains are shown for all residues, main chains are shown for Q41 in order to highlight the position of atoms involved in interactions with neighbouring residues. The RNA used in the crystallisations has a guanine on the 3′-end originating from the template for in vitro transcription, but this base may be in multiple conformations.
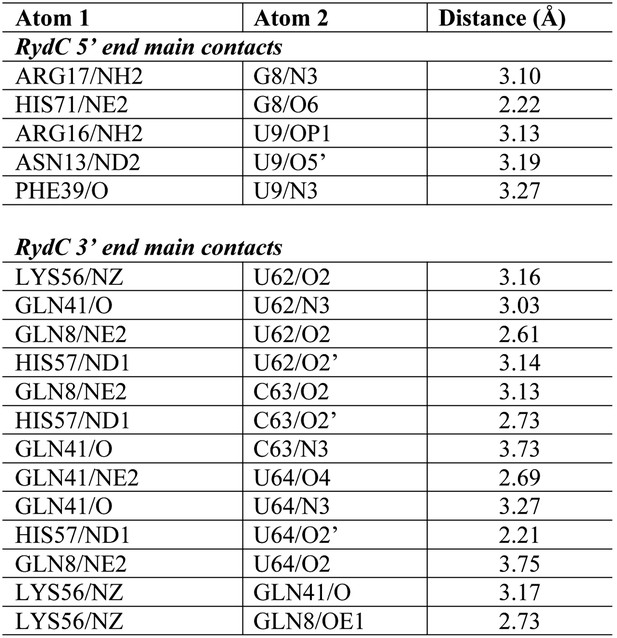
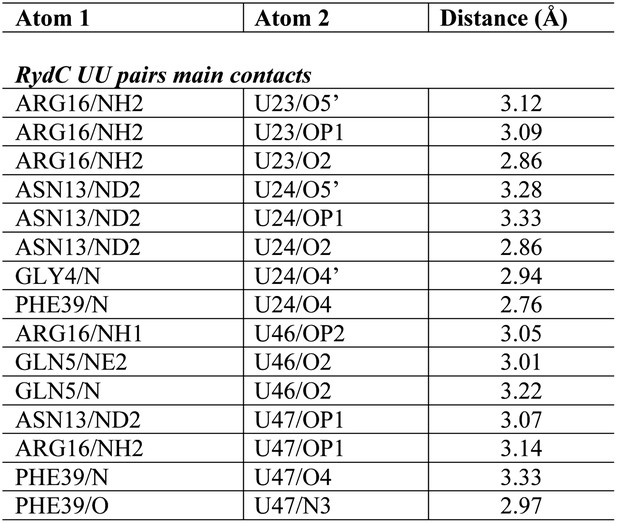
Distances of the main contacts between Hfq and the 5′- and 3′-ends of RydC.
https://doi.org/10.7554/eLife.05375.010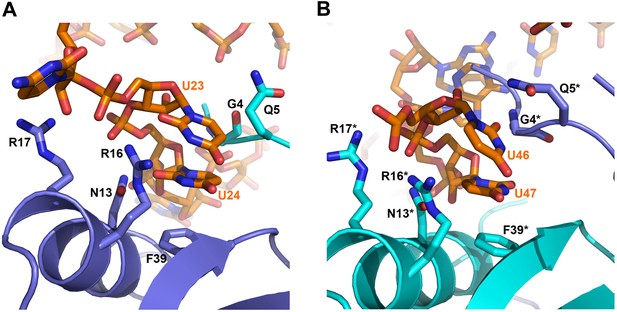
Contacts between the conserved U–U pairs of RydC and the Hfq rim.
(A) The U23/U24 pair interacts with F39, R16, R17, and N13 on the convex rim of Hfq. The U23/U24 interactions with the rim are consolidated by the N-terminal residues G4 and Q5 of a neighbouring Hfq protomer (coloured in cyan). (B) The U46/U47 pair makes similar interactions with Hfq in a neighbouring asymmetric unit. Residues belonging to the symmetry related Hfq molecule are labelled with an asterisk (*) in panel B. Side chains are shown for all residues, main chain is shown for G4 in both panels to help visualize the residue position.
Distances of the main contacts between Hfq and the U23/U24 and U46/U47 base steps of RydC.
https://doi.org/10.7554/eLife.05375.012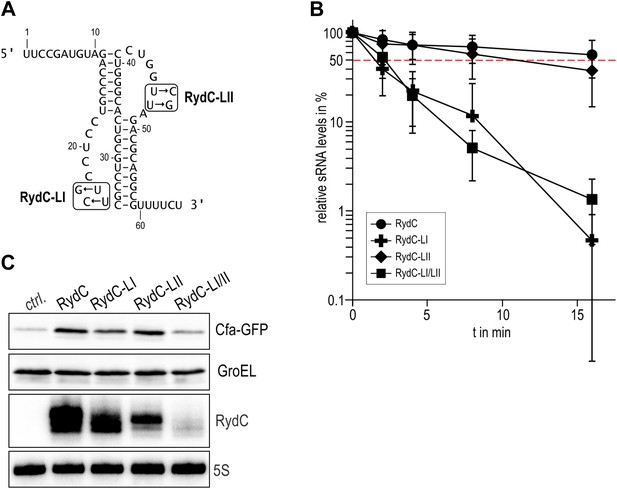
Requirement for the conserved UU steps for in vivo stability of RydC.
(A) Schematic of RydC pseudoknot fold with substitutions of U23G and U24G (RydC-LI) and U46G and U47G (RydC-LII). (B) Stabilities of RydC and its variants RydC-LI, RydC-LII, and RydC-LI/LII. RNA levels were determined by Northern blot analyses. Total RNA samples were extracted prior to and at indicated time-points after inhibition of transcription by rifampicin in late exponential phase (OD600 of 1) from Salmonella strain ΔrydC (JVS-0291) carrying plasmids expressing RydC (pKF42-1), RydC-LI (pKF224-9), RydC-LII (pKF223), or RydC-LI/II (pKF225) from the constitutive PL promoter. Error bars represent standard deviation calculated from three biological replicates. (C) Target regulation by RydC loop mutants. Salmonella ΔrydC (JVS-0291) express a plasmid-borne translational fusion of the RydC target cfa to the green fluorescent protein (Cfa–GFP; pKF31-1) in the presence of either a control (pJV300) or plasmids expressing RydC (pKF42-1), RydC-LI (pKF224-9), RydC-LII (pKF223), or RydC-LI/II (pKF225) from the constitutive PL promoter. Bacteria were grown to late exponential phase (OD600 of 1) and Cfa–GFP and GroEL proteins were detected by Western blot (upper two panels). RydC variants as well as 5S RNA were detected by Northern blot (lower two panels).
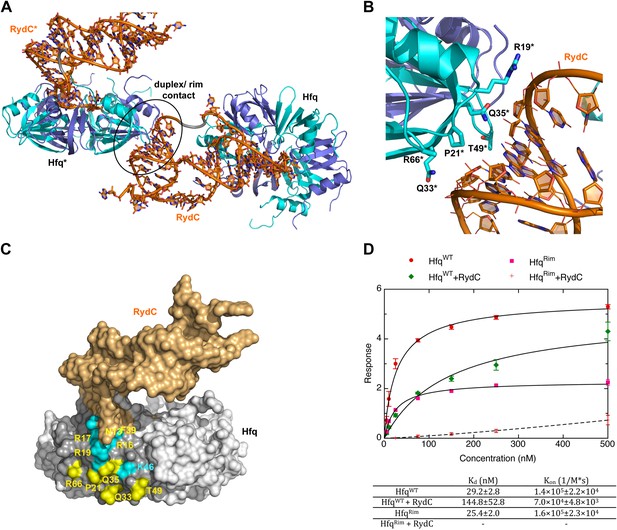
A potential RNA interaction surface on the circumferential rim of the Hfq hexamer.
(A) Interaction between a duplex region of RydC and the rim surface of an Hfq molecule belonging to the neighbouring asymmetric unit in the crystal. Symmetry related RydC and Hfq are labelled with an asterisk (*). (B) A detailed view into the interaction site, showing the main protein residues making contacts with the duplex RNA. The side chain of R66 is not shown as it is disordered in the structure. (C) A surface representation of the RydC/Hfq complex, highlighting in colour in one protomer, the residues of the proposed extended rim that recognises RNA. The residues coloured cyan are from earlier studies that identified the rim as an RNA binding region (R16, R17, R19, K46; Sauer et al., 2012), and the yellow residues include an extended surface proposed here to engage duplex RNA (P21, Q33, Q35, T49, R66). Also labelled are the residues of this extended surface that engage the U–U steps in RydC (N13, R16, R17, F39; F39 and N13 are not visible in this view). (D) Rates and equilibrium constants for binding of Hfq and Hfq/RydC to cfa for wild-type (HfqWT) and rim-mutant protein (HfqRim; R19A, P21A, Q33A, Q35A, T49A, R66A). The measurements were performed by fitting the averaged responses from three independent experiments. It was not possible to fit the weaker binding response of HfqRim binding RydC.
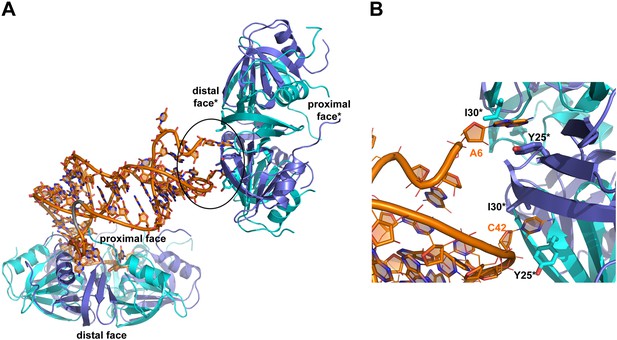
Interactions between RydC and the proximal face of a symmetric Hfq molecule.
(A) Hfq belonging to a neighbouring asymmetric unit is shown at the top right with the RNA contacting its distal face. (B) An expanded view into the distal face showing nucleotides A6 and C42 bound to the two pockets formed by residues I30 and Y25. Crystallographic and solution binding studies show that E. coli Hfq has preferences to bind an A-A-N motif on the distal face (Robinson et al., 2013). The contacts observed for RydC A6 and C42 with the distal are fortuitous lattice interactions, but they mimic interactions of the A-A-N repeats on the distal face.
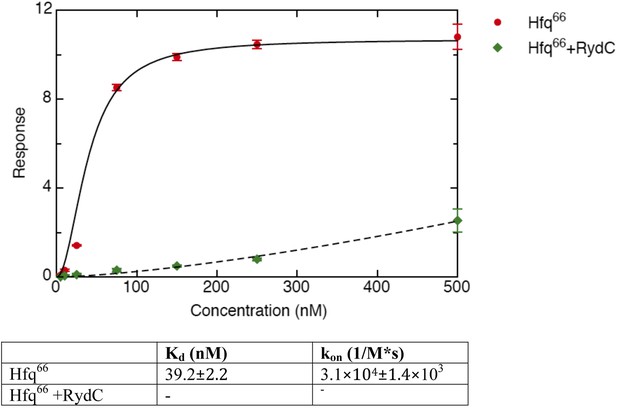
Binding rates and equilibrium constants for Hfq66 and Hfq66/RydC to immobilized cfa.
The measurements were performed by fitting the response from three independent experiments.
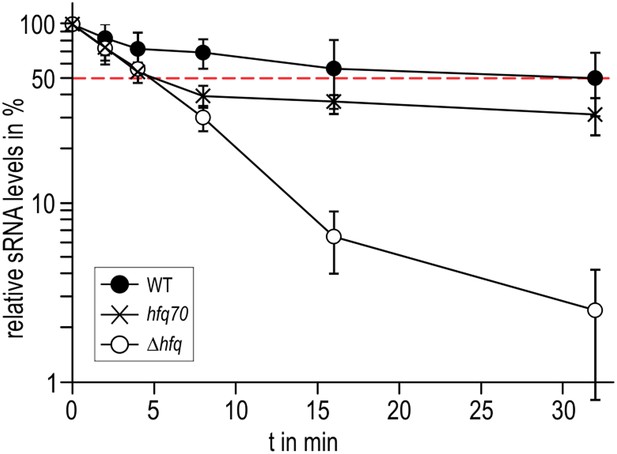
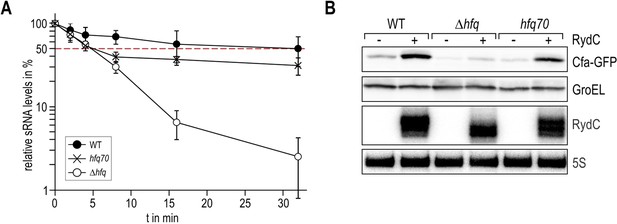
Role of the Hfq C-terminus in RydC stability in vivo.
Stabilities of RydC were determined by Northern blot analyses in Salmonella strains ΔrydC (WT; JVS-0291), ΔrydC Δhfq (Δhfq; JVS-10665), and ΔrydC hfq70 (hfq70; JVS-11150) expressing RydC from the constitutive PL promoter (pKF42-1). Total RNA samples were extracted prior to and at indicated time-points after inhibition of transcription by rifampicin in late exponential phase (OD600 of 1). Error bars represent standard deviations calculated from three biological replicates.
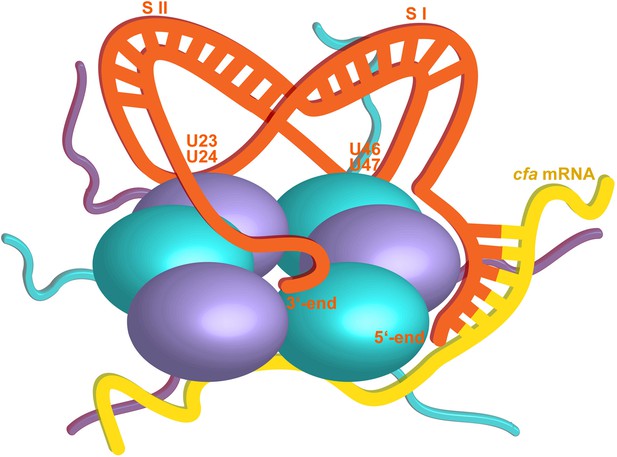
Hypothetical model of the RydC/mRNA/Hfq effector complex.
Portions of the protomers forming the Hfq core have been represented with six spheres, from which the disordered C-terminal tails extend radially. RydC (orange) sits on the proximal face of Hfq, with the 3′-end U-rich tail interacting with the central channel and the two conserved U–U pairs (U23/U24 and U46/U47) making contacts with the lateral face of the hexamer. The two double strands conferring the pseudoknot structure to RydC are indicated as S I and S II. The target mRNA cfa, depicted in yellow, associates with the distal face of Hfq, and it is proposed to form a duplex with the 5′-end ‘seed’ region of RydC that is recognised by the circumferential rim of Hfq. The six long C-termini are depicted in the cartoon, and the shorter N-termini are not shown for clarity. The association of the two RNA partners is aided by the C-terminal tails of Hfq, which extend towards the RNA molecules, embracing them and stabilizing the seed/target pairing. The natively unstructured termini make distributed interactions with the RNA, visiting dynamically multiple points on the nucleic acid.
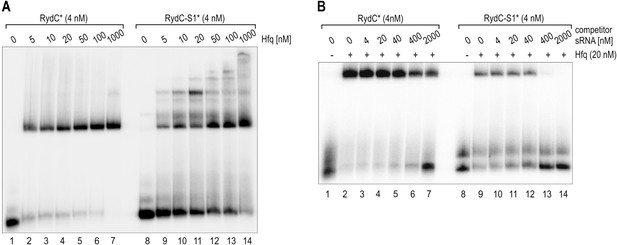
(A) Electrophoretic mobility shift assay (EMSA) with in vitro synthesized, 5' end-labelled RydC and RydC-S1 RNAs (RydC* and RydC-S1*, 4 nM) in the presence of increasing concentrations of Hfq protein as indicated. (B) Preformed Hfq/RNA*(RydC* and RydC-S1*, 4 nM) complexes were incubated with increasing concentrations of cold competitor RNA (RydC-S1 for RydC*; RydC for RydC-S1).
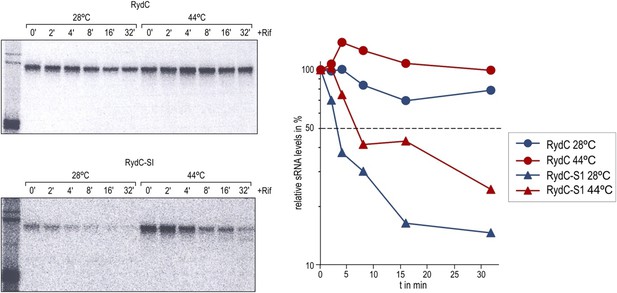
Stabilities of RydC and RydC-S1 were determined in Salmonella rne-ts by Northern blot analysis of total RNA samples withdrawn prior to and at indicated time-points after inhibition of transcription by rifampicin (OD600 of 1). In the temperature-sensitive rne-ts Salmonella strain, RNase E retains its normal capacity when grown at the permissive temperature of 28°C. As cells are shifted to 44°C, RNase E undergoes a conformational rearrangement rendering it catalytically inactive (Apirion, 1978; Figueroa-Bossi et al., 2009). To quantify the decay rates, the signal obtained at 0 min was set to 100, and the amount of RNA remaining at each time-point was plotted as a function of time. The time-point at which 50% of RNA had been decayed (dashed line) was calculated to determine the half-life (t1/2).
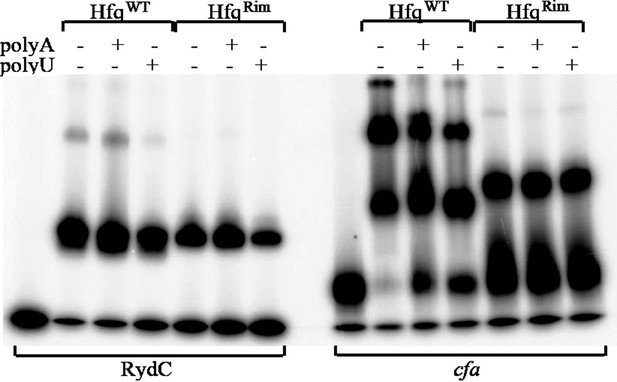
Hfq wild-type (HfqWT) and mutant (HfqRim) binding to RydC (left panel) and cfa (right panel). 10 nM of 5’ P32 labelled RNA was used with 20 nM Hfq and 25 nM polyA or polyU.
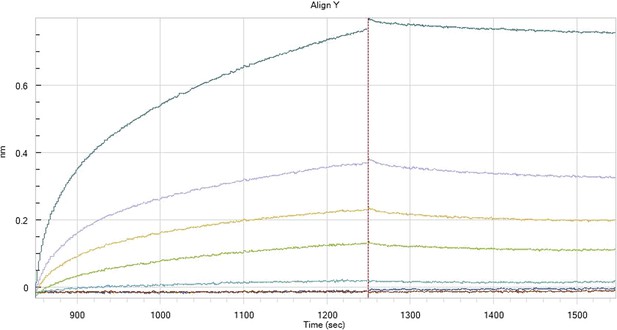
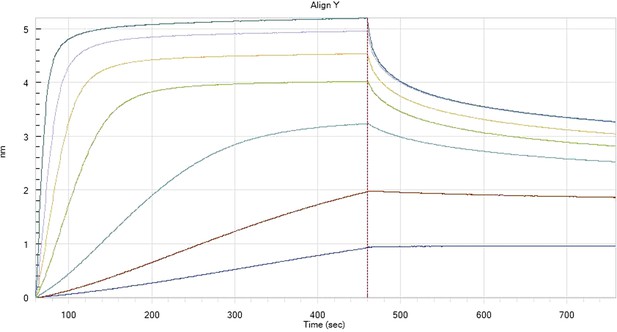
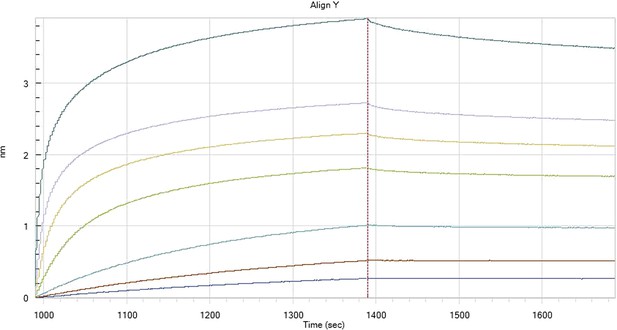
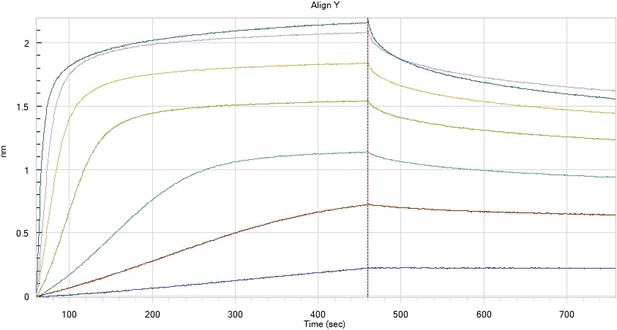
HfqRim binding to immobilised cfa in the presence of RydC: Traces correspond to different Hfq concentrations: 5 (dark blue), 10 (red), 25 (light blue), 75 (green), 150 (orange), 250 (light purple) and 500 nM (blue). First 400 seconds corresponds to association measurement, next 300 to dissociation.
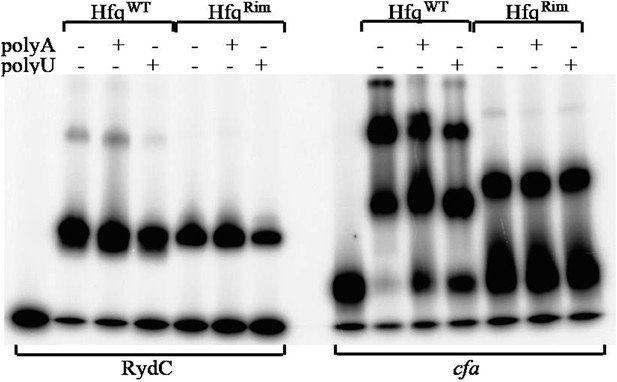
Hfq wild-type (HfqWT) and mutant (HfqRim) binding to RydC (left panel) and cfa (right panel). 10 nM of 5’ P32 labelled RNA was used with 20 nM Hfq and 25 nM polyA or polyU.
Tables
Oligonucleotides
| Name | Sequence 5′-3′ |
|---|---|
| RydC_T7_fwd | GTTTTTTTTTTAATACGACTCACTATAGGCTTCCGATGTAGACCCGT |
| RydC_T7_rev | CAGAAAACGCCTGCGTCTAACCAGGACCCG |
| JVO-0322 | CTACGGCGTTTCACTTCTGAGTTC |
| JVO-4363 | AGAAAACGCCTGCGTC |
| JVO-4558 | GTTTTTTTTAATACGACTCACTATAGGTTGTTTATATTACGATAATT |
| JVO-4721 | GTTTTTTTTTTAATACGACTCACTATAGGTTCCGATGTAGACCCGTCC |
| JVO-4722 | AGAAAACGCCTGCGTCTAAC |
| JVO-5165 | GTTTTTTTTTTAATACGACTCACTATAGGTTCCGATGTAGAGCGGTCC |
| JVO-9044 | CCCACGGACAATTCCGT |
| JVO-10909 | CAGCAACAATGCCGGTGGCGGCGCCAGCAATAACTACCATTAAGGTCCATATGAATATCCTCCTTAG |
| JVO-10910 | ATTATCCGACGCCCCCGACATGGATAAACAGCGCGTGAACGTGTAGGCTGGAGCTGCTTC |
| JVO-10913 | CGAGACGCAGGCGTTTTC |
| JVO-10914 | P-CCAGGACCCGTGACG |
| JVO-10915 | GCCGCCTGCGTCACGG |
| JVO-10916 | P-GGAGGACGGGTCTACATC |
| P: 5′ phosphorylation |
X-ray data collection and refinement statistics
| Data collection | |
| Space group | P212121 |
| Cell dimensions | |
| a, b, c (Å) | 71.94, 73.36, 137.95 |
| α, β, γ (°) | 90, 90, 90 |
| Resolution (Å) | 24.47–3.48 (3.69–3.48)* |
| Rmerge | 0.152 (0.886) |
| I/σI | 9.6 (2.3) |
| Completeness (%) | 99.1 (99.4) |
| Multiplicity | 5.8 (5.9) |
| Refinement | |
| Resolution (Å) | 24.32 (3.48) |
| No. reflections | 9190 |
| Rwork/Rfree | 0.22/0.28 |
| No. atoms | 4470 |
| Protein | 3238 |
| RNA | 1187 |
| Water | 45 |
| B-factors | |
| Overall | 94.69 |
| Protein | 93.36 |
| RNA | 122.60 |
| Water | 54.90 |
| R.m.s. deviations | |
| Bond lengths (Å) | 0.010 |
| Bond angles (°) | 1.475 |
-
*
Values in parentheses are for the highest-resolution shell.
Plasmids
| Plasmid trivial name | Plasmid stock name | Relevant fragment | Comment | Origin, marker | Reference | Used in Figure |
|---|---|---|---|---|---|---|
| ctrl. | pJV300 | pPL control plasmid, expresses an ∼50 nt nonsense transcript derived from rrnB terminator | ColE1, AmpR | (Sittka et al., 2007) | 5C | |
| pPL-RydC | pKF42-1 | RydC | Expresses Salmonella RydC from constitutive PLlacO promoter | ColE1, AmpR | (Fröhlich et al., 2013) | 1B/C; 5B/C; 7 |
| pPL-RydC-SI | pKF60-1 | RydC-SI | Expresses Salmonella RydC-SI (SNEs G37C; G39C) from constitutive PLlacO promoter | ColE1, AmpR | (Fröhlich et al., 2013) | 1B/C |
| pPL-RydC-LI | pKF224-9 | RydC-LI | Expresses Salmonella RydC-LI (SNEs U23G; U24C) from constitutive PLlacO promoter | ColE1, AmpR | This study | 5B/C |
| pPL-RydC-LII | pKF223 | RydC-LII | Expresses Salmonella RydC-LII (SNEs U46C; U47G) from constitutive PLlacO promoter | ColE1, AmpR | This study | 5B/C |
| pPL-RydC-LI/LII | pKF225 | RydC-LI/LII | expresses Salmonella RydC-LI/LII (SNEs U23G; U24C and U46C; U47G) from constitutive PLlacO promoter | ColE1, AmpR | This study | 5B/C |
| cfa::gfp | pKF31-1 | cfa::gfp | expresses cfa::gfp translational fusion (−210 rel. to AUG + 15 codons of cfa) from constitutive PLtetO-1 promoter | pSC101*, CmR | (Fröhlich et al., 2013) | 5C |
| pKD4 | Template plasmid for KmR mutant construction | oriRγ, AmpR | (Datsenko and Wanner, 2000) | |||
| pKD46 | γ-β-exo | temperature-sensitive plasmid to express λRED-recombinase from arabinose-inducible ParaB promoter | oriR101, AmpR | (Datsenko and Wanner, 2000) | ||
| pCP20 | FLP–ci857 | Temperature-sensitive Flp recombinase expression plasmid | pSC101, AmpR, CamR | (Cherepanov and Wackernagel, 1995) | ||
| Hfq wild-type | pEH–10–(hfq) | Hfq | Expresses wild-type Hfq protein without tags | AmpR | Kind gift of I. Moll (Max F. Perutz Laboratories, University of Vienna, Austria) | 2, 3, 4, 6 |
| HfqRim | pBAD-Rim | Hfq R19A,P21A,Q33A,Q35A,T49A,R66A | Expresses mutant of Hfq protein without tags | pMB1, AmpR | This study | 6 |
| Hfq66 | pBAD-66 | Hfq amino aids 1-66 | Expresses truncated Hfq protein without tags | pMB1, AmpR | This study | S5 |
Strains
| Trivial name | Stock | Genotype; relevant markers | Source/reference | Used in Figure |
|---|---|---|---|---|
| Salmonella | ||||
| JVS-0007 | SL1344; StrR hisG rpsL xyl | Laboratory stock | ||
| ΔrydC | JVS-0291 | ΔrydC::KanR | (Papenfort et al., 2008) | 1B/C; 5B/C; 7 |
| Δhfq | JVS-0584 | Δhfq | (Sittka et al., 2007) | 1B/C |
| ΔrydC Δhfq | JVS-10665 | ΔrydC::KanR Δhfq | (Fröhlich et al., 2013) | 7 |
| JVS-10701 | hfq70::STOP | This study | ||
| ΔrydC hfq70 | JVS-11150 | hfq70::STOP ΔrydC::KanR | This study | 7 |
| E. coli | ||||
| TOP10 | JVS-2000 | F− mcrA Δ(mrr-hsdRMS-mcrBC) Φ80lacZΔM15 ΔlacX74 recA1 araD139 Δ(ara-leu)7697 galU galK rpsL endA1 nupG λ− | Invitrogen | |