De novo synthesis of a sunscreen compound in vertebrates
Figures
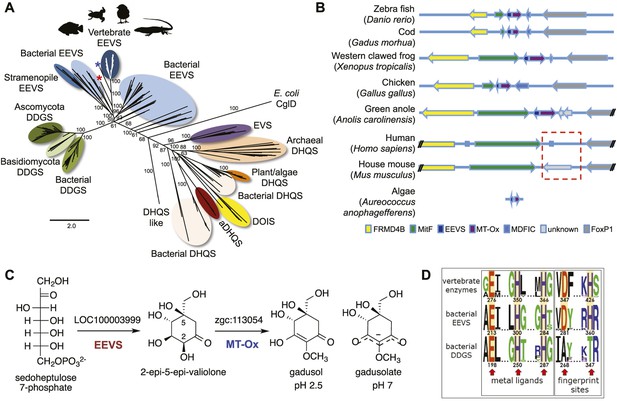
Bioinformatic analysis of sugar phosphate cyclases in prokaryotes and eukaryotes and biochemical characterization of gadusol biosynthetic enzymes.
(A) Bayesian phylogenetic tree of SPCs. Numbers represent posterior probability. The stramenopile Aureococcus anophagefferens, denoted by the blue star, has EEVS and MT-Ox proteins strikingly similar (over 50% identical) to those in vertebrates. The micro algae Coccomyxa subellipsoidea, denoted by the red star, also has EEVS and MT-Ox. (B) Genetic organizations of EEVS and MT-Ox genes in fish, amphibians, birds, and reptiles. Humans and other mammals lack these genes (indicated in dashed red box). For a complete list of vertebrates whose genomes are known to contain EEVS and MT-Ox genes, see Table 1. FRMD4B, FERM domain-containing protein 4B; MitF, microphtalmia-associated transcription factor; MDFIC, MyoD-family inhibitor domain-containing protein-like; and FoxP1, Forkhead-related transcription factor 1. (C) Biochemical characterization of recombinant LOC100003999 and zgc:113054 proteins. (D) WebLogo (Crooks et al., 2004) images of residue conservation patterns at the three metal ligand positions and two active site fingerprint sites known (Kean et al., 2014) to distinguish bacterial EEVSs from DDGSs. The residue numbers given correspond to the reference proteins D. rerio EEVS, ValA, and A. variabilis DDGS, respectively. WebLogos were based on 126 vertebrate, 63 bacterial EEVS, and 160 bacterial DDGS sequences, respectively, that had BLASTP E-values <10−120 in searches using the reference proteins noted above as queries. Each group was aligned using ProMals (Pei and Grishin, 2007).
-
Figure 1—source data 1
Amino acid sequences of the Sugar Phosphate Cyclases (SPCs).
- https://doi.org/10.7554/eLife.05919.004
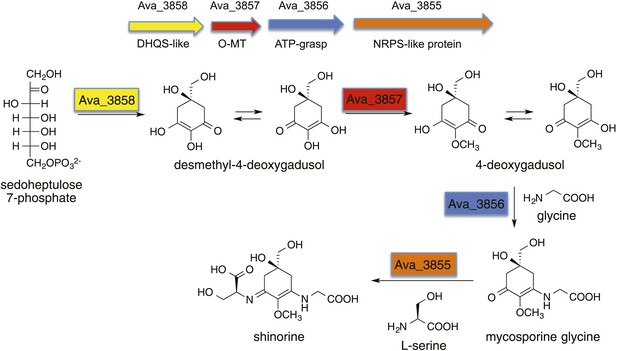
Biosynthetic pathway to shinorine in Anabaena variabilis (Balskus and Walsh, 2010).
https://doi.org/10.7554/eLife.05919.005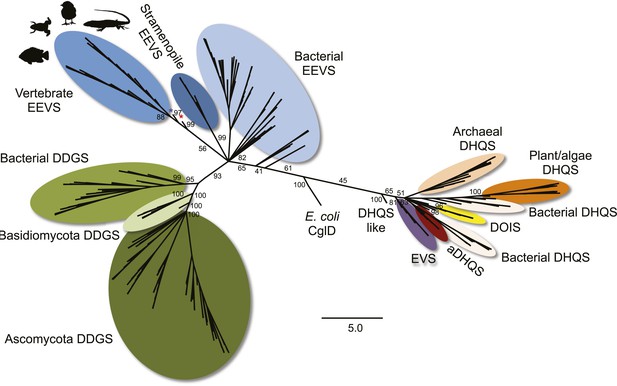
Maximum likelihood phylogenetic tree of sugar phosphate cyclases.
Numbers represent bootstrap confidence values. Blue star denotes the stramenopile A. anophagefferens, and red star denotes the micro algae C. subellipsoidea.
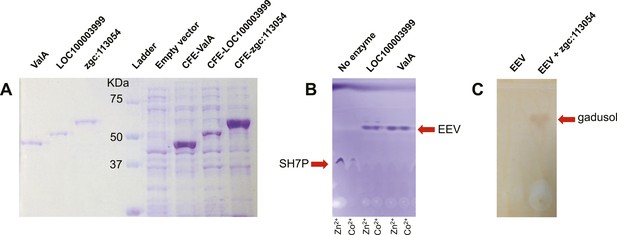
SDS PAGE of EEVS and MT-Ox proteins and thin-layer chromatography (TLC) analysis of their products.
(A) SDS PAGE of purified recombinant ValA, LOC100003999, and zgc:113054, and E. coli cell free extracts containing the enzymes. (B) TLC analysis of enzymatic products of purified ValA and purified LOC100003999. (C) TLC analysis of EEV and the zgc:113054 product (gadusol) using FeCl3 solution as a spraying agent. Only gadusol, but not EEV, can be detected as a maroon spot.
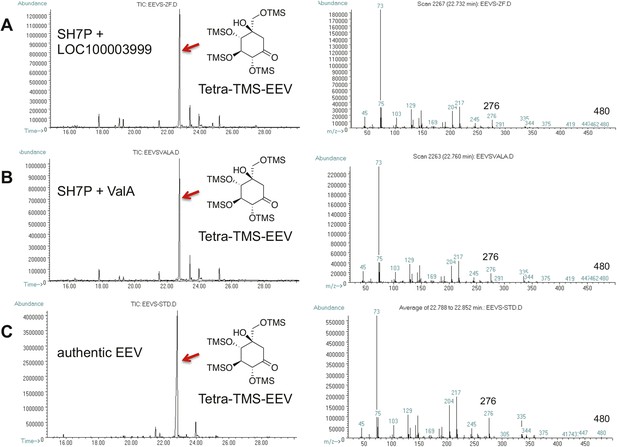
GC-MS analysis of LOC100003999 and ValA reaction products.
(A) SH7P with E. coli cell free extracts containing LOC100003999. (B) SH7P with purified ValA. (C) Authentic EEV. Samples were converted to their trimethylsilyl derivatives using Tri-Sil HTP (Thermo Scientific).
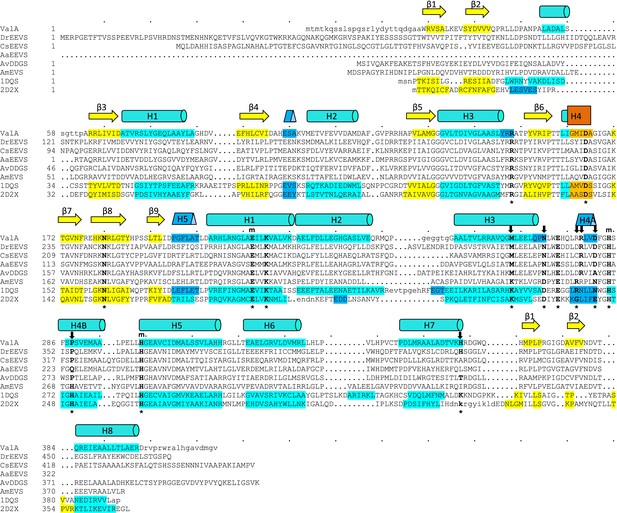
Sequences of the vertebrate clade of EEVS-related proteins have characteristic features of an EEVS.
Alignment of the sequence of the only structurally known EEVS–ValA from Streptomyces hygroscopicus (ShEEVS; PDB entry 4P53) with representative related enzymes. The family consensus secondary structure elements shown schematically shown above the sequences and within each sequence the residues involved in secondary structures are color coded: β-strands (yellow), α-helices (teal), 310 helices (blue), and π helices (orange). ValA is shown first, and other sequences in order are DrEEVS (Danio rerio EEVS), CsEEVS (C. subellipsoidea EEVS), AaEEVS (A. anophagefferens EEVS), AvDDGS (A. variabilis DDGS; Ava_3858), AmEVS (Actinosynnema mirum EVS; Amir_2000), PDB entry 1DQS (Aspergillus nidulans DHQS), and PDB entry 2D2X (Bacillus circulans DOIS). The 14 ‘fingerprint’ active site positions identified by Kean et al. (Kean et al., 2014) are indicated by an asterisk (*) below the sequences. Above the sequences are indicated a subset of those residues that ligate the catalytic metal (m) and another subset with notable variation between the family members that have different types of activity (↓). The DrEEVS sequence and those of all known vertebrate homologs (not shown) match ShEEVS at all fourteen active site positions. The putative algal EEVS enzymes, AaEEVS and CsEEVS, sequences match the EEVS residues at the positions thought to be important for distinguishing the EEVS activity, but respectively differ at one and two positions; these positions are the ones that are of unknown importance for EEVS activity.
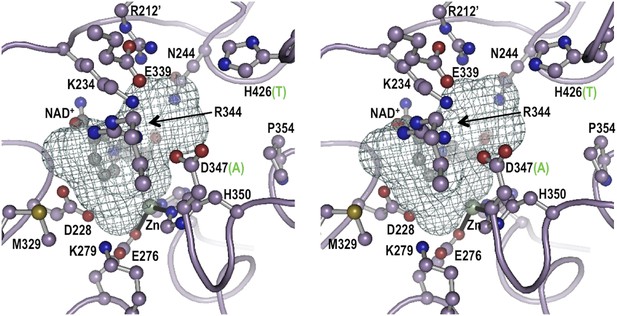
Stereoview for the modeled active site geometry of LOC100003999.
Shown are the 13 active-site residues conserved in all EEVS enzymes (pale purple carbons), the NAD+ (gray carbons), and the Zn2+ atom (silver sphere with black coordination bonds), along with a mesh that delineates the pocket suitable for binding an SH7P substrate. Residue numbers in LOC100003999 identifying twelve of the active site residues are shown; the third metal ligand, H366, is behind the zinc and is not labeled. For the two positions that differ between EEVS and DDGS, the residue type present in DDGS is shown in green in parentheses. Figure made with Pymol (Schrödinger L. The PyMOL Molecular Graphics System, Version 1.3.) using the coordinates of ValA, an EEVS from S. hygroscopicus subsp. jinggangensis 5008 (Kean et al., 2014).
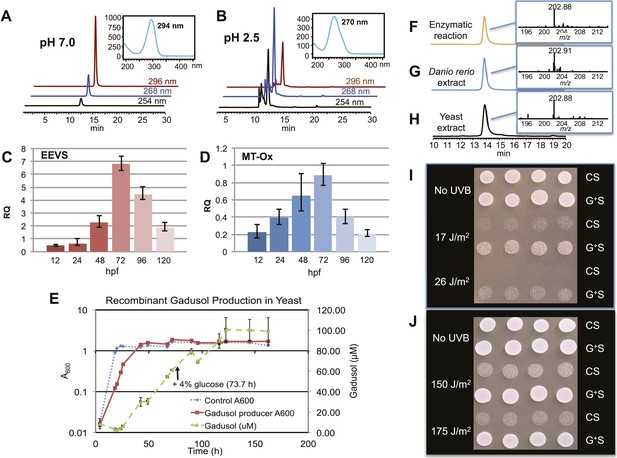
Production of gadusol in zebrafish and yeast and its sunscreen activity.
(A–B) HPLC traces and UV absorptions of gadusol produced from Escherichia coli cell-free extract containing EEVS and purified MT-Ox protein at pH 7.0 and 2.5. (C–D) Transcription patterns of EEVS and MT-Ox genes during zebrafish embryonic development. qRT-PCR analysis of mRNA isolated from zebrafish embryos (n = 3) at 12, 24, 48, 72, 96, and 120 hpf. (E) Time course of gadusol production in yeast harboring the zebrafish genes. The yeast was cultured in YNB + 2% glucose supplemented with leucine and lysine at 30°C for 2 days, and growth was monitored as A600 values (control, dotted blue line; gadusol producer, solid red line). Gadusol concentration in the supernatant of 20 ml cultures (n = 3) was monitored as A296 values in 50 mM phosphate buffer, pH 7.0 (dashed green line) corrected for non-gadusol background absorbance in the control supernatant, normalized to A600 value. Gadusol was quantified based on an extinction coefficient of 21,800 M−1 cm−1 in 50 mM phosphate buffer, pH 7. (F–H) Comparative HPLC analysis of gadusol from recombinant enzymatic reaction, zebrafish extract, and yeast extract. (I) Gadusol suppresses the UVB sensitivity of a rad1∆ yeast mutant; and (J) Gadusol increases the UVB tolerance of a wild-type (RAD1) strain. Cells suspended in control supernatant (CS) or gadusol+ supernatant (G+S) were irradiated with UVB and subsequently spotted in 3 µl aliquots (n = 4) onto YEPD plates, which were incubated at 30°C for 24 hr.
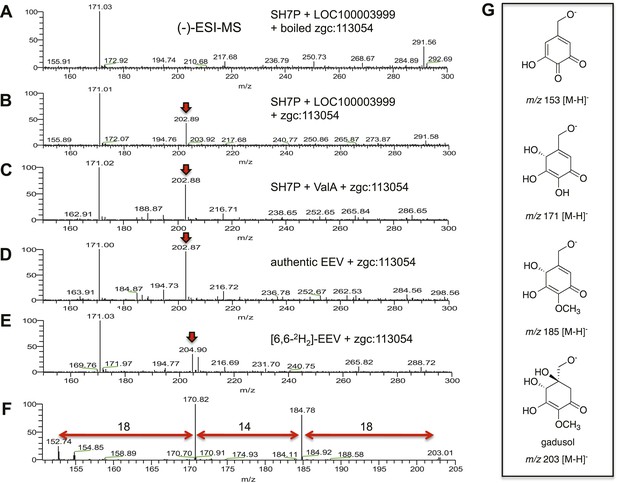
(−)-ESI-MS analysis of LOC100003999 and zgc:113054 reaction products.
(A) SH7P incubated with E. coli cell free extracts containing LOC100003999 (zebrafish EEVS) and boiled zgc:113054 (MT-Ox). (B) SH7P incubated with E. coli cell free extracts containing EEVS and purified MT-Ox. (C) SH7P incubated with purified ValA (a bacterial EEVS) and boiled MT-Ox. (D) Synthetic EEV incubated with purified MT-Ox. (E) Deuterated EEV incubated with purified MT-Ox. (F) MS/MS spectrum of gadusol obtained from the EEVS and MT-Ox reaction. (G) Proposed ion species of gadusol observed in F.
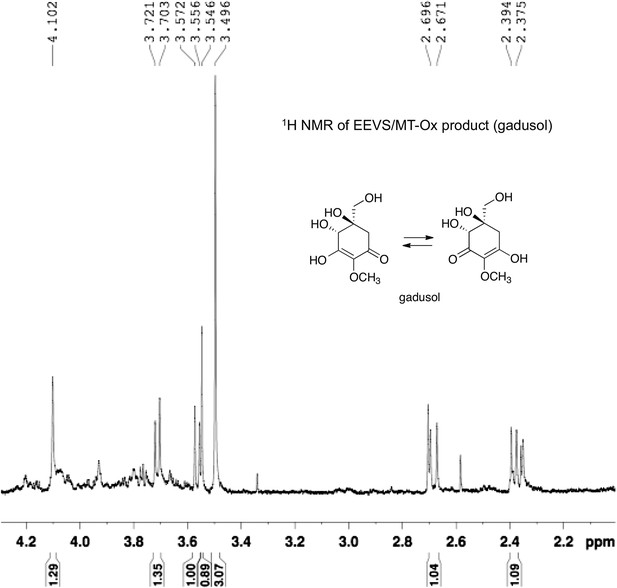
1H NMR spectrum of gadusol obtained from E. coli cell free extracts containing LOC100003999 and zgc:113054 reactions.
https://doi.org/10.7554/eLife.05919.014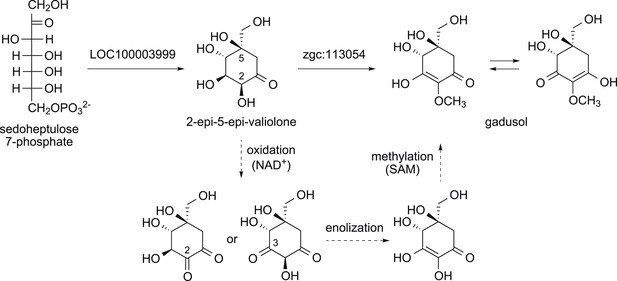
Proposed mode of formation of gadusol in vertebrates.
The dotted arrows show the three steps thought to be a part of the MT-Ox catalyzed reaction.
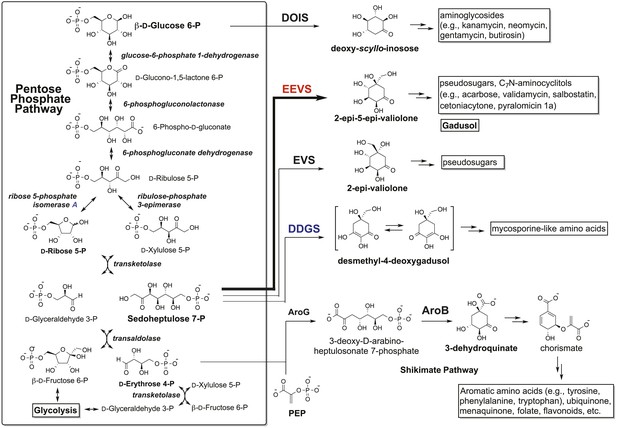
The Pentose Phosphate Pathway (Asamizu et al., 2012) and the shunt pathway to gadusol.
https://doi.org/10.7554/eLife.05919.016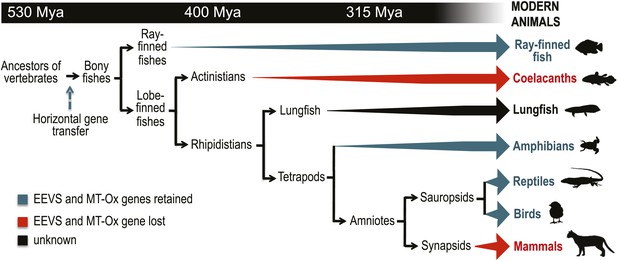
Model timeline for the evolution of the EEVS and MT-Ox genes in vertebrates.
The genes entered an early vertebrate genome as a linked pair (vertical blue arrow) and were retained in the modern ray-finned fish, amphibians, reptiles, and birds as indicated by thick dark cyan arrows. Coelacanths and mammals lost the genes (thick red arrows). No full genome sequence is available for assessing the presence of EEVS and MT-Ox in lungfish. The phylogenetic trees of the EEVS and MT-Ox proteins or mRNA from a selected set of vertebrates can be found in Figure 3—figure supplements 1–3 and Supplementary files 1, 2.
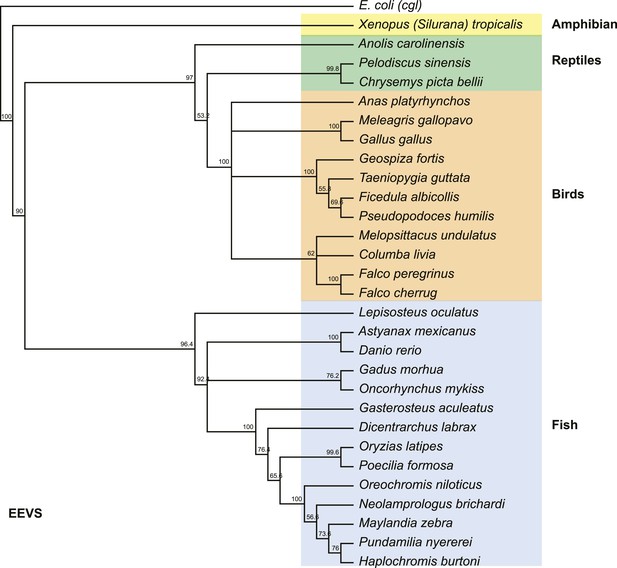
Phylogenetic tree of the EEVS proteins from a selected set of vertebrates.
The distantly related E. coli protein CglD was used as an out-group. Numbers represent bootstrap confidence values.
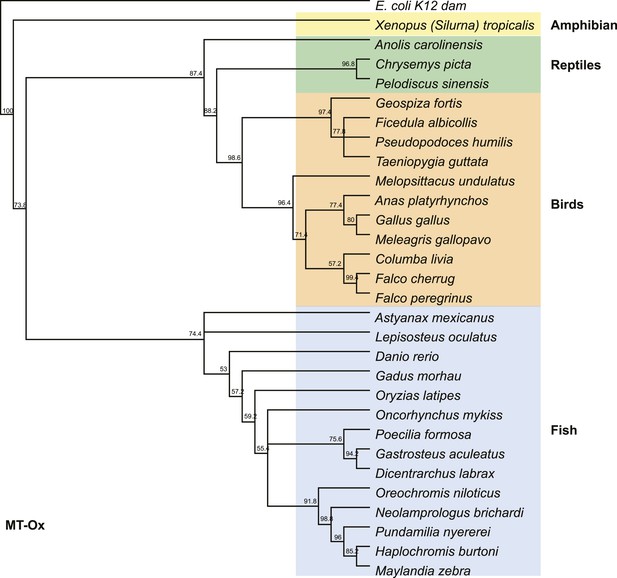
Phylogenetic tree of the MT-Ox proteins from a selected set of vertebrates.
E. coli K12 dam was used as an out-group. Numbers represent bootstrap confidence values.
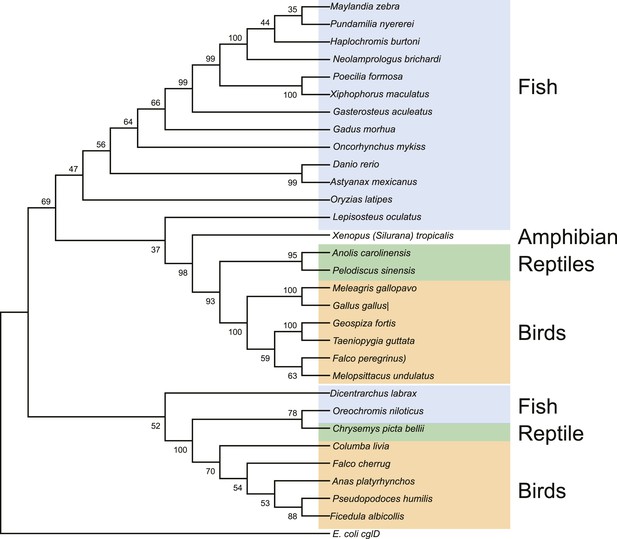
Maximum likelihood tree of vertebrate EEVS mRNA sequences.
E. coli cglD was used as an out-group. Numbers represent bootstrap confidence values.
Tables
Genetic analysis of EEVS and MT-Ox genes in some invertebrates and chordates
| Class | Species | Common name | FRMD4B | MitF | EEVS | MT-Ox | MDFIC | FoxP1 |
|---|---|---|---|---|---|---|---|---|
| Cnidaria | Acropora digitifera | Coral* | – | – | – | – | – | – |
| Nematostella vectensis | Starlet sea anemone* | √ | √ | – | – | – | √ | |
| Hydra vulgaris | Common hydra | – | √ | – | – | – | √ | |
| Invertebrate chordates | Ciona intestinalis | Vase tunicate | √ | √ | – | – | – | √ |
| Branchiostoma floridae | Florida lancelet | √ | √ | – | – | – | √ | |
| Agnatha | Petromyzon marinus | Lamprey† | ‡ | √ | ‡ | ‡ | ‡ | √ |
| Chondrichthyes | Callorhinchus milii | Australian ghostshark† | √ | √ | ‡ | ‡ | √ | √ |
| Osteichthyes (Actinopterygii) | Astyanax mexicanus | Mexican tetra | √ | √ | √ | √ | √ | √ |
| Danio rerio | Zebra fish | √ | √ | √ | √ | √ | √ | |
| Dicentrarchus labrax | European seabass | √ | √ | √ | √ | √ | √ | |
| Fundulus heteroclitus | Mummichog | √ | √ | √ | √ | √ | √ | |
| Gadus morhua | Atlantic cod | √ | √ | √ | √ | √ | √ | |
| Gasterosteus aculeatus | Three-spined stickleback | √ | √ | √ | √ | √ | √ | |
| Lepisosteus oculatus | Spotted gar | √ | √ | √ | √ | √ | √ | |
| Maylandia zebra | Zebra mbuna | √ | √ | √ | √ | √ | √ | |
| Neolamprologus brichardi | African cichlid | √ | √ | √ | √ | √ | √ | |
| Oncorhynchus mykiss | Rainbow trout | √ | √ | √ | √ | √ | √ | |
| Orechromis niloticus | Nile tilapia | √ | √ | √ | √ | √ | √ | |
| Oryzias latipes | Japanese rice fish | √ | √ | √ | √ | √ | √ | |
| Poecilia formosa | Amazon molly | √ | √ | √ | √ | √ | √ | |
| Pundamilia nyererei | African cichlid | √ | √ | √ | √ | √ | √ | |
| Salmo salar | Atlantic salmon | √ | √ | √ | √ | √ | √ | |
| Takifugu rubripes | Japanese puffer | √ | √ | – | – | – | √ | |
| Tetraodon nigroviridis | Green spotted puffer | √ | √ | – | – | – | √ | |
| Xiphophorus maculatus | Southern platyfish | √ | √ | √ | √ | √ | √ | |
| Sarcopterygii | Latimeria chalumnae | West African Coelacanth | √ | √ | – | – | √ | √ |
| Amphibia | Xenopus tropicalis | Western clawed frog | √ | √ | √ | √ | √ | √ |
| Reptilia | Alligator mississippiensis | American alligator | √ | √ | √ | √ | √ | √ |
| Anolis carolinensis | Green anole | √ | √ | √ | √ | √ | √ | |
| Chelonia mydas | Green sea turtle | √ | √ | √ | √ | √ | √ | |
| Chrysemys picta bellii | Western painted turtle | √ | √ | √ | √ | √ | √ | |
| Ophiophagus hannah | King cobra† | √ | √ | ‡ | ‡ | √ | √ | |
| Pelodiscus sinensis | Chinese softshell turtle | √ | √ | √ | √ | √ | √ | |
| Python bivittatus | Burmese python† | √ | √ | ‡ | ‡ | √ | √ | |
| Aves | Anas platyrhynchos | Mallard | √ | √ | √ | √ | √ | √ |
| Columba livia | Rock dove | √ | √ | √ | √ | √ | √ | |
| Falco cherrug | Saker falcon | √ | √ | √ | √ | √ | √ | |
| Ficedula albicollis | Collard flycatcher | √ | √ | √ | √ | √ | √ | |
| Gallus gallus | Chicken | √ | √ | √ | √ | √ | √ | |
| Geospiza fortis | Medium ground-finch | √ | √ | √ | √ | √ | √ | |
| Meleagris gallopavo | North American wild turkey | √ | √ | √ | √ | √ | √ | |
| Melopsittacus undulates | Common pet parakeet | √ | √ | √ | √ | √ | √ | |
| Pseudopodoces humilis | Tibetan ground-tit | √ | √ | √ | √ | √ | √ | |
| Taeniopygia guttata | Zebra finch | √ | √ | √ | √ | ‡ | √ | |
| Zonotrichia albicollis | White-throated sparrow | √ | √ | √ | √ | √ | √ | |
| Mammalia | Balaenoptera acutorostrata scammoni | Minke whale | √ | √ | – | – | – | √ |
| Bos taurus | Cattle | √ | √ | – | – | – | √ | |
| Callithrix jacchus | White-tufted-ear marmoset | √ | √ | – | – | – | √ | |
| Capra hircus | Goat | √ | √ | – | – | – | √ | |
| Ceratotherium simum simum | Southern white rhinoceros | √ | √ | – | – | – | √ | |
| Chinchilla lanigera | Long-tailed chinchilla | √ | √ | – | – | √ | √ | |
| Chlorocebus sabaeus | Green monkey | √ | √ | – | – | √ | √ | |
| Cricetulus griseus | Chinese hamster | √ | √ | – | – | √ | √ | |
| Eptesicus fuscus | Big brown bat | √ | √ | – | – | – | √ | |
| Equus caballus | Horse | √ | √ | – | – | – | √ | |
| Erinaceus europaeus | Western European hedgehog | √ | √ | – | – | √ | √ | |
| Felis catus | Cat | √ | √ | – | – | – | √ | |
| Gorilla gorilla | Western gorilla | √ | √ | – | – | – | √ | |
| Heterocephalus glaber | Naked mole-rat | √ | √ | – | – | √ | √ | |
| Homo sapiens | Human | √ | √ | – | – | – | √ | |
| Lipotes vexillifer | Yangtze river dolphin | √ | √ | – | – | – | √ | |
| Loxodonta africana | African savanna elephant | √ | √ | – | – | – | √ | |
| Macaca fascicularis | Crab-eating macaque | √ | √ | – | – | √ | √ | |
| Mesocricetus auratus | Golden hamster | √ | √ | – | – | √ | √ | |
| Monodelphis domestica | Gray short-tailed opossum | √ | √ | – | – | √ | √ | |
| Mus musculus | House mouse | √ | √ | – | – | √ | √ | |
| Mustela putorius furo | Domestic Ferret | √ | √ | – | – | – | √ | |
| Myotis davidii | Mouse-eared bat | √ | √ | – | – | – | √ | |
| Myotis lucifugus | Little-brown bat | √ | √ | – | – | √ | √ | |
| Nomascus leucogenys | Northern white-cheeked gibbon | √ | √ | – | – | – | √ | |
| Odobenus rosmarus divergens | Pacific walrus | √ | √ | – | – | √ | √ | |
| Orcinus orca | Killer whale | √ | √ | – | – | – | √ | |
| Ornithorhynchus anatinus | Duck-billed platypus† | √ | √ | – | – | – | √ | |
| Orycteropus afer | Ardvark | √ | √ | – | – | – | √ | |
| Otolemur garnettii | Small-eared galago | √ | √ | – | – | – | √ | |
| Ovis aries | Sheep | √ | √ | – | – | – | √ | |
| Pan troglodytes | Chimpanzee | √ | √ | – | – | – | √ | |
| Papio anubis | Olive baboon | √ | √ | – | – | – | √ | |
| Peromyscus maniculatus bairdii | Prairie deer mouse | √ | √ | – | – | √ | √ | |
| Pteropus alecto | Black flying fox | √ | √ | – | – | – | √ | |
| Rattus norvegicus | Norway rat | √ | √ | – | – | – | √ | |
| Sarcophilus harrisii | Tasmanian devil | √ | √ | – | – | √ | √ | |
| Trichechus manatus latirostris | Florida manatee | √ | √ | – | – | – | √ | |
| Vicugna pacos | Alpaca | √ | √ | – | – | – | √ |
-
*
Harbors genes for MAA biosynthesis.
-
†
Incomplete genome sequence; no EEVS and MT-Ox genes identified.
-
‡
The presence of EEVS or MT-Ox genes is unknown due to missing contigs or sequence information.
Additional files
-
Supplementary file 1
SPCs used for multiple sequence alignment and phylogenetic tree construction.
- https://doi.org/10.7554/eLife.05919.021
-
Supplementary file 2
MT-Ox proteins used for multiple sequence alignment and phylogenetic tree construction.
- https://doi.org/10.7554/eLife.05919.022
-
Supplementary file 3
Reciprocal best hit analysis.
- https://doi.org/10.7554/eLife.05919.023
-
Supplementary file 4
Yeast strains used.
- https://doi.org/10.7554/eLife.05919.024
-
Supplementary file 5
Plasmids used.
- https://doi.org/10.7554/eLife.05919.025
-
Supplementary file 6
Primers used.
- https://doi.org/10.7554/eLife.05919.026





