QIL1 is a novel mitochondrial protein required for MICOS complex stability and cristae morphology
Figures
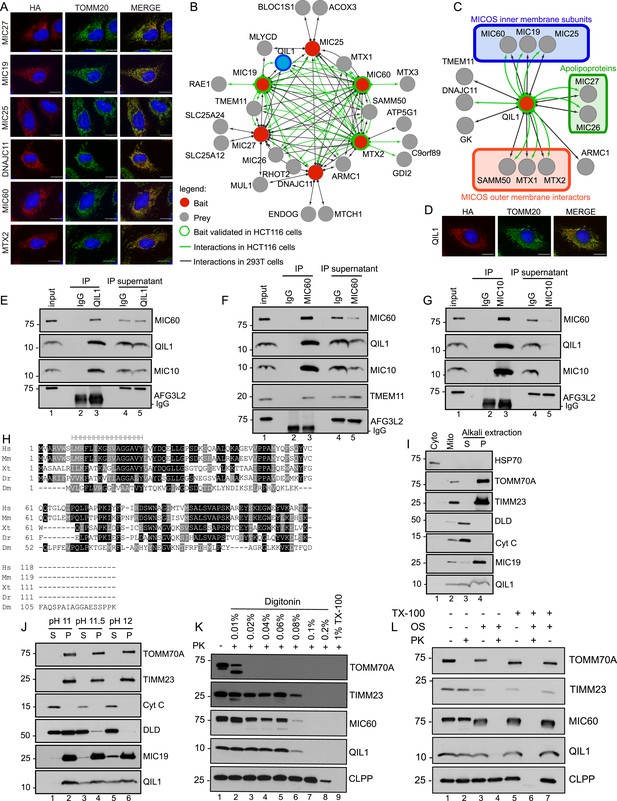
Interaction proteomics of the MICOS complex reveals QIL1 as a novel interactor.
(A) Immunofluorescence analysis of the subcellular localization of tagged proteins. Bars, 20 µm. (B) Overview of the MICOS interaction network obtained from IP-MS analysis. (C) Validation of QIL1 protein interactions by IP-MS analysis. Figure 1—figure supplement 1 shows a schematic representation of the MICOS complex highlighting the subunits and interactors analyzed by IP-MS in red and the expression levels of C-terminally tagged proteins compared to the endogenous version. Figure 1—figure supplement 2 contains the entire IP-MS data set. (D) Immunofluorescence analysis of the subcellular localization of C-terminally tagged QIL1. Bars, 20 µm. Endogenous QIL1 (E), MIC60 (F), or MIC10 (G) was immunopurified from crude mitochondria isolated from 293T cells. Endogenous interactions with MIC60, MIC10, AFG3L2, TMEM11, or QIL1 were assessed. (H) Alignment of QIL1 orthologs using the ClustalW software. ‘H’ indicates amino acids present within a conserved predicted transmembrane region (TMPred algorithm). (I) Cytoplasmic and mitochondrial fractions were separated in 293T lysates. Soluble and membrane fractions were separated by alkaline extraction. S indicates soluble, P indicates pellet. (J) Alkaline extraction was performed at increasing pH (pH11, pH11.5, pH12). (K) Proteinase K (PK) and increasing concentrations of digitonin were used to assess QIL1 sub-mitochondrial localization. (L) Proteinase K (PK), osmotic shock (OS), and Triton X-100 (TX100) were used to assess QIL1 sub-mitochondrial localization.
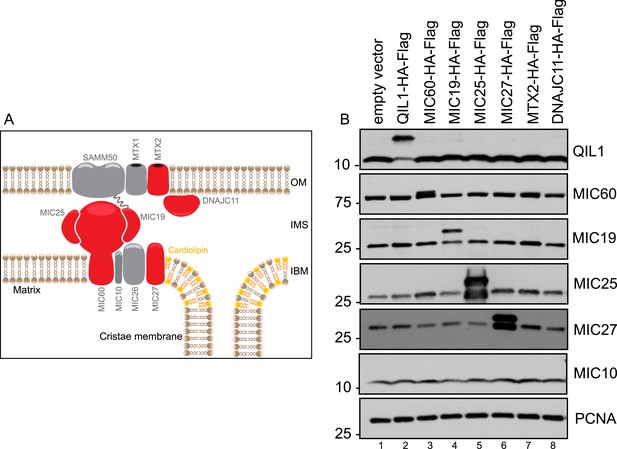
Expression levels of C-terminally tagged MICOS subunits and related proteins analyzed by IP-MS in this study compared to the endogenous version.
(A) Schematic representation of the MICOS complex and its interactors. Proteins C-terminally tagged and subjected to IP-MS analysis are represented in red. (B) Immunoblot analysis was performed to examine the protein levels of the overexpressed C-terminally tagged proteins compared to the endogenous levels in each stable cell line.
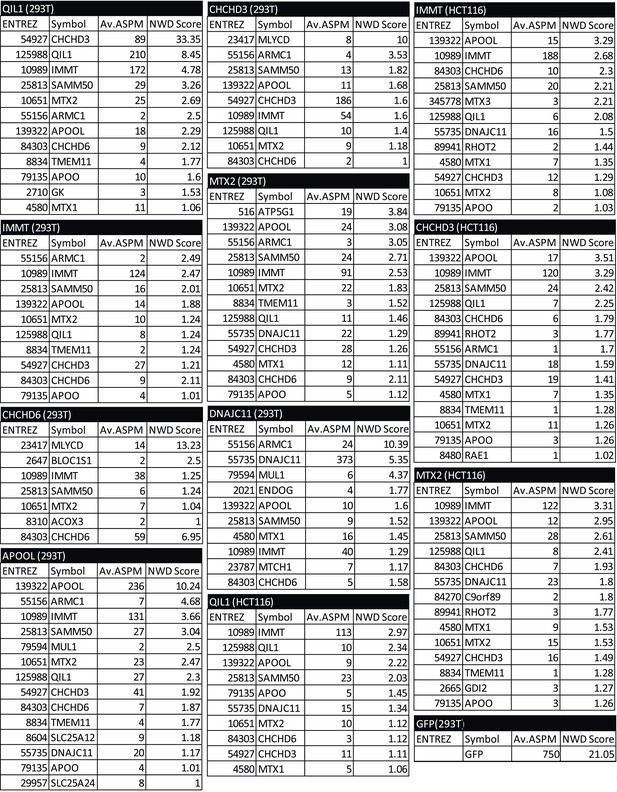
Mass spectrometry analysis of immunopurified protein interactors for QIL1, MIC27, MIC19, MIC25, MIC60, MTX2, DNAJC11 and GFP in 293T and/or HCT116 cells.
Spectra search with Sequest, target-decoy peptide filtering, and linear discriminant analysis (Huttlin et al., 2010) was performed on raw data obtained from technical duplicate runs on either an Thermo LTQ or a LTQ Orbitrap Elite mass spectrometer. Protein Assembler was used to convert spectral counts to APSMs. Peptide data (APSMs) were uploaded into the CompPASS algorithm housed within the CORE environment. Interaction confidence was ranked according to the NWD scores (Sowa et al., 2009). The results obtained from the CompPASS analysis were filtered to determine proteins with a NWD score>1 or high confidence interactors (HCIPs). HCIPs were then filtered for mitochondrial proteins based on the proteins present in Mitocarta (Pagliarini et al., 2008). For the 293T data set, HCIPs with 2 or more ASPMs were included.
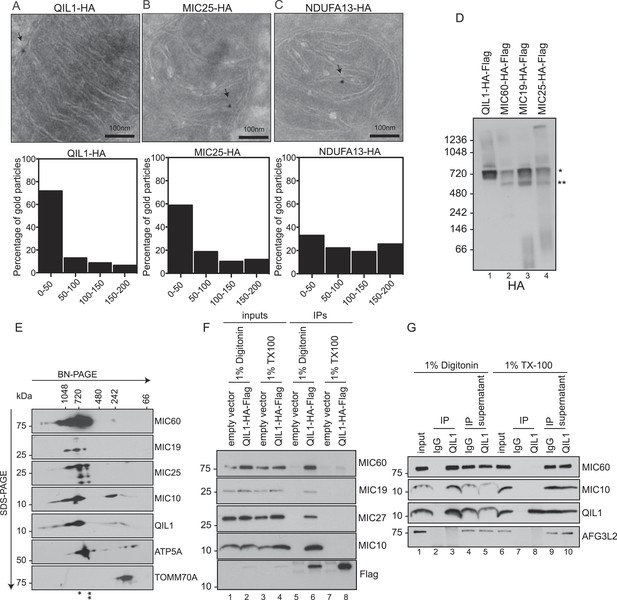
QIL1 localizes at cristae junctions and is present in the mature MICOS complex.
(A–C) Immunogold labeling of 293T cells overexpressing C-terminally tagged QIL1 (A), MIC25 (B), or NDUFA13 (C) using an α-HA antibody coupled to 10-nm gold particles. Bars, 100 nm. Representative mitochondria are shown. The arrows point to the position of a gold particle. The distance in nanometers between the gold particles and the nearest CJ was measured using the ImageJ software. The histograms show the fraction of gold particles within the indicated distance to the crista junction in nanometers in QIL1-HA (n = 231 gold particles), MIC25-HA (n = 192 gold particles), and NDUFA13-HA (n = 309 gold particles) expressing cells. (D) Mitochondria isolated from stable 293T cell lines expressing C-terminally tagged QIL1, MIC60, MIC19, or MIC25 were lysed in 1% digitonin, subjected to BN-PAGE followed by immunotransfer to nitrocellulose membranes and probing with α-HA antibody. The mature ∼700 kDa MICOS complex is highlighted with an asterisk. MIC60, MIC19, and MIC25 were also detected in a sub-complex of ∼500 kDa (two asterisks). (E) Two-dimensional blue native electrophoresis of 293T mitochondrial lysates. Endogenous QIL1 is present in the mature ∼700 kDa MICOS complex (asterisk). MIC60, MIC19, and MIC25 were also present in a smaller sub-complex (two asterisks). (F–G) C-terminally tagged (F) or endogenous QIL (G) was immunopurified from mitochondria lysed in 1% digitonin or 1% Triton X-100 (TX100). Immunoblot analysis was performed to detect interaction with other MICOS subunits.
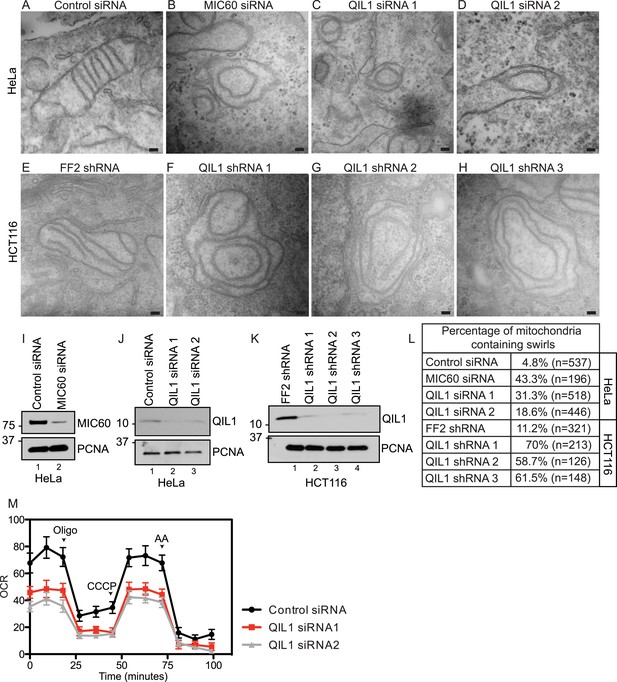
QIL1 deletion alters CJs formation, cristae morphology, and mitochondrial respiration.
Electron microscopy of HeLa cells transfected with Control siRNA (A), MIC60 siRNA (B), two independent QIL1 siRNAs (C and D), FF2 shRNA (E), or QIL1 shRNAs (F–H). Bars, 100 nm. (I–K) Knock-down levels are shown by immunoblot analysis. (L) Quantification of the number of mitochondria containing membrane swirls based on electron microscopy images. (M) Oxygen consumption rate (pmoles/min) was measured at baseline conditions, after oligomycin, FCCP, and antimycin A injections as indicated by arrowheads in HeLa cells transfected with control or QIL1 siRNAs.
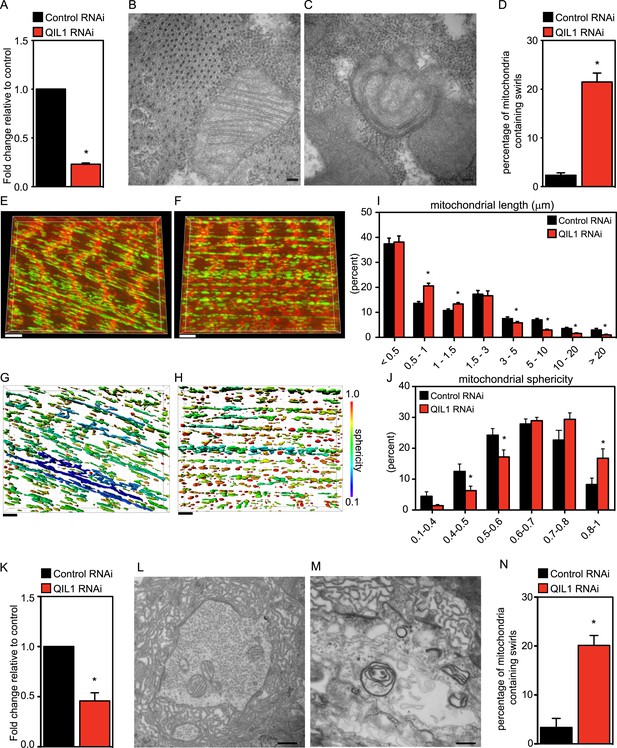
Drosophila QIL1 is required for mitochondrial homeostasis and CJ formation in vivo.
(A) DMef2-driven QIL1 knock-down efficiency in three-instar larvae muscles was assessed by qPCR. Ultrastructural analysis of Control (B) vs QIL1 RNAi (C) third instar larval bodywall muscles in transversal sections. Bars, 100 nm. (D) The number of mitochondria containing IM swirls was quantified. Immunofluorescence analysis of muscles dissected from Control (E) or QIL1 RNAi (F) third instar larvae stained with Phalloidin (red) and α-ATP5A (green). Bars, 10 µm. (I–J) Mitochondrial length (I) and sphericity (G-H, J) of each individual mitochondria were measured. (K) Elav-driven QIL1 knock-down efficiency in three-instar larvae neurons was assessed by qPCR. Ultrastructural analysis of Control (L) vs QIL1 RNAi (M) third instar larval neuromuscular junctions in longitudinal sections. Bars, 500 nm. (N) The number of mitochondria containing IM swirls was quantified.
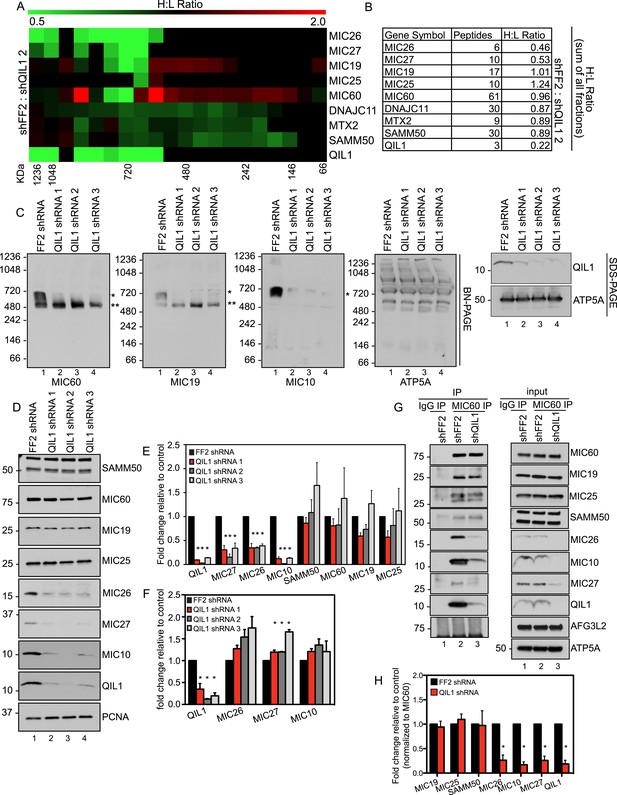
QIL1 deficiency impairs MICOS assembly.
(A) Light-labeled (K0) shFF2 and heavy (K8)-labeled shQIL1 stable cell lines were mixed at a 1:1 ratio and mitochondria subsequently isolated and lysed with 1% Digitonin. Mitochondrial protein complexes were separated using BN-PAGE and sliced in 20 gel pieces ranging from >1 MDa to <60 kDa. Proteins were subjected to tryptic digestion and analyzed by quantitative mass spectrometry. Heavy to light (H:L) ratios were calculated for MICOS components. The ratios were plotted in heatmaps where values <1 are represented in green and values >1 are represented in red. (B) Ratios from the summed heavy and light intensities for each peptide separately across all BN-PAGE fractions from Figure 4A. (C) BN-PAGE followed by immunotransfer to nitrocellulose membranes. QIL1 knockdown lead to a decrease in MIC60, MIC19, and MIC10 in the ∼700 kDa mature MICOS complex (asterisk) and accumulation of MIC60 and MIC19 in a smaller ∼500 kDa sub-complex (two asterisks). (D) Immunoblot analysis of MICOS subunits. (E) Densitometry analysis was performed using ImageJ. (F) qPCR analysis. (G) Endogenous MIC60 was immunopurified from crude mitochondria isolated from HCT116 cells stably expressing FF2 or QIL1 shRNA. Endogenous interactions with MIC19, MIC25, SAMM50, MIC26, MIC10, MIC27, AFG3L2, or QIL1 were assessed. (H) Densitometry analysis was performed using ImageJ.
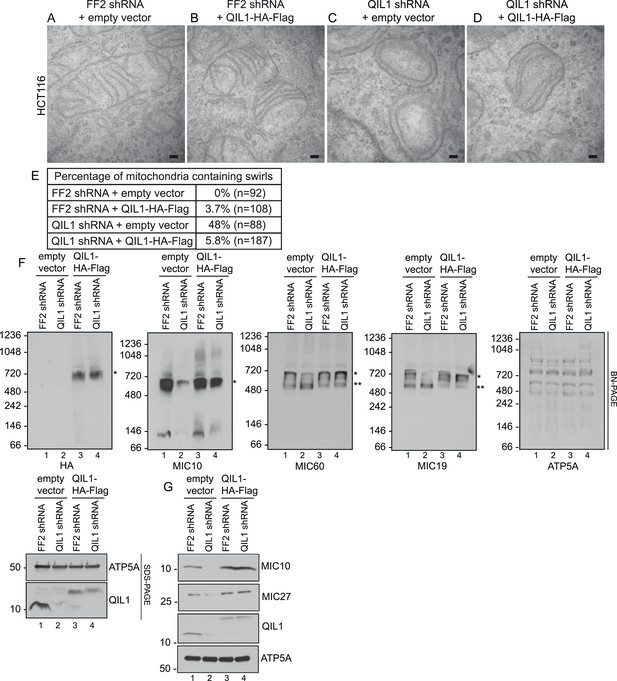
Cristae morphology defects, MICOS disassembly and MIC10 and MIC26 degradation can be rescued by QIL1 overexpression.
Electron microscopy of HCT116 cells transfected with FF2 shRNA and empty vector (A) FF2 shRNA and C-terminally tagged QIL1 (B), a QIL1 shRNA targeting the 3′ untranslated region and empty vector (C) or QIL1 shRNA and C-terminally tagged QIL1 (D). Bars, 100 nm. (E) Quantification of the number of mitochondria containing membrane swirls based on electron microscopy images. (F) BN-PAGE followed by immunotransfer to nitrocellulose membranes. Overexpression of C-terminally tagged QIL1 in cells expressing a QIL1 shRNA targeting the 3′ untranslated region restored the levels of MIC10, MIC19, and MIC60 in the mature ∼700 kDa MICOS complex and reversed the accumulation of MIC19 and MIC60 in the ∼500 kDa assembly intermediate sub-complex (two asterisks). It also restored MIC10 and MIC26 protein levels as observed in immunoblot analysis of total cell lysates (G).
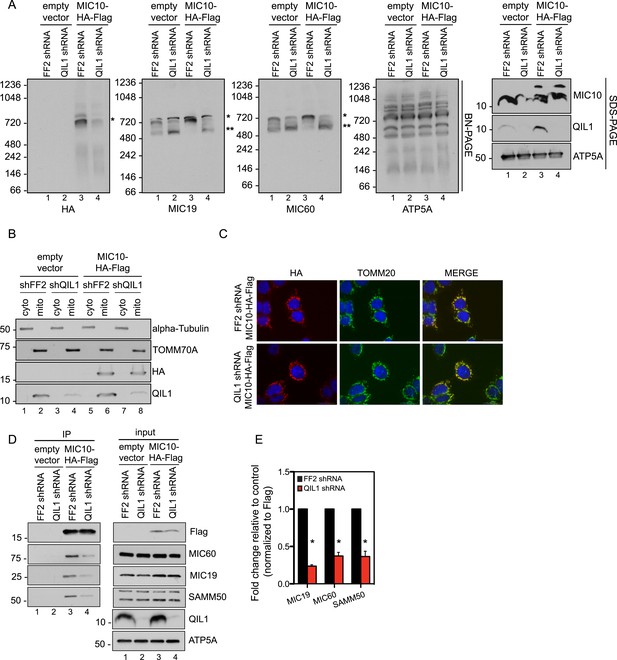
QIL1 is required for the binding of MIC10 to the MICOS complex.
(A) BN-PAGE followed by immunotransfer to nitrocellulose membranes. Incorporation of C-terminally tagged MIC10 into the mature ∼700 kDa MICOS complex (asterisk) was decreased in cells expressing QIL1 shRNA compared to those expressing FF2 shRNA. (B) Cytoplasmic and mitochondrial fractions were separated in lysates obtained from HCT116 cells stably expressing FF2 or QIL1 shRNA and transiently expressing empty vector or C-terminally tagged MIC10. (C) Immunofluorescence analysis of the subcellular localization of C-terminally tagged MIC10. Bars, 20 µm. (D) C-terminally tagged MIC10 was transiently expressed in HCT116 cell lines expressing FF2 shRNA or QIL1 shRNA and immunopurified from mitochondrial lysates. Immunoblot analysis was performed to detect interaction with other MICOS subunits. (E) Densitometry analysis was performed using ImageJ. Figure 7—figure supplement 1 shows the analysis of cardiolipin content and species distribution by LC-MS/MS in mitochondria obtained from cells stably expressing FF2 or 3 different shRNAs targeting QIL1.
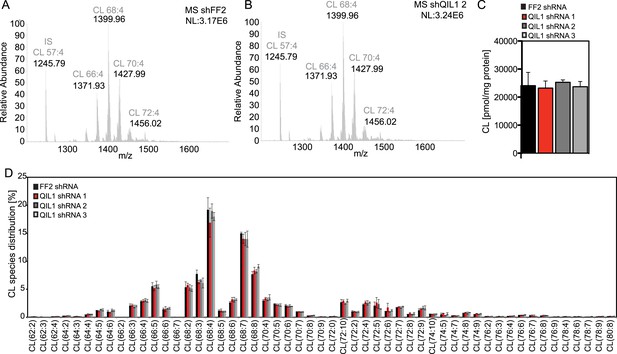
Analysis of cardiolipin content and species distribution by LC-MS/MS in mitochondria obtained from cells stably expressing FF2 shRNA or 3 different shRNAs targeting QIL1.
(A–B) Typical spectra of cardiolipin (CL) obtained from mitochondria isolated from HCT116 cells expressing control (A) or QIL1 shRNA (B). (C) Bar graph showing the quantification of cardiolipin (CL) content in pmol/mg protein by LC-MS/MS. (D) Acyl chain distribution of cardiolipin. First numbers represent the sum of the carbon atoms in the four acyl chains; the second numbers indicate the total number of double bonds.
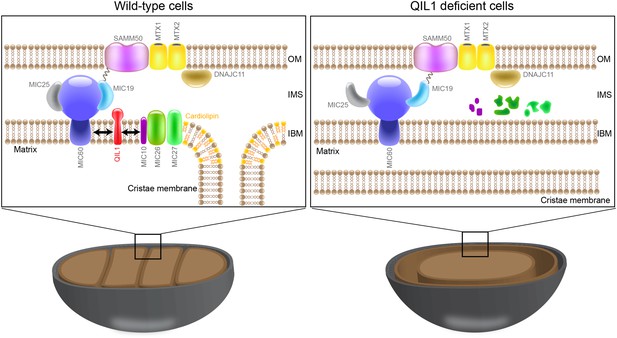
QIL1 is a novel MICOS subunit required for MICOS assembly.
Model for QIL1 incorporation into the MICOS complex and the effect of loss of QIL1 on the composition of MICOS subunits. We propose that MIC10, MIC26, and MIC27 fail to assemble into a stable MICOS sub-complex containing MIC60, MIC19, and MIC25 and are degraded when QIL1 is depleted leading to MICOS disassembly, loss of CJs and cristae morphology defects.





