2.8 Å resolution reconstruction of the Thermoplasma acidophilum 20S proteasome using cryo-electron microscopy
Figures
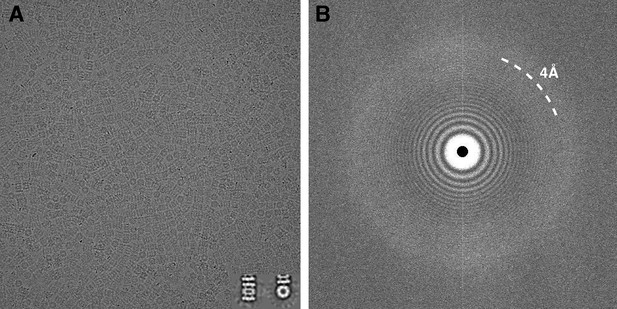
Typical micrograph of ice-embedded T20S proteasome and corresponding power spectrum.
(A) Micrograph of T20S after movie-frame alignment. Inset: Reference free 2D class averages showing a side view (left) and a top view (right). (B) Thon rings are visible well beyond 4 Å−1 resolution in the power spectrum of the micrograph shown in (A).
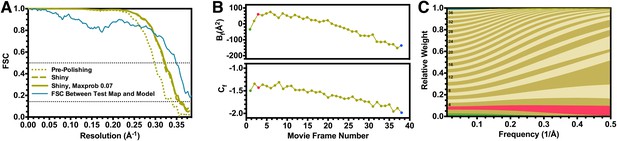
Fourier shell correlation curves and radiation damage weighting plots.
(A) Gold-standard FSC curves for the T20S reconstructions before and after particle polishing as well as after excluding particles based on the uncertainty of the angular assignments. Estimated resolutions are 2.98 Å, 2.83 Å, and 2.81 Å, respectively. The FSC curve computed between the atomic model and the test map is shown in blue. The FSC reaches 0.5 at 2.86 Å resolution, in agreement with the resolution estimated by gold standard refinement in Relion. (B) Estimated values for Bf (top) and Cf (bottom) during the particle polishing procedure. (C) Frequency-dependent relative weights for all movie frames. The first, third, and final movie frames are highlighted in green, red, and blue, respectively.
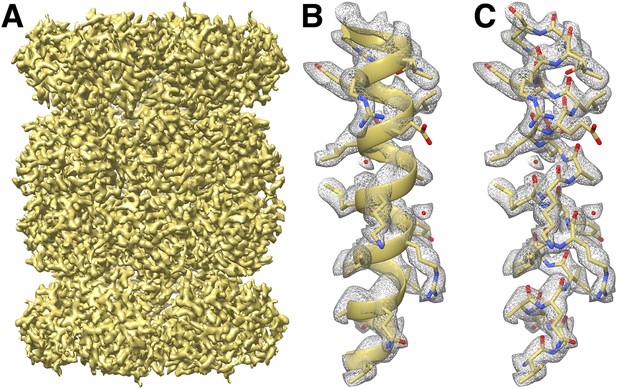
CryoEM reconstruction of the T20S at 2.8 Å resolution.
(A) Isosurface representation of the T20S map. (B) An α-helical segment from one β subunit is shown in ribbon representation docked into the corresponding region of the reconstruction. (C) Same α-helical segment as in (B) shown in atom representation docked into the corresponding region of the reconstruction. Several water molecules are visible.
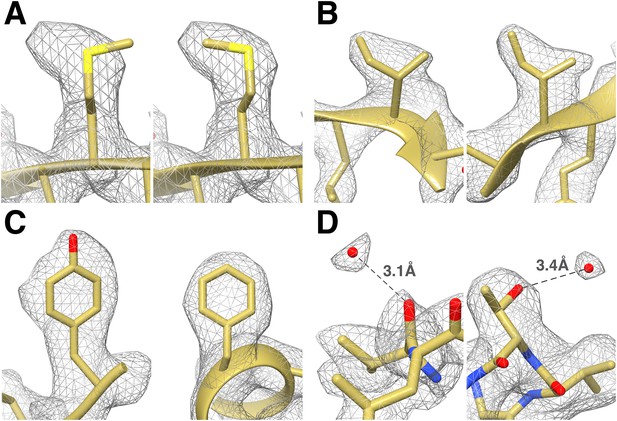
Identification of the rotameric conformation of amino acid side chains and resolving of ordered water molecules in the T20S cryoEM reconstruction at 2.8 Å resolution.
(A) Different rotameric conformations adopted by the Met-14 side chain (β-subunit) between a crystal structure of the T20S determined at 3.4 Å resolution (left, PDB 1PMA) and the EM density (right). (B) Unambiguous establishment of the rotameric conformations of two different isoleucine residues: Ile70 (left, α-subunit) and Ile37 (right, β-subunit). (C) The additional density accounting for the hydroxyl group of tyrosine side chains (left, Tyr132, α-subunit) is prominent when compared to phenylalanine side chains (right, Phe91, α-subunit). (D) Numerous water molecules are resolved in the T20S cryoEM map. Left: a water molecule hydrogen-bonded to the carbonyl group of Val87 (α-subunit). Right: a water molecule hydrogen-bonded to the side chain hydroxyl of Thr102 (α-subunit). The cross-validation of the assignment of these water molecules using gold-standard half maps can be found in Figure 4—figure supplement 1.
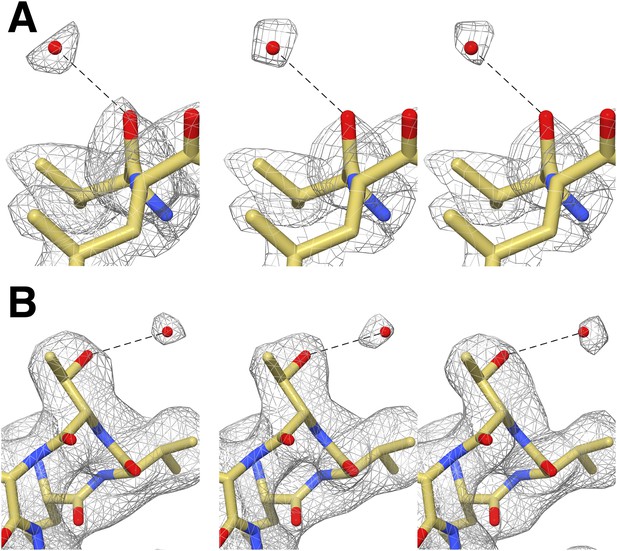
Cross-validation of water molecule assignment through comparison of gold standard half maps.
(A) Water molecule hydrogen-bonded to Val87. Left: The EM density reconstructed using all particles (49,954). Middle and right: gold standard half-maps. (B) Water molecule hydrogen-bonded to Thr102. Left: EM density reconstructed using all particles. Middle and right: gold standard half-maps. In each case, the density attributed to water is present in both half maps confirming that it is not due to random noise. Furthermore the position of these water molecules matches those observed in a crystal structure of the proteasome determined to 1.9 Å resolution (PDB 1YAR).