Structure of a type IV pilus machinery in the open and closed state
Figures
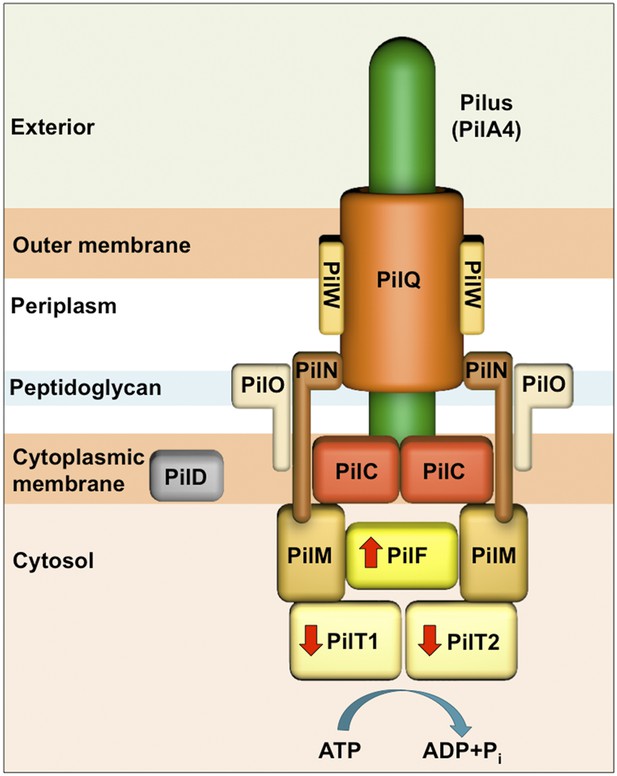
Schematic of the T4P machinery in T. thermophilus.
The type IV pilus machinery is a heterooligomer, formed from at least 10 different proteins. The PilQ secretin (orange) forms a channel in the outer membrane for secretion of the pilus-forming protein PilA4 (green), which is processed by the prepillin peptidase PilD (grey) (Friedrich et al., 2002; Schwarzenlander et al., 2009). The membrane protein PilW (light orange) plays a role in DNA transport, PilQ assembly, and pilus extrusion (Rumszauer et al., 2006; Schwarzenlander et al., 2009). The dimeric complex PilC (red) is located in the inner membrane and is essential for pilus formation (Friedrich et al., 2002; Karuppiah et al., 2010). PilM (light brown), PilN (dark brown), and PilO (beige) are suggested to form the inner membrane assembly platform and connect the periplasmic and cytoplasmic sides of the complex (Rumszauer et al., 2006; Schwarzenlander et al., 2009; Karuppiah and Derrick, 2011; Karuppiah et al., 2013). The cytoplasmic ATPases PilF (bright yellow) and PilT1/PilT2 (pale yellow) drive pilus extension and retraction, respectively (indicated with red arrows) (Rose et al., 2011; Salzer et al., 2014b).
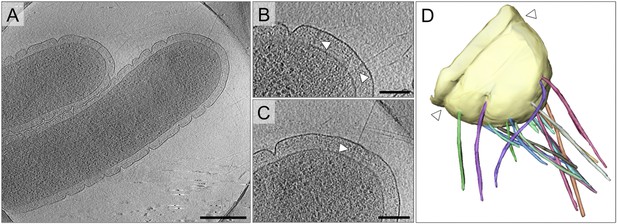
Cell morphology and pili of T. thermophilus.
(A–C) Tomographic slices through T. thermophilus cells show invaginations in the outer membrane and large protein complexes crossing the periplasm (white arrowheads), which are associated with pili. Scale bar = 500 nm in A, 100 nm in B and C. (D) Volume rendering shows the distribution of pili (multi-coloured), protruding from the outer membrane (pale yellow). A concentric invagination of the outer membrane is indicated (white arrowheads).
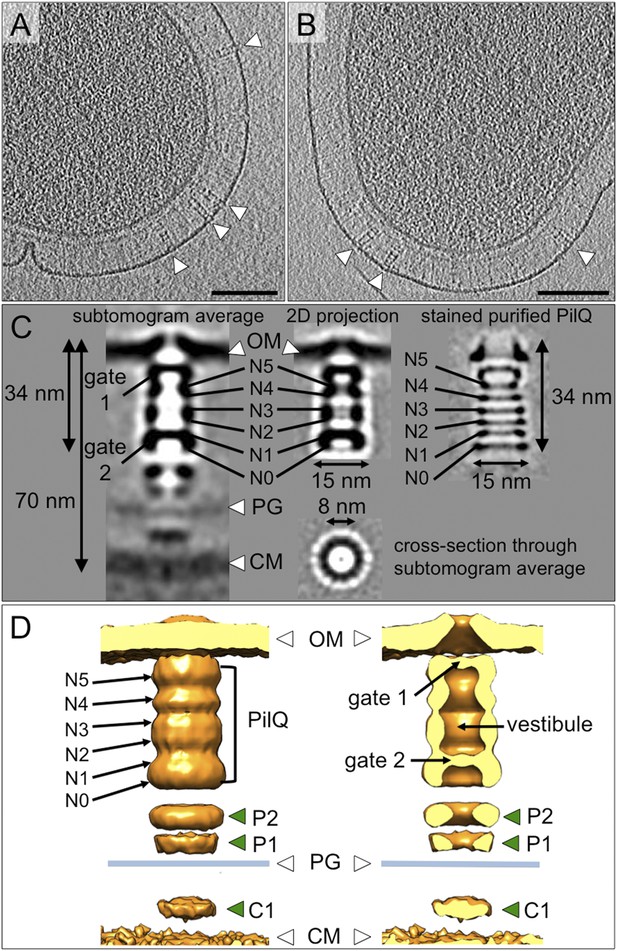
Structure of the T4P machinery in the closed state.
(A and B) Tomographic slices of T. thermophilus cells show large protein complexes crossing the periplasm in the absence of pili (white arrowheads). Scale bars = 100 nm. (C) Resulting subtomogram average (left panel) and its 2D projection (centre) are compared to the previously determined projection map of isolated and stained PilQ (right panel) (Burkhardt et al., 2011). The contrast of the stained PilQ has been inverted. This image was originally published in The Journal of Biological Chemistry. Janin Burkhardt, Janet Vonck, and Beate Averhoff. Structure and Function of PilQ, a Secretin of the DNA Transporter from the Thermophilic Bacterium T. thermophilus HB27. JBC. 2011; 286:9977–9984, the American Society for Biochemistry and Molecular Biology. The putative N0–N5 domains of PilQ (Burkhardt et al., 2012) are marked. (D) 3D surface rendering of the average reveals that PilQ has a periplasmic vestibule closed at both ends by two gates. Additional protein densities distinct from PilQ (green arrowheads; C1 = proximal to the cytoplasmic membrane, P1 = central periplasmic ring 1, P2 = central periplasmic ring 2) are also shown. OM, outer membrane; PG, peptidoglycan; CM, cytoplasmic membrane.
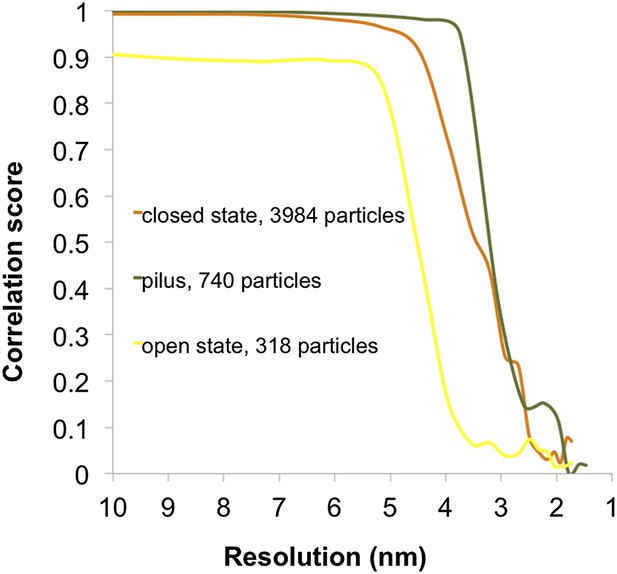
Fourier shell correlation curves for subtomogram averages.
Resolution estimates were based on conventional Fourier shell correlation (FSC) measurements and the 0.5 criterion in IMOD. Calculations for the closed state of the T4P machinery (orange, 35 Å), open state with pilus extended (yellow, 45 Å), and the pilus (green, 32 Å) are shown.
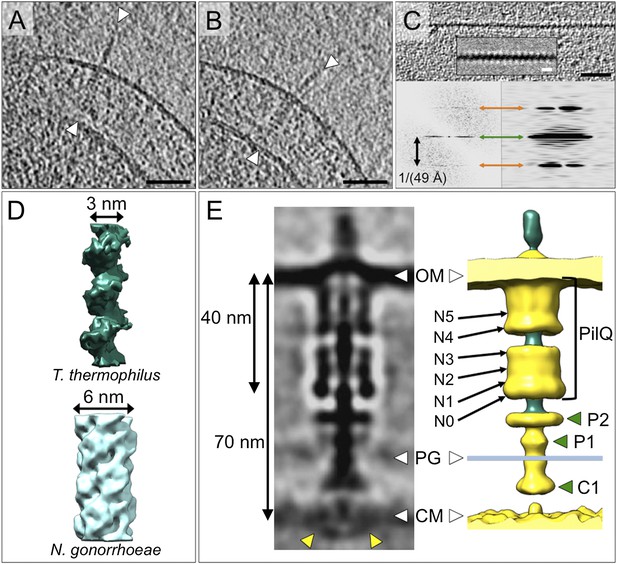
Structure of the T4P and the assembled machinery in the open state.
(A and B) Tomographic slices show close-up views of the T4P machinery with assembled pili (white arrowheads). Scale bar = 50 nm. (C) Upper panel, a tomographic slice of the pilus shows that the structure is periodic. Scale bar = 20 nm. A slice through the subtomogram average (inset) shows the repeat more clearly. Scale bar = 5 nm. Lower panels, power spectra of the tomographic slice (left) and the average (right) depict layer lines (orange arrows) at a distance of 1/(49 Å) from the equator (green arrow), corresponding to a helical pitch of 4.9 nm. The contrast has been inverted. (D) Subtomogram average of the ∼3 nm wide T. thermophilus pilus (green, top panel), compared to the previously determined ∼6 nm wide structure from N. gonorrhoeae (blue, bottom panel, EMDB 1236) (Craig et al., 2006). (E) Subtomogram average (left panel) and 3D surface rendering (right panel) of the T4P machinery in the open state with the central pilus (green). The putative N0–N5 (Burkhardt et al., 2012) domains of PilQ are marked. Additional protein densities distinct from PilQ (green arrowheads; C1 = proximal to the cytoplasmic membrane, P1 = central periplasmic ring 1, P2 = central periplasmic ring 2) are also shown. Compared to the closed state of the complex, additional protein densities (left panel, yellow arrowheads) are visible in the cytoplasm. See Video 1 for details. OM, outer membrane; PG, peptidoglycan; CM, cytoplasmic membrane.
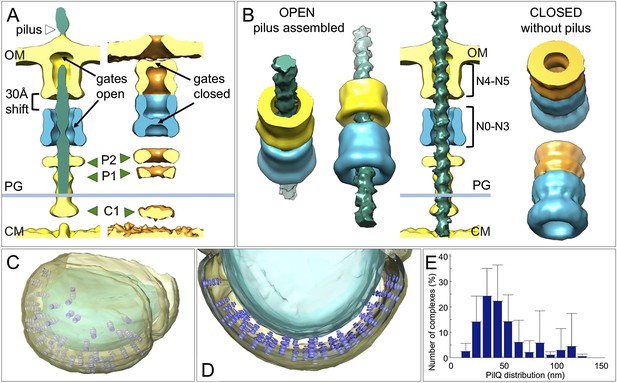
Changes between the open and closed state of the T4P machinery and its distribution in situ.
(A and B) Comparisons between the PilQ components of the T4P machinery reveal large conformational changes whereby both gates open and domains N0–N3 (now shown in blue for both states) shift by ∼30 Å towards the cytoplasm on pilus extrusion. Green arrowheads indicate additional protein densities (C1 = proximal to the cytoplasmic membrane, P1 = central periplasmic ring 1, P2 = central periplasmic ring 2). In (B) the structure of the T4P has been docked into the open state. OM, outer membrane; PG, peptidoglycan; CM, cytoplasmic membrane. (C and D) Docking subtomogram averages (purple) into the tomographic volume of a cell reveals the distribution of the closed T4P machinery in situ with respect to the outer membrane (pale yellow) and cytoplasmic membrane (blue). See Video 2 for details. (E) Averaged histogram of nearest-neighbour distance between protein complexes, calculated from 9 cells, with a total of 332 data points. Error bars indicate the standard deviation of the frequency distribution for each minimal distance.
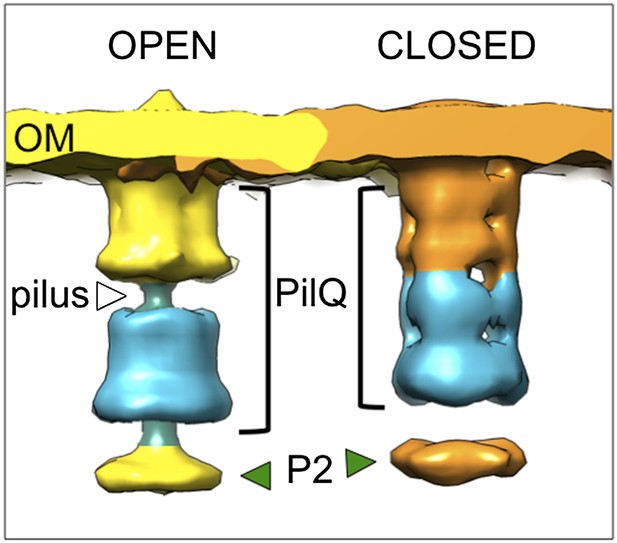
Subtomogram averages from cells with high periplasmic protein content.
Subtomogram averages were calculated for the open and closed state of the T4P machinery from the tomogram shown in Figure 2A–C, where the periplasm contains high-protein levels. The resolution of the subtomogram averages is limited due to the low number of protein complexes available from a single tomogram. Therefore, data for PilQ and protein P2 (green arrowhead) only are shown, which indicate essentially the same large conformational change in domains N0–N3 (blue) as seen in Figure 5A,B.
Videos
Comparison of the T4P machinery in the open and closed state.
The video was generated by morphing the two subtomogram averages in ImageJ.
Distribution of T4P complexes in situ.
The video shows the rendered tomographic volume of a T. thermophilus cell. The outer membrane (pale yellow), peptidoglycan (orange), and cytoplasmic membrane (blue) are shown with the closed T4P complexes (purple).





