Asymmetric division triggers cell-specific gene expression through coupled capture and stabilization of a phosphatase
Figures
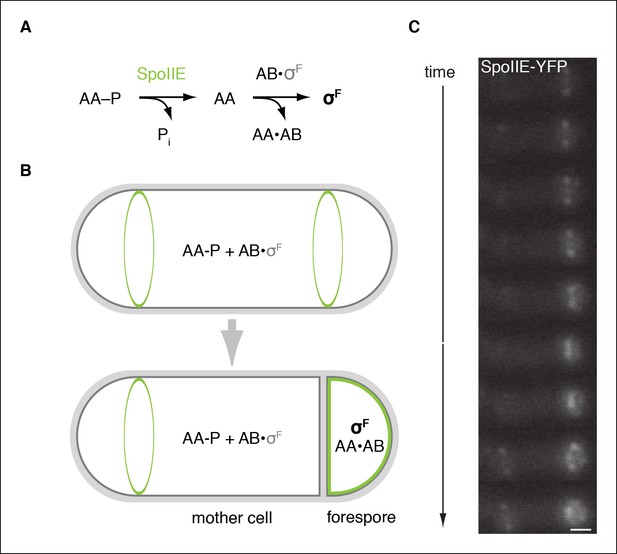
SpoIIE is compartmentalized in the forespore and activates σF.
(A) Diagram of the pathway for σF activation. Phorphorylated SpoIIAA (AA-P) is dephosphorylated by SpoIIE. Dephosphorylated SpoIIAA (AA) then binds to SpoIIAB (AB) displacing σF and leading to σF-directed transcription. (B) SpoIIE (green), AA, AB and σF are produced in predivisional cells. Prior to completion of asymmetric cell division SpoIIE associates with the polar divisome near one or both cell poles (the pole at which division initiates is chosen randomly). Following completion of cytokinesis, SpoIIE is enriched in the forespore where it dephosphorylates AA-P to activate σF. (C) A montage of images taken every 6 min from a single sporulating cell (strain RL5876) producing SpoIIE-YFP. Cells are oriented with the forespore on the right as in the diagram. A movie of this sporulating cell is provided as Video 1. Scale bar: 0.5 µm.
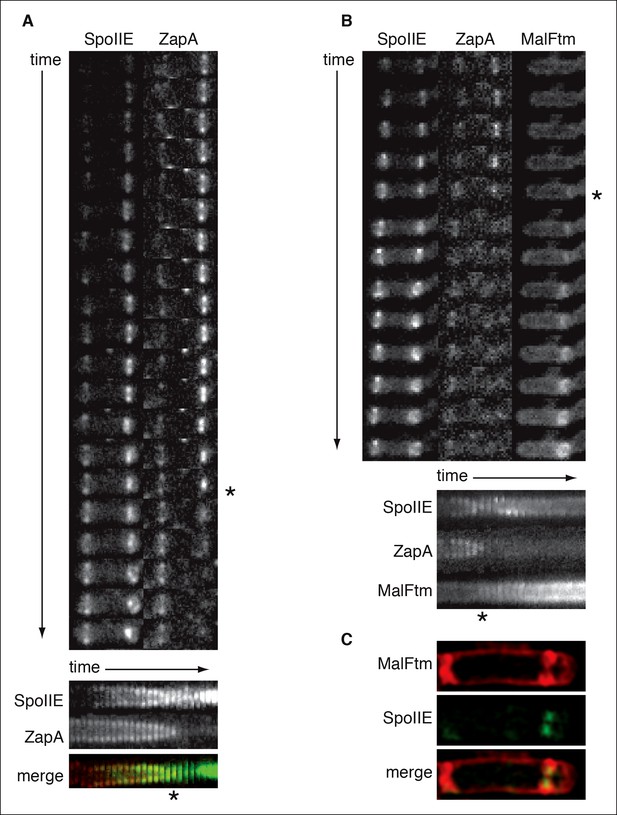
SpoIIE constricts along with FtsZ during asymmetric cell division.
(A) Montage of images of SpoIIE-YFP and CFP-ZapA in a sporulating cell (strain RL6042) from timelapse microscopy. Images were acquired at 5 min intervals on agarose pads following resuspension to induce sporulation. The lower panel is a kymograph of a slice through the asymmetric FtsZ ring from the images above. The merge shows SpoIIE in green and ZapA in red. An asterisk marks the frame with maximal constriction of the FtsZ ring. A movie of this sporulating cell is provided as Video 2 . (B) Montage of images of SpoIIE-YFP, CFP-ZapA, and MalFtm-mNeptune in a sporulating cell (strain RL6043) from timelapse microscopy. Images were acquired at 7 min intervals on agarose pads following resuspension to induce sporulation. The lower panel is a kymograph of a slice through the asymmetric FtsZ ring from the images above. An asterisk marks the frame with maximal constriction of the FtsZ ring. A movie of this sporulating cell is provided as Video 3. (C) Structured illumination microscopy image of a sporulating cell (strain RL6044) expressing SpoIIE-GFP (green) and MalFtm-mNeptune (red). Of 16 such cells observed with incomplete polar septa, 81% showed clear constriction of the SpoIIE ring.
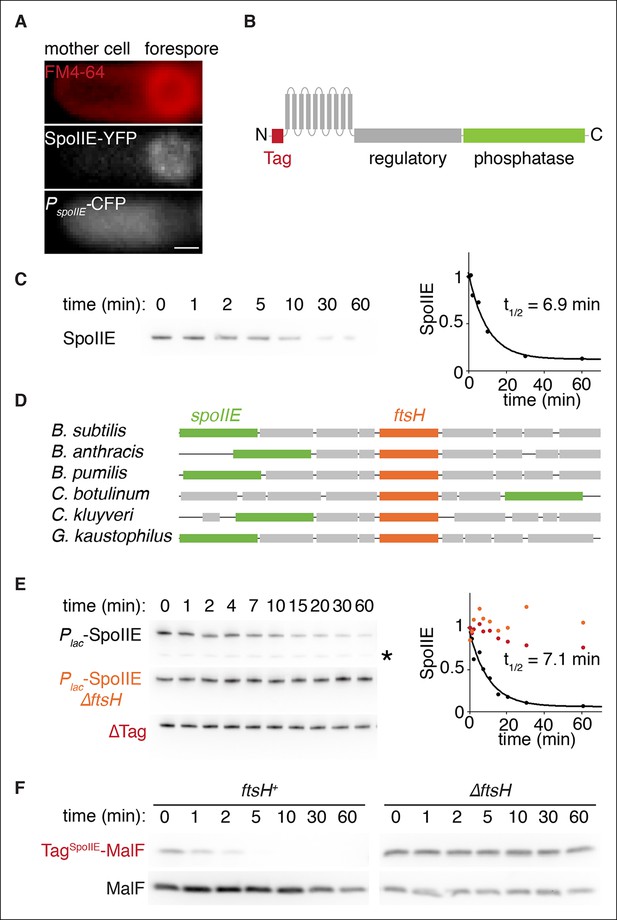
SpoIIE degradation depends on FtsH.
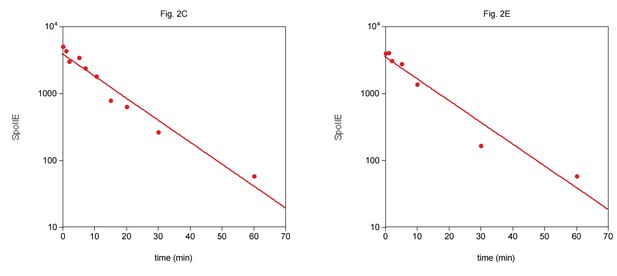
(A) SpoIIE is compartmentalized to the forespore. A single sporulating cell (strain RL5874) is shown following the completion of asymmetric septation. The membrane stained with FM4-64 is shown in red above SpoIIE-YFP and CFP driven by an in frame fusion to the start of the spoIIE open reading frame. Scale bar: 0.5 µm. (B) The domain architecture of SpoIIE. The N-terminal cytoplasmic tail (red), followed by 10 transmembrane-spanning segments, the regulatory region (amino acids 320–589, gray), and the phosphatase domain (amino acids 590–827, green). (C) SpoIIE is degraded during sporulation. Translation was arrested (by addition of 100 µg/ml chloramphenicol) in sporulating cells producing SpoIIE-FLAG (strain RL5877), and samples were withdrawn at the indicated times. SpoIIE was detected by western blot using α-FLAG monoclonal antibody (left). Quantitation of the western (right) fit to a single exponential equation. (D) The genes for spoIIE and ftsH are near each other in the genome with conserved synteny. The diagram shows genomic organization of diverse endospore forming species. Filled boxes indicate genes with spoIIE in green, ftsH in orange, and other genes in gray. (E) SpoIIE degradation requires FtsH, and FtsH mediated degradation requires the N-terminal cytoplasmic tail (TagSpoIIE) of SpoIIE. Translation was arrested in vegetatively growing cells producing SpoIIE-FLAG with an IPTG inducible promoter (strain RL5878 wt, RL5879 ∆ftsH, and RL5880 SpoIIE-∆Tag (residues 11–37 were deleted). Degradation was monitored (left) and quantitated (right) as in panel C. (F) TagSpoIIE is sufficient to target a heterologous protein for FtsH-dependent degradation. Degradation of MalF-TM-FLAG fused at the N-terminus to either TagSpoIIE (wt RL5888, ∆ftsH RL5889) or the first 10 amino acids of SpoIIE (wt RL5890, ∆ftsH RL5891) produced during exponential growth was monitored as in panel C.

SpoIIE degradation depends on FtsH.
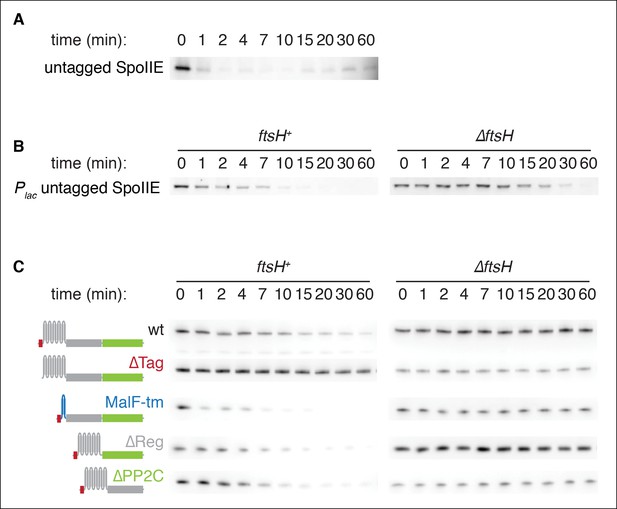
(A) Translation was arrested (by addition of 100 µg/ml chloramphenicol) in sporulating wild-type PY79 cells (strain RL3), and samples were withdrawn at the indicated times. SpoIIE was detected by western blot using affinity purified rat α-SpoIIE polyclonal antibody. (B) Degradation of SpoIIE was monitored in vegetative cells induced to produce untagged SpoIIE (strain RL6040, wt) (strain RL6041, ∆ftsH) as in A. (C) Domains of SpoIIE-FLAG were serially deleted or replaced as diagramed at left and expressed in wt or ∆ftsH vegetatively growing cells. Translation was arrested by chloramphenicol addition, and samples were collected at times indicated. SpoIIE was detected by western blotting with α-FLAG monoclonal antibody. ∆Tag removes residues 11–37 (strain RL5880, wt) (strain RL5884, ∆ftsH), MalF-tm replaces the transmembrane domain (residues 38–319) with the first two transmembrane domains of E. coli MalF (strain RL5881, wt) (strain RL5885, ∆ftsH), ∆Reg removes the regulatory domain (residues 320–568) (strain RL5882, wt) (strain RL5886, ∆ftsH), and ∆PP2C removes the phosphatase domain (residues 590–827) (strain RL5883, wt) (strain RL5887, ∆ftsH). Western blots for strains shown in Figure 2E are duplicated here for comparison.
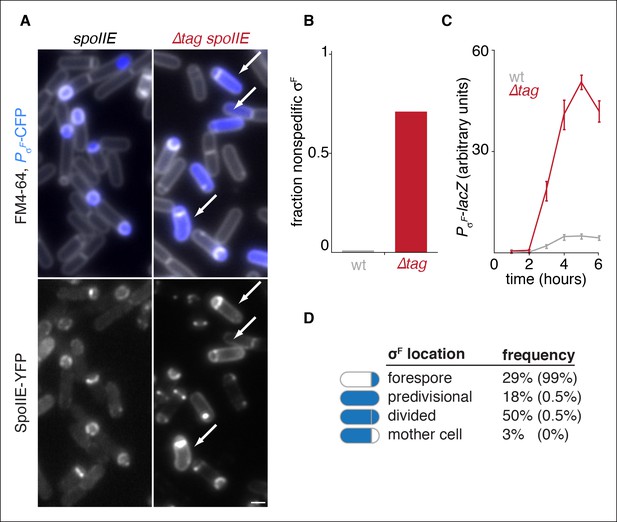
Degradation of SpoIIE is required to compartmentalize SpoIIE and σF activity.
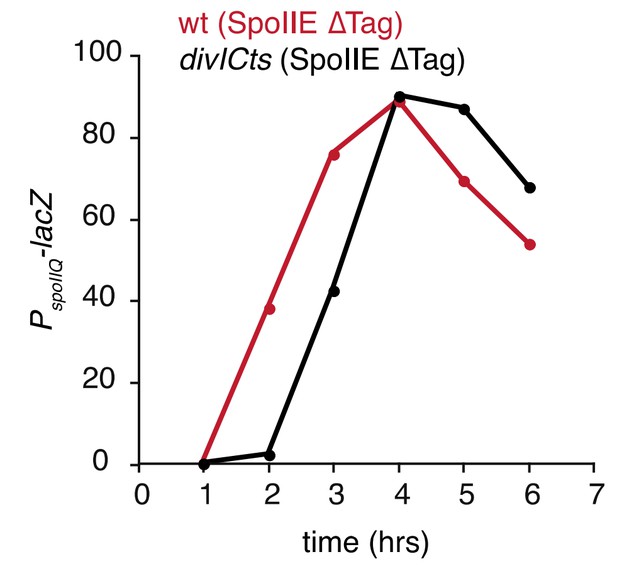
(A) Images of sporulating cells producing SpoIIE-YFP (left, strain RL5876) or ∆Tag SpoIIE-YFP (right, strain RL5892). Top images show CFP (blue) produced under the control of the σF dependent spoIIQ promoter and FM4-64-stained membrane (white); bottom images show SpoIIE-YFP. White arrows indicate cells with uncompartmentalized σF activity and SpoIIE in the mother cell. The contrast for images of SpoIIE-YFP has been adjusted to approximately 5X brighter than for ∆Tag-SpoIIE-YFP for display purposes. Scale bar: 1 µm. (B) Quantification of the forespore specificity of σF activity from hundreds of cells from images as shown in panel A. (C) σF activity was measured during sporulation using a translational fusion of the σF dependent SpoIIQ promoter to LacZ (wt SpoIIE strain RL5893, ∆Tag SpoIIE strain RL5894). Time after initiation of sporulation is indicated, and error bars represent the standard deviation from three biological replicates. (D) Quantification of the dependence of σF activation on asymmetric cell division driven by ∆Tag-SpoIIE. Hundreds of cells from images as shown in panel A were manually assessed for completion of asymmetric cell division and compartmentalization of σF activity. The percent of cells with each pattern of σF activity is indicated for ∆tag-spoIIE cells (values for wt spoIIE cells are indicated in parenthesis).
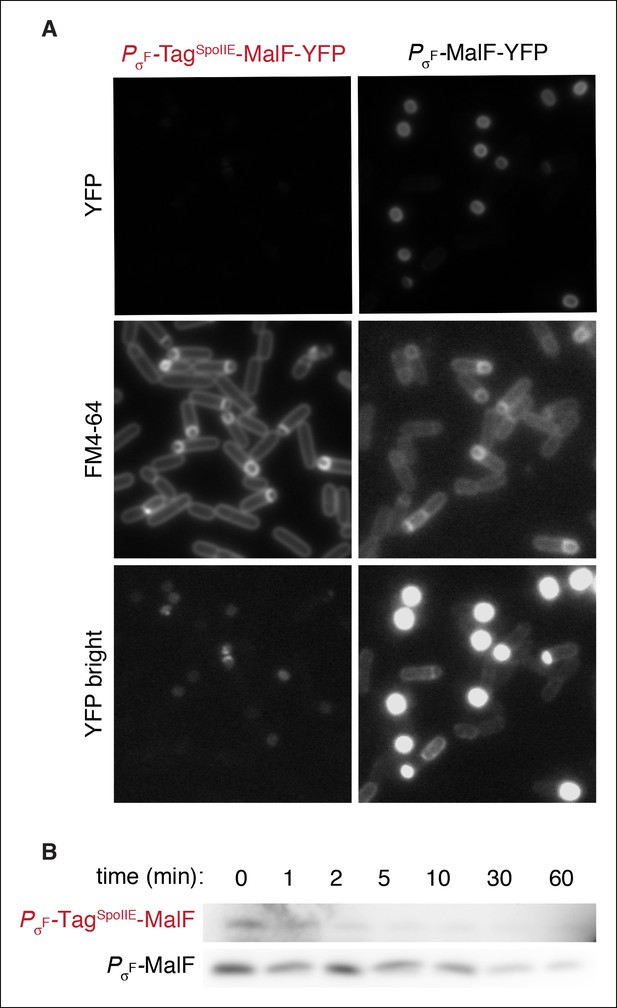
FtsH is active in the forespore.
(A) Images of sporulating cells expressing MalF-TM-YFP (the first two transmembrane segments of E. coli MalF fused to YFP (strain RL5932) or TagSpoIIE-MalF-TM-YFP (strain RL5933) under the control of the σF dependent spoIIQ promoter. Cells were stained with FM4-64 (middle images). Images of PσF-MalF-TM-YFP and PσF-TagSpoIIE-MalF-TM-YFP are shown with identical exposure times and contrast settings; bottom images are adjusted to higher contrast than top images to show limited accumulation of PσF-TagSpoIIE-MalF-TM- YFP in the forespore. (B) Sporulating cells expressing MalF-TM-FLAG (strain RL5934) or TagSpoIIE-MalF-TM-FLAG (strain RL5935) under the control of the σF dependent spoIIQ promoter were translationally arrested, and samples were taken at the indicated timepoints. MalF-TM-FLAG was detected by western blotting with α-FLAG monoclonal antibody.
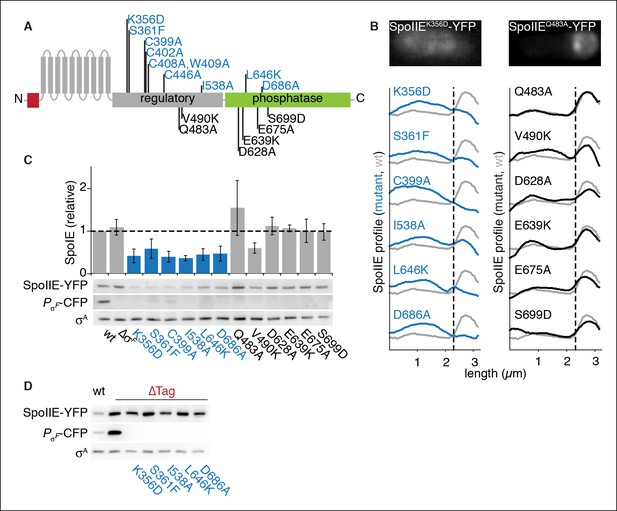
SpoIIE stabilization and localization mutants.
(A) Diagram of SpoIIE mutants with sporulation defects. Variants with localization and σF activation defects are shown above the diagram in blue, and variants with normal localization but defects in σF activation are shown below the diagram in black. (B) Localization of SpoIIE mutants. Hundreds of asymmetrically divided cells were aligned at the forespore pole to generate average profiles of SpoIIE-YFP localization for each SpoIIE mutant (strains RL5895- 5909) with a reference plot (gray) from wild-type SpoIIE-YFP from σF mutant cells (strain RL5910). The dashed line represents the approximate position of the asymmetric septum. Images of representative cells with the mislocalized variant SpoIIEK356D (left, strain RL5895) and forespore-localized SpoIIEQ483A (right, strain RL5904) are shown. (C) Western blots of protein levels in SpoIIE mutant strains shown in panel B probed for SpoIIE-YFP (with α-GFP antibody), CFP produced under the control of a σF-driven promoter, and σA as a loading control. Levels of each SpoIIE variant were normalized to wt SpoIIE (strain RL5876). Error bars represent the standard deviation from three biological replicates. All localization mutants (shown in blue) are different from the σF mutant control with p values less than 0.0025 from a paired t-test. (D) Western blots of SpoIIE compartmentalization mutants with TagSpoIIE removed (strains RL5911-5916, with strain RL5876 as a reference) as in panel C.
-
Figure 4—source data 1
Sporulation efficiency of SpoIIE mutants.
Cultures of indicated strains with were grown in DSM for 28 hr at 37°C followed by heat killing for 20 min at 85°C. The number of spores was determined by counting viable cells.
- https://doi.org/10.7554/eLife.08145.011
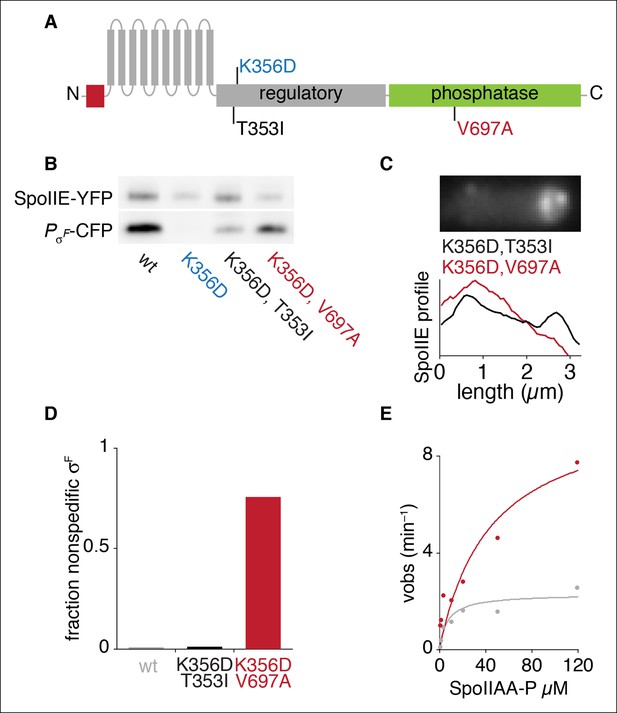
An allele specific suppressor of SpoIIEK356D rescues compartmentalization and stabilization.
(A) Diagram of SpoIIE with the position of the mutation K356D shown above and the suppressor mutations T353I and V697A shown below. (B) SpoIIE variants were detected in extracts of sporulating cells by western blotting using an α-GFP antibody (wt strain RL5876, SpoIIEK356D strain RL5895, SpoIIEK356D, T353I strain RL5936, SpoIEK356D, V697A strain RL5939). σF activity was measured in these samples using a CFP reporter fused to the σF dependent spoIIQ promoter. (C) Localization of SpoIIE mutants. Average profiles of SpoIIE-YFP localization from hundreds of asymmetrically divided cells aligned at the forespore pole are shown for SpoIIEK356D, T353I (strain RL5943), and for SpoIIEK356D, V697A (strain RL5944). An image of a representative cell expressing SpoIIEK356D, T353I (strain RL5943) following asymmetric septation is shown above. (D) Quantification of the forespore specificity of σF activity based on CFP fused to the σF dependent spoIIQ promoter using strains as in C. (E) Multiple turnover measurements of dephosphorylation of SpoIIAA-P. Data were fit to the Michaelis-Menton equation; Vmax = 7.0 min–1 for SpoIIE320-827 and 44.3 min–1 for SpoIIE320-827, V697A.
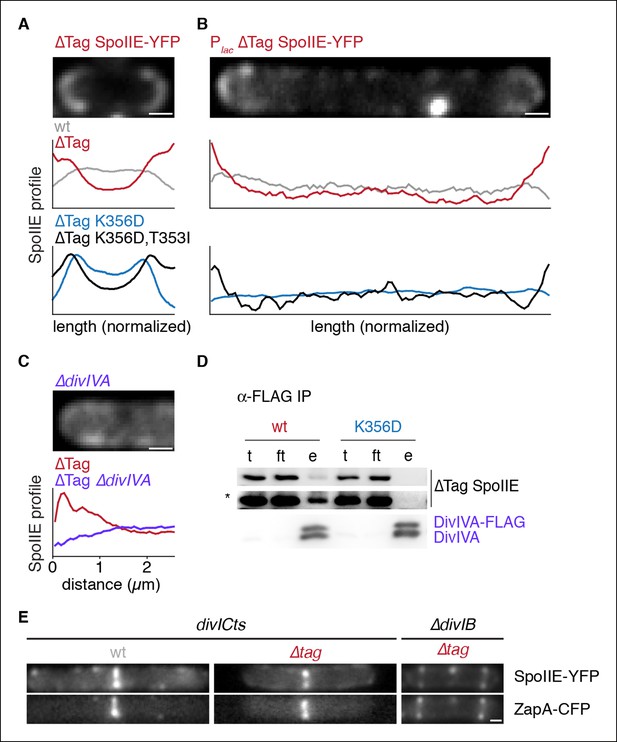
Stabilized SpoIIE localizes to the cell pole.
(A) Average profiles of SpoIIE-YFP from undivided sporulating cells lacking σF activity are shown (strains RL5910, 5917, 5912, 5918), with a representative cell with ∆Tag-SpoIIE-YFP (strain RL5917) displayed above. (B) Vegetatively growing cells expressing SpoIIE-YFP (strains RL5919-5922) displayed as in panel A; variants as indicated) were imaged 30 min after induction of expression of the FtsZ polymerization inhibitor MciZ. (C) MciZ-expressing cells (ΔdivIVA [strain RL5923] or otherwise wildtype [strain RL5924]) were imaged as in panel B and average profiles of ∆Tag-SpoIIE-YFP were generated from 20 randomly selected cell poles. (D) DivIVA-FLAG was immunoprecipitated with α-FLAG magnetic beads from extracts of sporulating cells expressing SpoIIE variants as indicated (strains RL5925, 5926), and detected by Western blot. The elution (e) shown is 100X concentrated relative to the load (l) and flowthrough (ft) samples. Blots were probed with α-GFP antibody (top two images; lower image in high contrast*) and α-DivIVA antisera (below). Because DivIVA oligomerizes, untagged DivIVA is also co-immunoprecipitated. (E) SpoIIE preferentially localizes to the divisome rather than the pole. Representative cells expressing SpoIIE-YFP and CFP-ZapA are shown. Left images show exponentially growing divICts cells (strains RL5927, 5928), and right images show a sporulating ΔdivIB cell (strain RL5929). Scale bars indicate 0.5 µm in all panels.
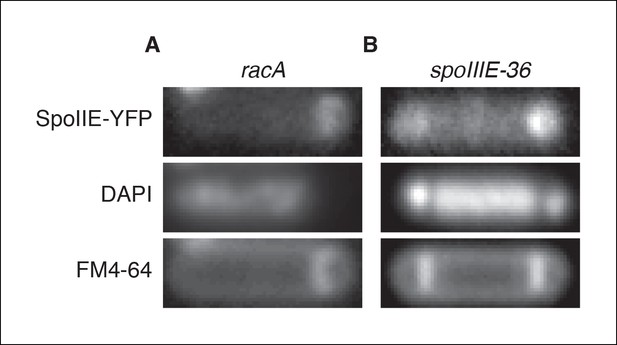
Transcription of spoIIE in the forespore is not required to compartmentalize SpoIIE.
(A) Representative image of SpoIIE-YFP in a sporulating ∆racA mutant cell (strain RL6045) that failed to capture the chromosome in the forespore. The chromosome was stained with DAPI and the membrane was stained with FM4-64. (B) Representative image of a SpoIIE-YFP in a sporulating cell harboring the spoIIIE36 mutation to block pumping of the chromosome to the forespore and with the spoIIE-YFP gene near the terminus (strain RL6046).
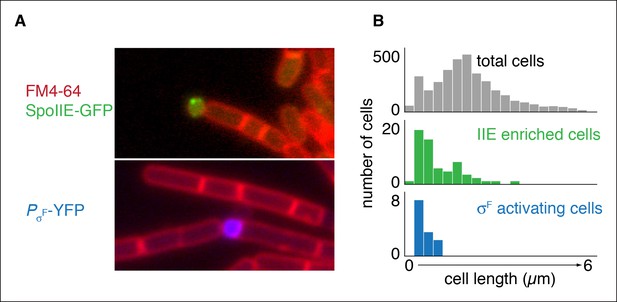
Repositioning the septum in vegetative cells is sufficient for compartmentalization of SpoIIE.
(A) In the top image, vegetatively growing cells producing SpoIIE-GFP and overexpressing the ftsZ operon formed minicells enriched for SpoIIE-GFP (strain RL5930). In the lower image, cells additionally expressed the spoIIA operon and harbored a YFP reporter for σF activity (strain RL5931). (B) Quantification of images as shown in panel A. SpoIIE enriched cells (representing 56 out of 3168 total cells) are shown in green in the middle plot and are defined as cells with 2 standard deviations above the mean SpoIIE-GFP intensity. Cells that had active σF were rare in the population (12 cells) and were identified based on detectable levels of YFP fluorescence and their sizes were measured. They are shown in blue at the bottom.
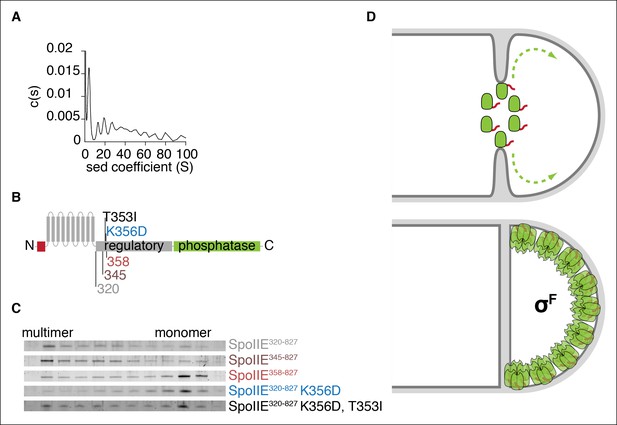
Multimerization is required for compartmentalization of SpoIIE and σF activation.
(A) Purified soluble SpoIIE320-827 was analyzed by sedimentation velocity analytical ultracentrifugation detected by absorbance at 280nm and fitted to c (s) using Sedfit. Peaks corresponding to the predicted sedimentation coefficient for monomeric SpoIIE, hexamers, and multimers of hexamers were observed. (B) Diagram of the mutations and truncations in SpoIIE analyzed in panel C. (C) Multimerization of SpoIIE variants was analyzed by gel filtration with a 24 ml Superose 6 column. 1 ml fractions from 7-–18 ml of the run were collected, run on SDS-PAGE gels, and stained with SYPRO Ruby. (D) Model for handoff of SpoIIE from the divisome to the adjacent cell pole. SpoIIE (green) initially accumulates at the divisome and constricts along with FtsZ during cytokinesis. Prior to the completion of cell division, SpoIIE is degraded by FtsH through its TagSpoIIE (red). (We cannot distinguish whether association with the divisome protects SpoIIE from proteolysis or if SpoIIE turns over while associated with the divisome.) Upon the completion of cytokinesis, SpoIIE transfers to the adjacent cell pole where multimerization protects it from proteolysis (as depicted by the light orange Tags) and leads to phosphatase activation. We propose that close proximity favors transfer to the immediately adjacent pole and that concentration of SpoIIE in the forespore, which is almost entirely derived from the pole, promotes multimerization.
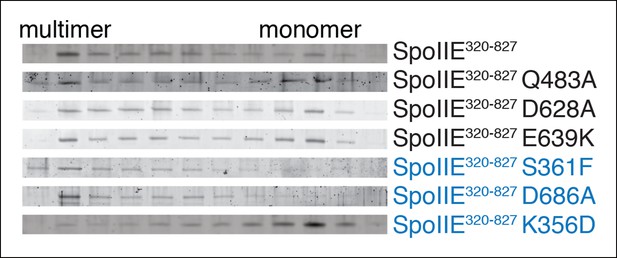
SpoIIE variants defective for sporulation multimerize.
Multimerization of SpoIIE variants was analyzed by gel filtration with a 24 ml Superose 6 column. 1 ml fractions from 7–18 ml of the run were collected, run on SDS-PAGE gels, and stained with SYPRO Ruby. SpoIIE variants that fail to accumulate in the forespore are marked in blue. Gels of wt SpoIIE and SpoIIEK356D are reproduced from Figure 7C for comparison.
Videos
Movie file of the sporulating cell shown in Figure 1C (2fps).
https://doi.org/10.7554/eLife.08145.005Movie file of the sporulating cell shown in Figure 1—figure supplement 1A (2fps).
SpoIIE-YFP is shown in grey, and the divisome marked by CFP-ZapA is shown in blue.
Movie file of the sporulating cell shown in Figure 1—figure supplement 1B (2fps).
SpoIIE-YFP is shown in grey, the divisome marked by CFP-ZapA is shown in blue, and the membrane marked by MalFtm-mNeptune is shown in red.
Additional files
-
Supplementary file 1
Table of strains.
- https://doi.org/10.7554/eLife.08145.020
-
Source code 1
Matlab scripts.
- https://doi.org/10.7554/eLife.08145.021





