YAP controls retinal stem cell DNA replication timing and genomic stability
Figures
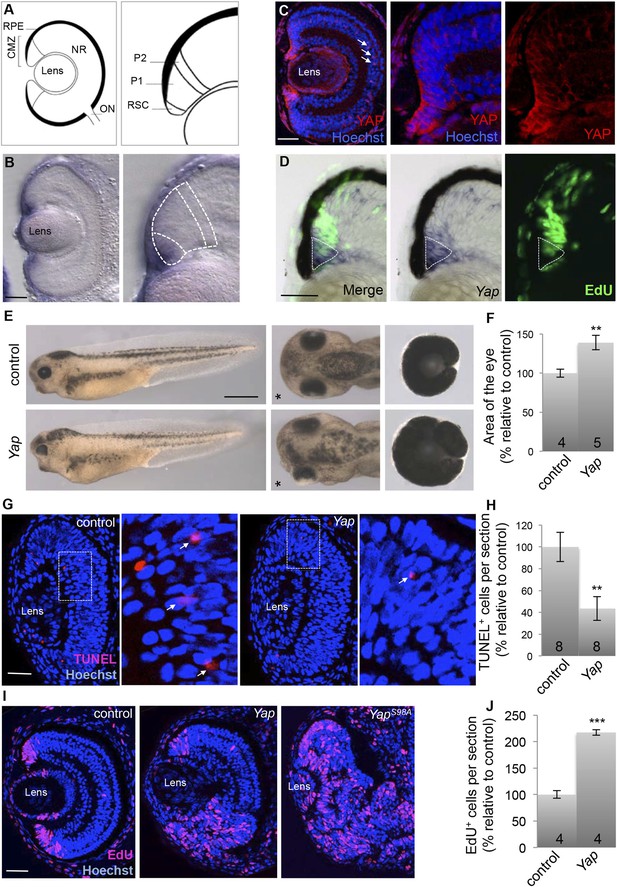
Yap overexpression expands the proliferating cell population in the post-embryonic retina.
(A) Schematic transversal section of a Xenopus tadpole retina (RPE: retinal pigment epithelium; NR: neural retina; ON: optic nerve). Within the CMZ (right panel), retinal stem cells (RSC) reside in the most peripheral margin while actively dividing progenitors (P1) and their post-mitotic progeny (P2) are localized more centrally. (B) In situ hybridization analysis of Yap expression on stage 40 retinal sections. The image on the right is a higher magnification of the CMZ (dashed lines represent the different zones as in a). (C) Immunostaining with anti-YAP antibody on stage 42 retinal sections. YAP labeling is detected in the CMZ as well as in Müller glial cells (arrows). Images on the right are higher magnifications of the CMZ. (D) EdU labeling (3-hr pulse) following in situ hybridization with a Yap probe (dotted line) on stage 40 retinal sections. (E) Lateral views (left panels), head dorsal views (middle panels) and dissected eyes (right panels) of stage 40 tadpoles following two-cell stage microinjection of GFP mRNA as a lineage tracer with either ß-gal (control) or Yap mRNA. The asterisk indicates the injected side. (F) Quantification of dissected eye area. (G–J) TUNEL (G, H; stage 33/34) or EdU incorporation (I, J; 3-hr pulse at stage 40) assays analyzed on retinal sections from tadpoles injected as in (E). Arrows point to TUNEL-positive cells in higher magnifications of the area delineated with dotted line (G). The number of analyzed retinas is indicated in each bar. Data are represented as mean ± SEM. Scale bar = 1 mm in (E) and 40 µm in other panels.
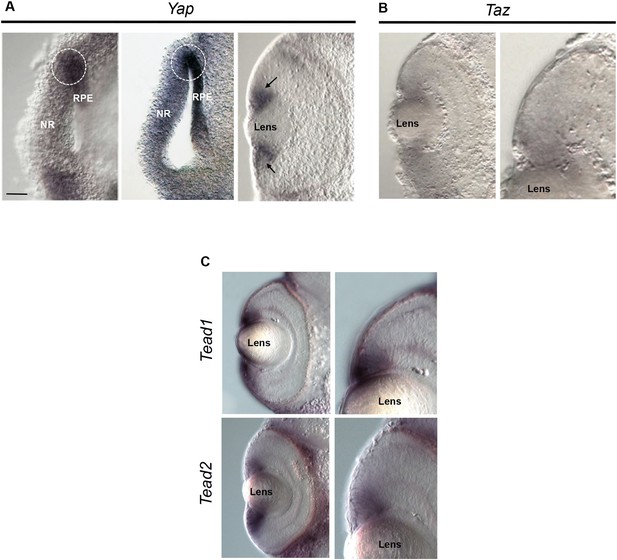
Yap, Taz and Tead expression.
(A) In situ hybridization analysis of Yap expression on stage 24–25 (left), 26–27 (middle) and stage 35 (right) retinal sections. Yap is expressed in the presumptive neural retina (NR) of the optic vesicle but is more strongly detected in the presumptive retinal pigmented epithelium (RPE), consistently with previous data in fish (Miesfeld and Link, 2014). It also labels the RPE/NR border region (delineated with dotted lines), which is believed to give rise to the CMZ (El Yakoubi et al., 2012). At latter stages, Yap gets restricted to the ciliary margin (black arrows). (B, C) In situ hybridization analysis of Taz (B), Tead1 and Tead2 (C) expression on stage 40 retinal sections. Images on the right are higher magnifications of the ciliary margin. Scale bar = 40 µm.
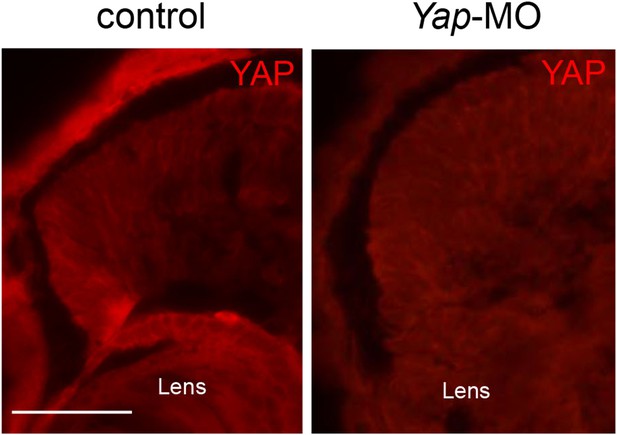
Validation of YAP antibody specificity.
Immunostaining with anti-YAP antibody on retinal sections from stage 42 tadpoles following microinjection of either Yap-5-mismatch-MO (control) or Yap-MO. YAP is undetectable in Yap morphants. Scale bar = 40 µm.
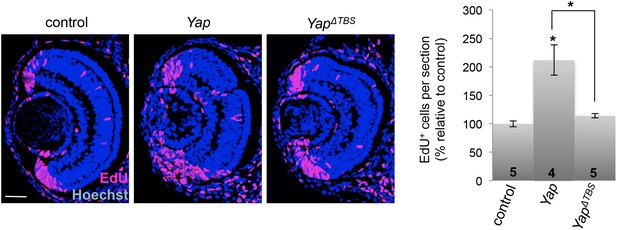
YapΔTBS does not promote CMZ cell proliferation.
EdU incorporation assays (3-hr pulse at stage 40) analyzed on retinal sections from tadpoles injected as in Figure 1E. The number of analyzed retinas is indicated in each bar. Data are represented as mean ± SEM. Scale bar = 40 µm.
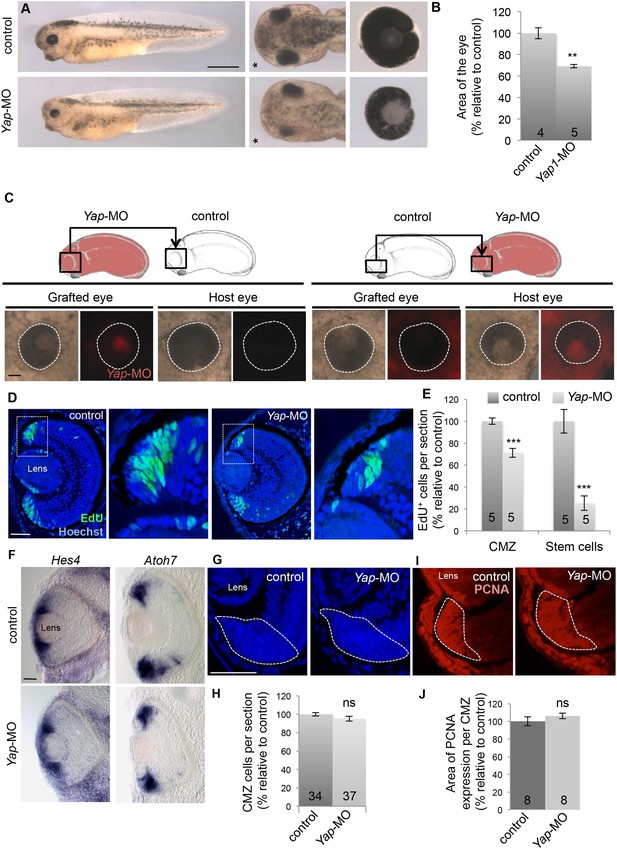
Yap knockdown decreases eye size and EdU incorporation in the post-embryonic retina.
(A) Lateral views (left panels), head dorsal views (middle panels) and dissected eyes (right panels) of stage 40 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO (control) or Yap-MO. The asterisk indicates the injected side. (B) Quantification of dissected eye area. (C) Eyes of stage 40 tadpoles following optic vesicle transplantation at stage 25 as shown in the schematic. Dotted lines delineate the eye circumference. (D) EdU incorporation assay (6-hr pulse) analyzed on retinal sections from stage 40 tadpoles injected as in (A). Higher magnifications of the CMZ (delineated by dotted lines) are shown for each condition. (E) Quantification of EdU-positive cells within the whole CMZ or within the most peripheral stem cell compartment. (F) In situ hybridization analysis of Hes4 (retinal stem cell marker; El Yakoubi et al., 2012) and Atoh7 (progenitor cell marker; Kanekar et al., 1997) expression on stage 40 retinal sections from tadpoles injected as in (A). (G–J) Hoechst staining and PCNA immunolabeling on stage 40 retinal sections from tadpoles injected as in (A). The CMZ is delineated with dotted lines. The number of analyzed retinas per condition is indicated in each bar. Data are represented as mean ± SEM. Scale bar = 1 mm in (A) and 40 µm in other panels.
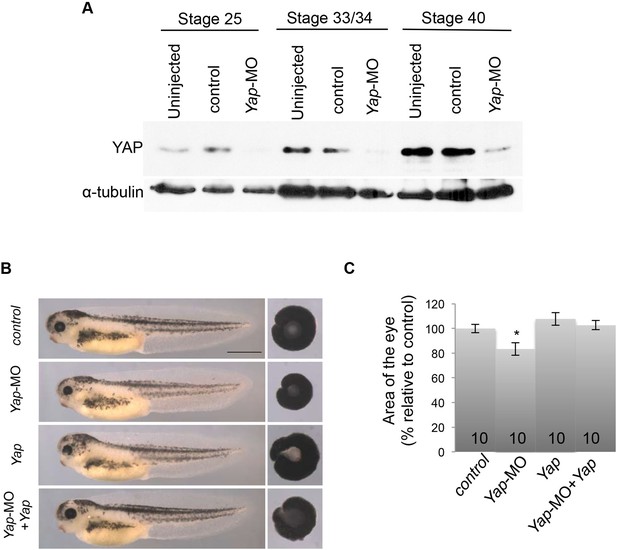
Validation of Yap-MO efficiency and specificity.
(A) Western blot analysis showing YAP expression decrease at different stages following microinjection of either Yap-5-mismatch-MO (control) or Yap-MO. α-tubulin is used as a loading control. (B) Lateral views and dissected eyes of stage 40 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO + ß-gal mRNA (control), Yap-MO + ß-gal mRNA (Yap-MO), Yap-5-mismatch-MO + Yap mRNA (Yap), Yap-MO + Yap mRNA (Yap-MO + Yap). (C) Quantification of dissected eye area. The Yap-MO-induced small eye phenotype is rescued by co-injection of Yap mRNA. Of note a suboptimal dose of Yap mRNA was used for the rescue experiment so that it does not alone give any eye phenotype. The number of analyzed tadpoles is indicated for each bar. Scale bar = 1 mm.
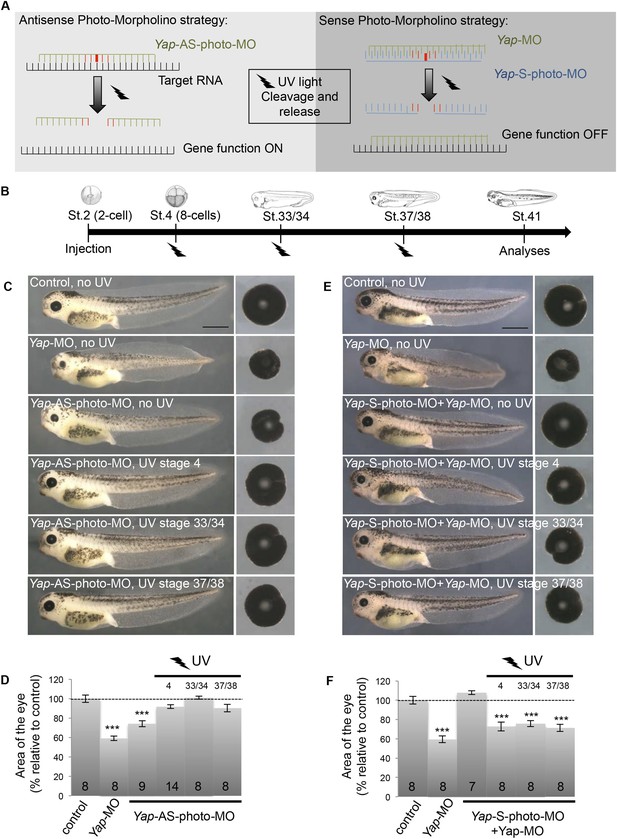
Conditional Yap knockdown in the retina.
(A) Principle of reversible and inducible gene knockdown using photo-Morpholinos (photo-MO). Photo-MO contains a photo-sensitive subunit cleaved by 365 nm light. Yap-AS-photo-MO is degraded upon UV light exposure and its translation blocking activity is thus interrupted. Unmodified Yap-MO is rendered inactive by binding to Yap-S-photo-MO. It therefore cannot bind its mRNA target until light-induced cleavage of the sense MO. (B) Diagram of the experimental design. Embryos are microinjected with MO at the two-cell stage, subjected to UV exposure at different developmental stages as indicated (black flashes) and then sacrificed for analyses. (C–F) Analysis of reversible (C, D) and inducible (E, F) Yap knockdown. (C, E) Lateral views and dissected eyes of stage 41 tadpoles following two-cell stage microinjection of the indicated MO (see table in B). (D, F) Quantification of dissected eye area. The stage at which UV exposure was performed is indicated above each bar. The Yap-AS-photo-MO (without any UV exposure) shows the same efficiency as the Yap-MO in reducing eye size. It is efficiently cleaved by UV light since exposure right after injection (stage 4) leads to a wild type phenotype. Restoring Yap function from stage 33/34 or even from stage 37/38 onwards leads to normal eye sized embryos, demonstrating that restricting Yap knockdown to embryogenesis is not sufficient to affect tadpole eye growth. The Yap-S-photo-MO efficiently blocks Yap-MO since their co-injection does not affect eye size. It is efficiently cleaved by UV light since exposure right after co-injection (stage 4) leads to a small eye phenotype, as observed in Yap-MO-injected embryos. Conditional Yap knockdown by light exposure from stage 33/34 or even from stage 37/38 onwards is sufficient to reduce eye size, suggesting that Yap is required at post-embryonic stages to maintain CMZ-dependent eye growth. Of note, in our experimental conditions, UV light exposure does not generate any significant effects on eye size (data not shown). The number of analyzed tadpoles is indicated within each bar. Scale bar = 1 mm.
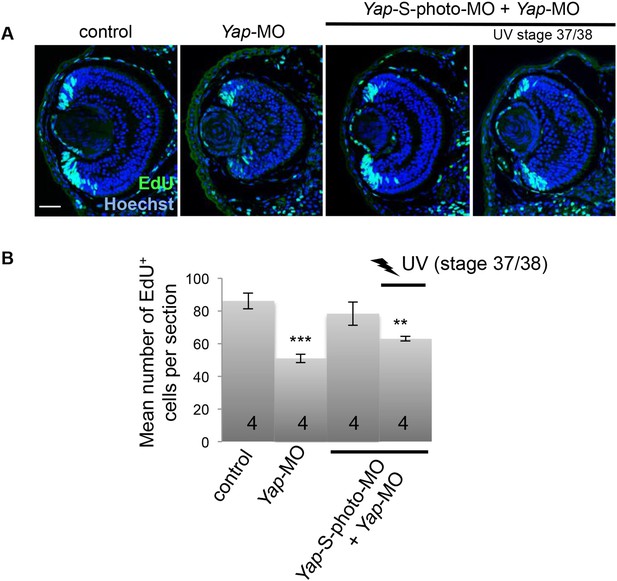
Inducible Yap knockdown at post-embryonic stages reduces EdU incorporation in the CMZ.
(A) EdU incorporation assay (3-hr pulse) analyzed on retinal sections from stage 41 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO (control), Yap-MO and/or Yap-S-photo-MO, as indicated in the table. (B) Quantification of EdU-positive cells within the CMZ. Conditional Yap knockdown by light exposure from stage 37/38 onwards is sufficient to reduce EdU incorporation. The total number of analyzed retinas per condition is indicated in each bar. Scale bar = 40 µm.
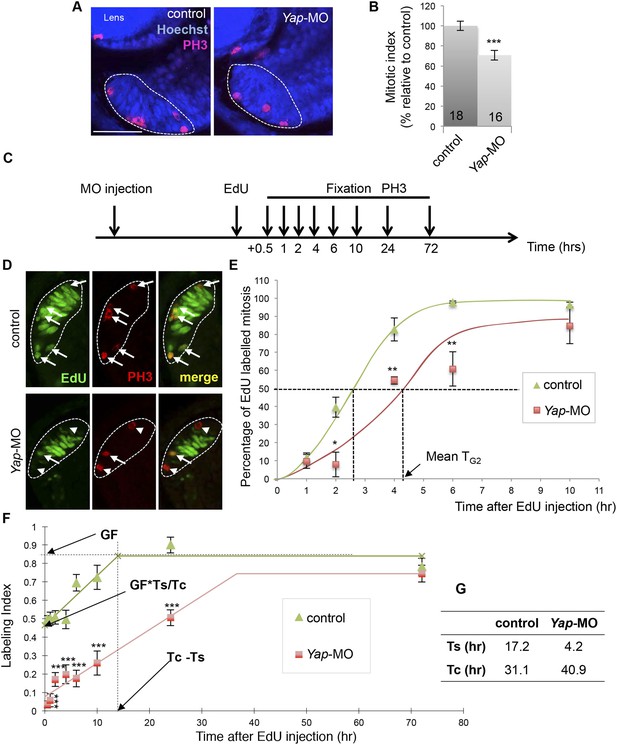
Yap loss of function slows down cell cycle kinetics of retinal stem cells.
(A) PH3 immunolabeling on retinal sections from stage 40 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO (control) or Yap-MO. The CMZ is delineated with dotted lines. (B) Quantification of the mitotic index within the CMZ. The number of analyzed retinas per condition is indicated in each bar. (C) Outline of the PLM and EdU cumulative labeling experiments: tadpoles injected as in (A) were fixed at different time points following EdU injection at stage 39. EdU and PH3 labeling was then analyzed on retinal sections. (D) Retinal sections stained for both PH3 and EdU. The CMZ is delineated with dotted lines. Arrows and arrowheads point to PH3+/EdU+ and PH3+/EdU− cells respectively. (E) Quantification of the PLM within the whole CMZ. TG2: G2-phase duration. (F) Quantification of the EdU labeling index within retinal stem cells along with increasing EdU exposure times. GF: growth fraction; TC: total cell cycle; TS: S-phase. (G) Estimation of TC and TS. Data are represented as mean ± SEM. Scale bar = 40 µm.
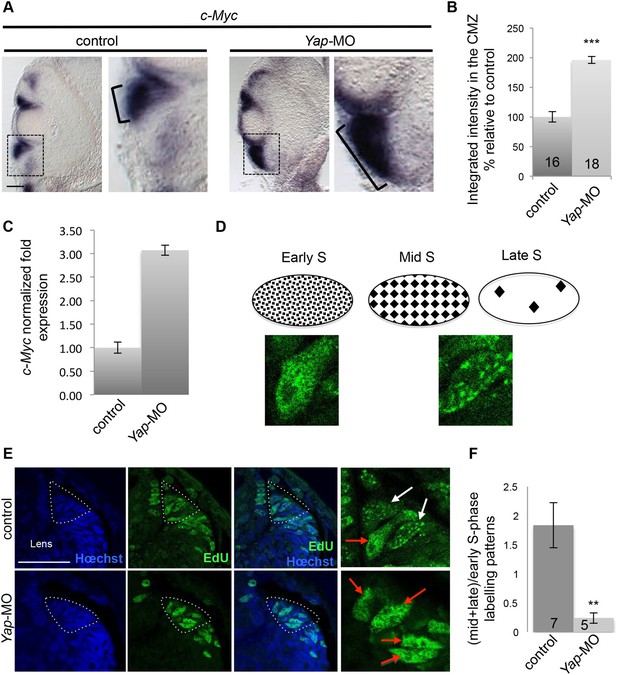
Yap loss of function affects the temporal program of retinal stem cell DNA replication.
(A) In situ hybridization analysis of c-Myc expression on stage 40 retinal sections following two-cell stage microinjection of Yap-5-mismatch-MO (control) or Yap-MO. Images on the right show higher magnifications of the CMZ (dotted lines). Note the strong expansion of c-Myc expression area (bracket). (B) Quantification of the staining in the CMZ. The number of analyzed retinal sections per condition is indicated in each bar. (C) qPCR analysis of c-Myc expression in the retina of tadpoles injected as in (A). (D) Schematic representation of replication foci observed during S-phase progression, as inferred from EdU labeling. Pictures illustrate two examples of EdU-labeled foci observed in control CMZ cells, one homogenous (early S-phase) and one with large dots (mid/late S-phase). (E) Analysis of EdU-labeled replication foci (1 hr-pulse) in the CMZ of stage 40 tadpoles injected as in (A). Enlargements of the CMZ tip (dotted lines) are shown on the right. Early (red arrows) and mid/late profiles (white arrows) were distinguished. (F) Corresponding quantification. The number of analyzed retinas per condition is indicated in each bar. Data are represented as mean ± SEM. Scale bar = 40 µm.
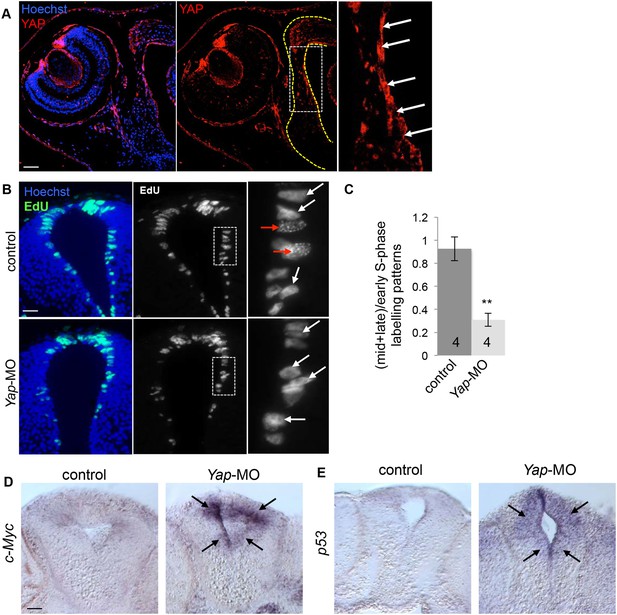
Effects of Yap knockdown in the neural tube.
(A) Immunostaining with anti-YAP antibody on stage 42 sections. The left side of the neural tube is delineated with yellow dotted line. A higher magnification of the ventricular zone (white dotted line) is provided in the right panel. YAP labeling is most strongly detected in this region where progenitor cells reside (arrows). (B) Analysis of EdU-labeled replication foci (1 hr-pulse) in the neural tube of stage 40 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO (control) or Yap-MO. Enlargements (dotted lines) are shown on the right. Early (red arrows) and mid/late profiles (white arrows) were distinguished. (C) Corresponding quantification. The number of analyzed tadpoles per condition is indicated in each bar. Data are represented as mean ± SEM. (D, E) In situ hybridization analysis of c-Myc or p53 expression in the neural tube of stage 40 tadpoles injected as in (B). Note the strong upregulation in the ventricular zone of the neural tube (black arrows). Scale bars = 40 µm.
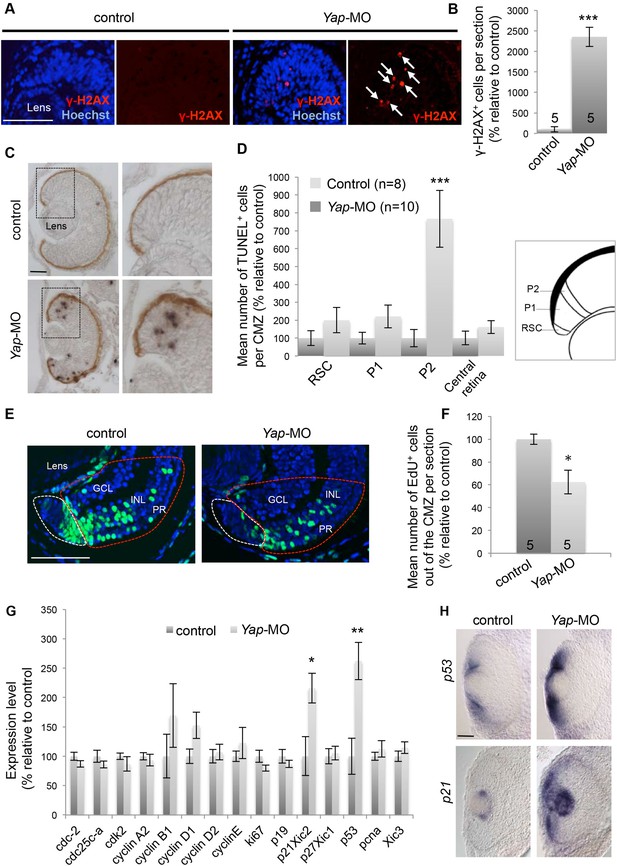
Yap loss of function induces DNA damage.
(A) γ-H2AX immunolabeling in the CMZ of retinal sections from stage 40 tadpoles following two-cell stage microinjection of Yap-5-mismatch-MO (control) or Yap-MO. Arrows point to γ-H2AX-positive cells. (B) Corresponding quantification. (C) TUNEL assay on retinal sections from stage 40 tadpoles injected as in (A). Images on the right show higher magnifications of the CMZ delineated with dotted lines. (D) Quantification of TUNEL-positive cells in the different compartments of the CMZ as illustrated on the schematic. (E) 2 days-chase of EdU-labeled cells in the CMZ of stage 42 tadpoles injected as in (A). EdU-positive cells inside the zone encircled with a red dotted line have exited the CMZ (white dotted lines) and integrated the different retinal layers. GCL: ganglion cell layer; INL: inner nuclear layer; PR: photoreceptor layer. (F) Quantification of EdU-positive cells in the neural retina layers. (G) mRNA expression levels of cell cycle genes as measured with the NanoString nCounter system in heads from stage 40 tadpoles following two-cell stage microinjection of Standard MO (control) or Yap-MO. Data are the mean of four independent experiments. (H) In situ hybridization analyses of p53 and p21 expression on retinal sections from stage 40 tadpoles injected as in (A). The number of analyzed retinas per condition is indicated in each bar (B, F) or on the graph (n in D). Data are represented as mean ± SEM. Scale bar = 40 µm.
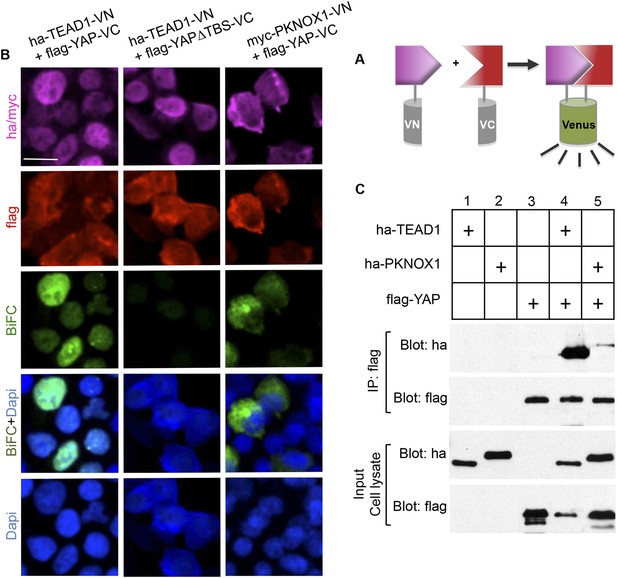
Physical interaction between YAP and PKNOX1.
(A) Schematic representation of BiFC principle. (B) Immunolabeling/BiFC analysis on HEK293T cells transfected with VN and VC chimeric constructs, as indicated. (C) Co-immunoprecipitation assay of 293T cells transfected with tagged constructs as indicated. Scale bars = 20 µm.
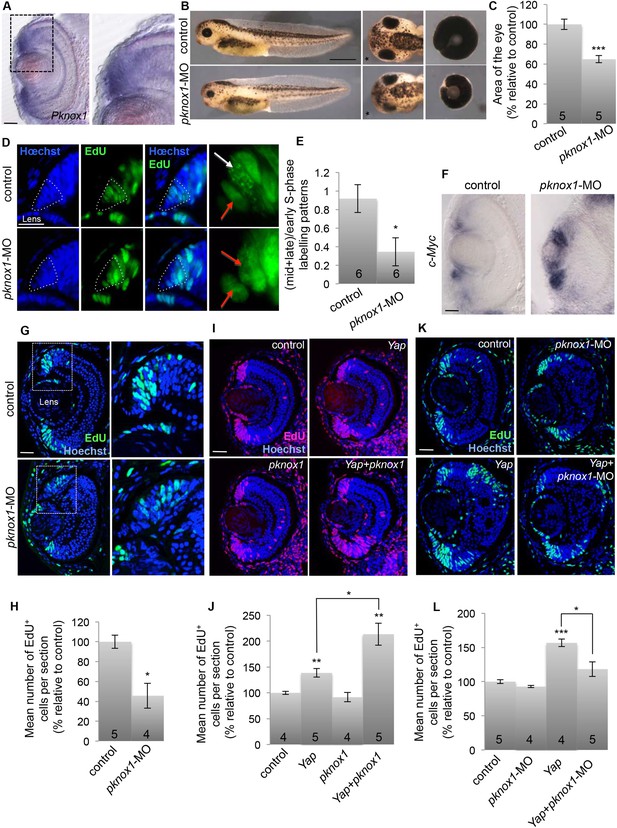
Functional interaction between YAP and PKNOX1.
(A) In situ hybridization analysis of pknox1 expression on stage 40 retinal sections. The right panels shows an enlargement of the CMZ region delineated with dotted lines. (B) Lateral views (left panels), head dorsal views (middle panels) and dissected eyes (right panels) of stage 40 tadpoles following two-cell stage microinjection of pknox1-5-mismatch-MO (control) or pknox1-MO. The asterisk indicates the injected side. (C) Quantification of dissected eye area. (D) Analysis of EdU-labeled replication foci (45 min-pulse) in the CMZ of tadpoles injected as in (B). Enlargements of the CMZ tip (dotted lines) are shown on the right. Early (red arrows) and mid/late profiles (white arrows) were distinguished. (E) Corresponding quantification. (F) In situ hybridization analysis of c-Myc expression on stage 40 retinal sections from tadpoles injected as in (B). (G–L) EdU incorporation assays (3-hr pulse) analyzed on retinal sections from stage 40 tadpoles. (G, H) shows the effect of pknox1 knockdown (injection of pknox1-5-mismatch-MO (control) or pknox1-MO). (I, J) shows the synergistic effects of pknox1 and Yap (injection of GFP mRNA and either ß-gal mRNA (control), Yap + ß-gal mRNA (Yap), pknox1 + ß-gal mRNA (pknox1), Yap + pknox1 mRNA (Yap + pknox1)). (K, L) shows the rescue of Yap overexpression by pknox1 knockdown (injection of either pknox1-5-mismatch-MO + ß-gal mRNA (control), pknox1-MO + ß-gal mRNA (pknox1-MO), pknox1-5-mismatch-MO + Yap mRNA (Yap), pknox1-MO + Yap mRNA (Yap + pknox-MO)). Of note, a suboptimal dose of pknox1-MO was used for the rescue experiment so that it does not alone give any eye phenotype. The total number of analyzed retinas per condition is indicated in each bar. Scale bar = 1 mm in (B) and 40 µm for all other panels.
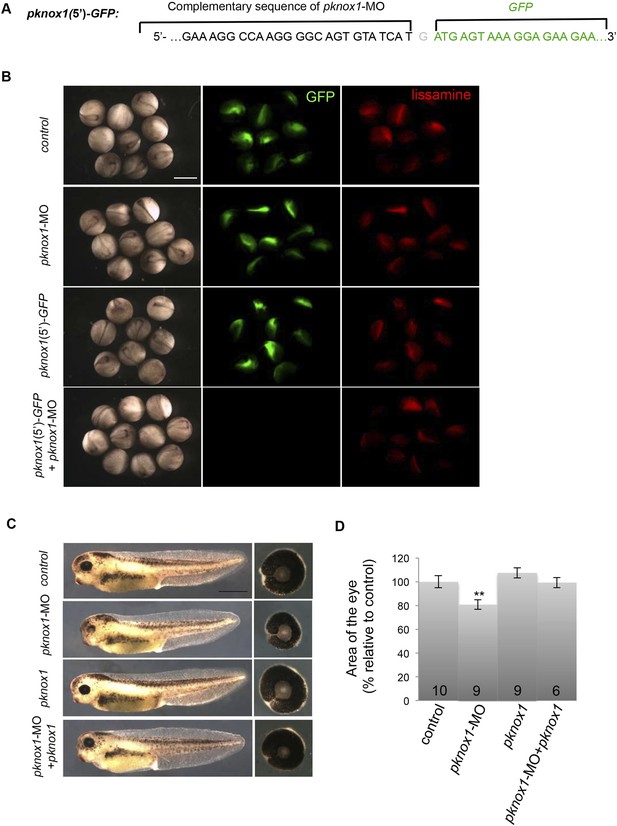
Validation of pknox1-MO efficiency and specificity.
(A) Schematic representation of the chimeric construct containing GFP downstream of pknox1-MO complementary sequence (pknox1(5′)-GFP). (B) GFP fluorescence analysis at stage 18 following two-cell stage microinjection of GFP mRNA + pknox1-5-mismatch MO (control), GFP mRNA + pknox1-MO (pknox1-MO), pknox1(5′)-GFP + pknox1-5-mismatch-MO (pknox1(5′)-GFP) or pknox1(5′)-GFP + pknox1-MO. Fluorescence imaging at 594 nm detects the MO-bound lissamine tag. pknox1-MO efficiently inhibits GFP translation of pknox1(5′)-GFP mRNA. (C) Lateral views and dissected eyes of stage 40 tadpoles following two-cell stage microinjection of pknox1-5-mismatch-MO + ß-gal mRNA (control), pknox1-MO + ß-gal mRNA (pknox1-MO), pknox1-5-mismatch-MO + pknox1 mRNA (pknox1), pknox1-MO + pknox1 mRNA (pknox1-MO + pknox1). (D) Quantification of dissected eye area. The pknox1-induced small eye phenotype is rescued by co-injection with pknox1 mRNA. The number of analyzed tadpoles is indicated for each bar. Scale bar = 1 mm.
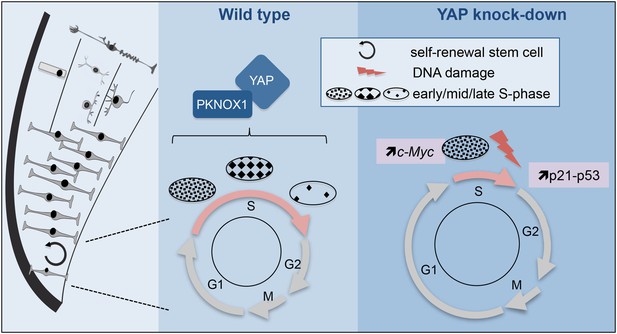
Model illustrating YAP function in retinal stem cells.
We found that YAP is expressed in CMZ retinal stem cells (left panel). The middle panel shows the cell cycle of wild type retinal stem cells and the putative role of the YAP/PKNOX1 complex in the control of S-phase temporal progression (represented by the distinct patterns of DNA replication foci). YAP knock-down (right panel) leads to a dramatic reduction of S-phase length likely due to c-Myc-dependent premature firing of late replication origins. This results in increased occurrence of DNA damage, enhanced p21 and p53 expression and eventually cell death.
Additional files
-
Supplementary file 1
Sequences of Morpholino oligonucleotides and primers used in the study.
- https://doi.org/10.7554/eLife.08488.019
-
Supplementary file 2
List of antibodies used in the study.
- https://doi.org/10.7554/eLife.08488.020
-
Supplementary file 3
Sequences used in the NanoString experiment.
- https://doi.org/10.7554/eLife.08488.021





