HIV Tat controls RNA Polymerase II and the epigenetic landscape to transcriptionally reprogram target immune cells
Figures
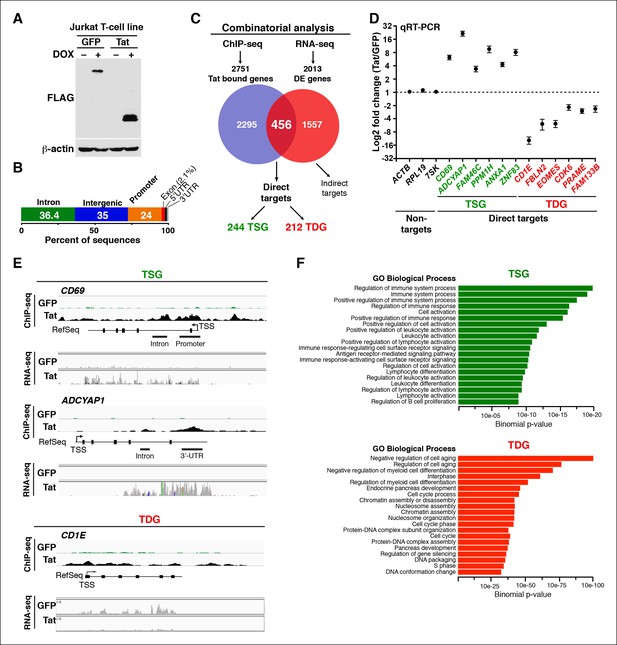
Genomic domains occupied and regulated by Tat in CD4+ T cells.
(A) Western blot of Jurkat-GFP and -Tat cell lines treated (+) or not (–) with DOX using the indicated antibodies. (B) Genome-wide distribution of Tat across the human genome. (C) Integration of the FLAG ChIP-seq and RNA-seq datasets defines a set of genes directly regulated by Tat. (D) Validation of the RNA-seq dataset using qRT-PCR on the indicated TSG, TDG or non-target genes as negative controls (mean ± SEM; n = 3). (E) Individual tracks showing FLAG ChIP-seq and the corresponding RNA-seq dataset in the GFP and Tat cell lines. (F) Functional annotation of biological processes enriched at TSG and TDG. This figure is associated with Figure 1—figure supplements 1–10. Direct targets, genes directly bound and regulated by Tat; ChIP-seq, chromatin immunoprecipitation sequencing; DOX, doxycycline; GFP, green fluorescent protein; RNA-seq, RNA sequencing; qRT-PCR, quantitative real time polymerase chain reaction; TDG, Tat downregulated genes; TSG, Tat stimulated genes; TSS, transcription start site.
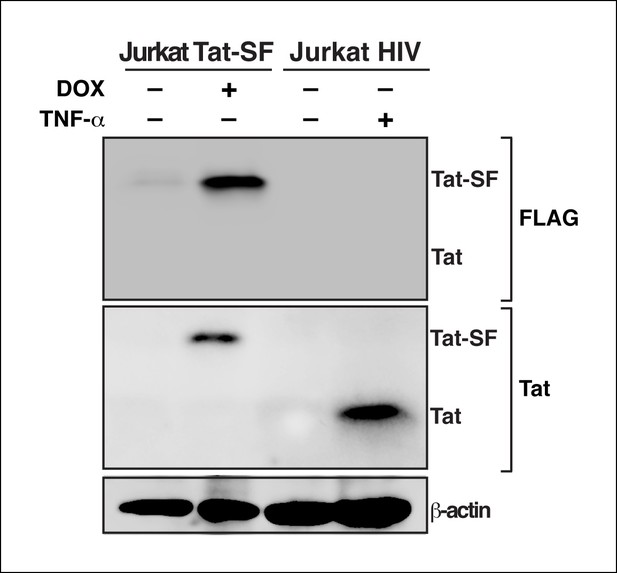
Tat protein expression levels in the Jurkat Tat-SF model matches the levels of Tat detected during HIV infection.
Western blot (FLAG and Tat) of Jurkat Tat-SF cell line treated with (+) or without (–) DOX, and Jurkat cells latently infected with HIV (clone E4) induced with (+) or without (–) TNF-α for 24 hr. A β-actin western blot is shown as loading control. DOX, doxycycline; TNF-α, tumor necrosis factor.
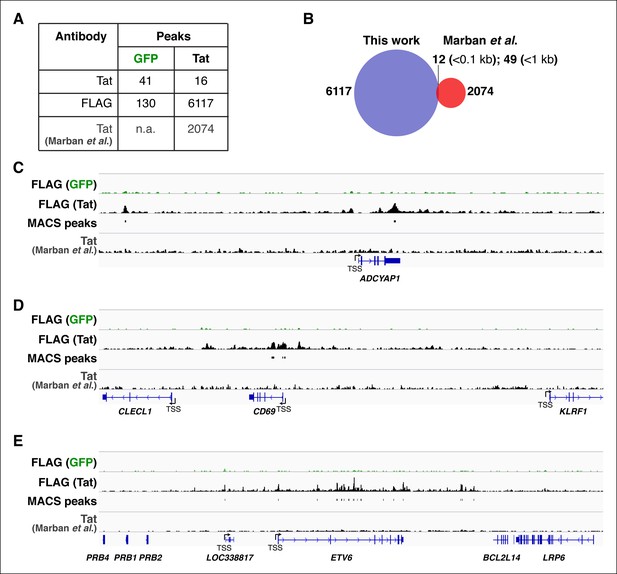
Technical improvement of Tat ChIP-seq in CD4+ T cells.
(A) Comparison of ChIP-seq Tat peak numbers in the Jurkat-GFP and -Tat cell lines using the Tat and FLAG antibodies. The number of Tat peaks in a similar Jurkat-Tat cell line from the study of Marban et al., (2011) is indicated. (B) Overlay of Tat peaks from this study and those identified by Marban et al., (2011)using ChIP-seq. The number of common Tat peaks between our ChIP-seq data and Marban’s at a distance of <0.1 kb or <1 kb from the Tat sites is shown. (C–E) Genome browser views of FLAG ChIP-seq tracks in the GFP and Tat cell lines along with the FLAG peaks called by MACS in the Tat cell line and the Tat track by Marban et al., (2011). ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; MACS, model-based analysis of ChIP-seq.
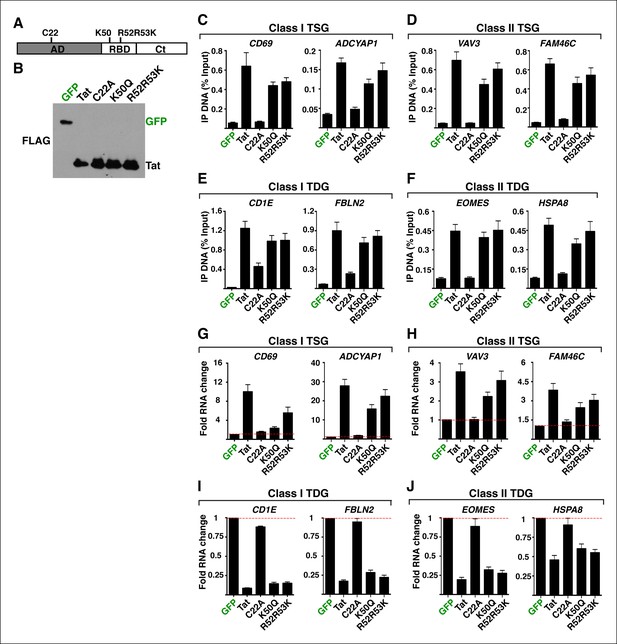
Non-functional Tat mutants have compromised chromatin interaction and modulation of cellular gene expression.
(A) Scheme of Tat showing the position of its domains (AD, RBD and Ct) along with the location of the mutated residues. (B) Western blots of Jurkat CD4+ T cell lines expressing GFP, wild-type Tat or non-functional mutants (C22A, K50Q, R52R53K) (D'Orso et al., 2012). Cells from panel (B) were used in ChIP assays to analyze the occupancy of GFP, Tat or the non-functional mutants at class I TSG promoters (C), class II TSG promoters (D), class I TDG promoters (E) and class II TDG promoters (F). Values representing the average of three independent experiments (mean ± SEM; n = 3). Cells from panel (B) were used to isolate total RNA and the expression of class I TSG (G), class II TSG (H), class I TDG (I) and class II TDG (J) was measured by qRT-PCR, normalized to RPL19, and plotted as fold RNA change over the GFP line arbitrarily set at 1 (mean ± SEM; n = 3). AD, activation domain; ChIP-seq, chromatin immunoprecipitation sequencing; Ct, C-terminal domain; GFP, green fluorescent protein; qRT-PCR, quantitative real time polymerase chain reaction; RBD, RNA-binding domain; TDG, Tat downregulated genes; TSG, Tat stimulated genes.
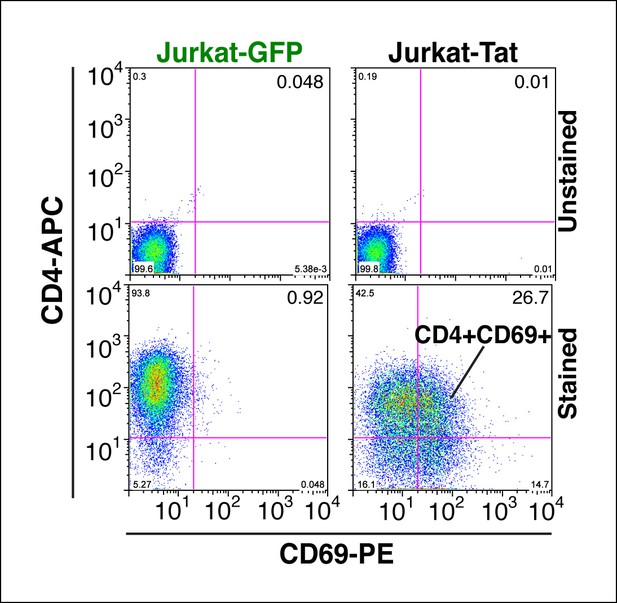
Tat-induced transcriptome changes are also observed at the protein level.
Flow cytometry analysis of Jurkat-GFP and -Tat cell lines induced with doxycycline for 24 hr and stained with CD4-APC and CD69-PE antibodies or unstained (negative control) to monitor levels of CD4 and CD69 proteins expressed at the cell surface. Note the increase in the CD69+ population in the presence of Tat. GFP, green fluorescent protein.
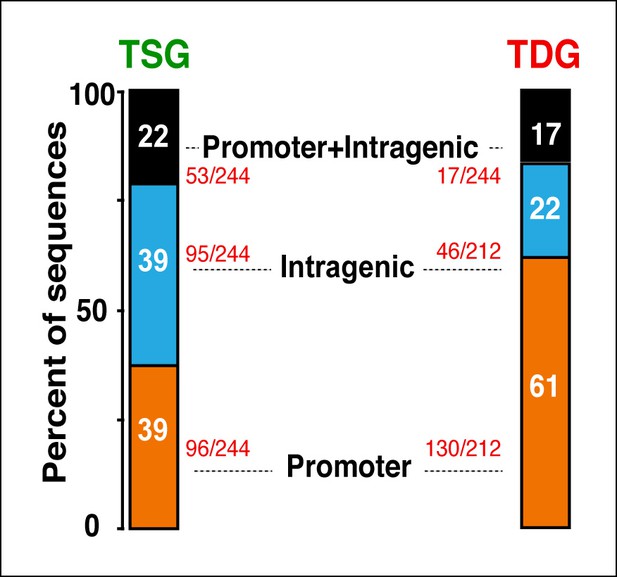
Distribution of Tat occupancy at promoter and/or intragenic domains in TSG and TDG.
The percentage of sequences from the total is indicated. TDG, Tat downregulated genes; TSG, Tat stimulated genes.
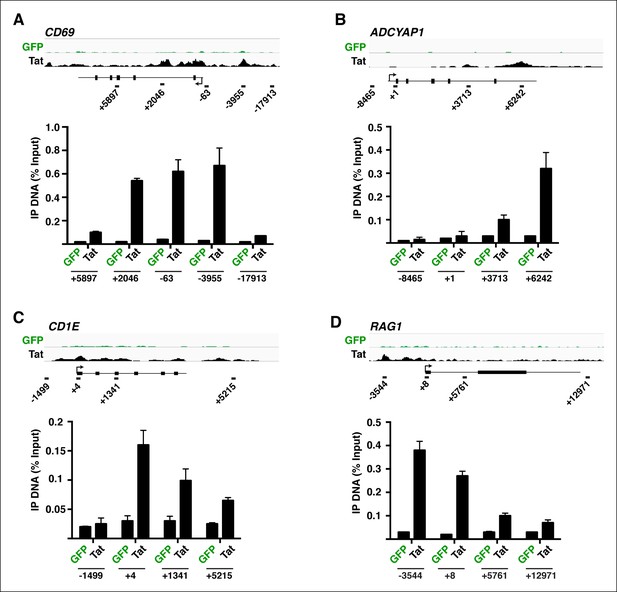
FLAG ChIP-qPCR analysis on the indicated genomic loci.
ChIP assay to analyze the distribution of Tat or GFP at the CD69 (A), ADCYAP1 (B), CD1E (C) and RAG1 (D) locus in the GFP (green) and Tat (black) cell lines. The position of the amplicons used in ChIP-qPCR and their distance to the TSS (arrow) is shown with the schematic of each locus. The schemes are not in real scale. Values represent the average of three independent experiments (mean ± SEM; n = 3). ChIP, chromatin immunoprecipitation; GFP, green fluorescent protein; qPCR, quantitative polymerase chain reaction; SEM, standard error of the mean; TSS, transcription start site.
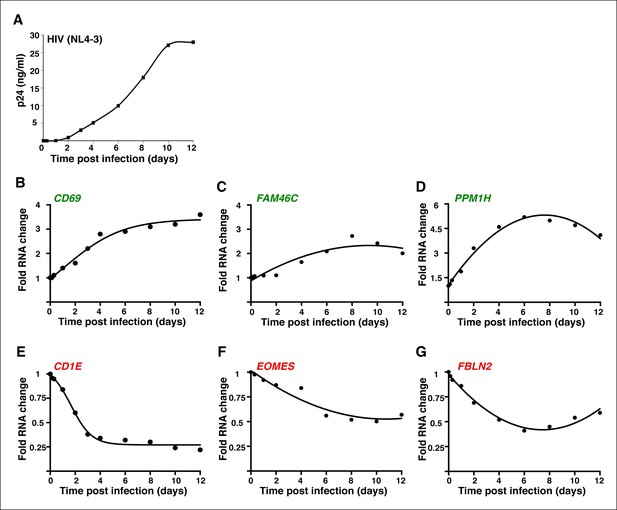
The genes modulated by ectopic expression of Tat are also detected during a time-course HIV infection experiment.
(A) Jurkat T cells were infected with HIV (NL4-3) and levels of p24/Capsid protein was quantified using ELISA at different time points post-infection (0, 3 and 7 hr; 1, 2, 4, 6, 8, 10, and 12 days). Values represent the average of three independent experiments (mean ± SEM; n = 3). Cells from panel (A) were used to isolate total RNA and the expression of three TSG: CD69 (B), FAM46C (C), and PPM1H (D); and three TDG: CD1E (E), EOMES (F) and FBLN2 (G) normalized to RPL19 was measured by qRT-PCR and plotted as fold RNA change over the GFP cell line arbitrarily set at 1 (mean ± SEM; n = 3). The points in the curve were fitted to a non-linear regression in GraphPad Prism. ELISA, enzyme-linked immunosorbent assay; GFP, green fluorescent protein; HIV, human immunodeficiency virus; qRT-PCR, quantitative real time polymerase chain reaction; SEM, standard error of the mean; TSG, Tat stimulated genes; TDG, Tat downregulated genes.
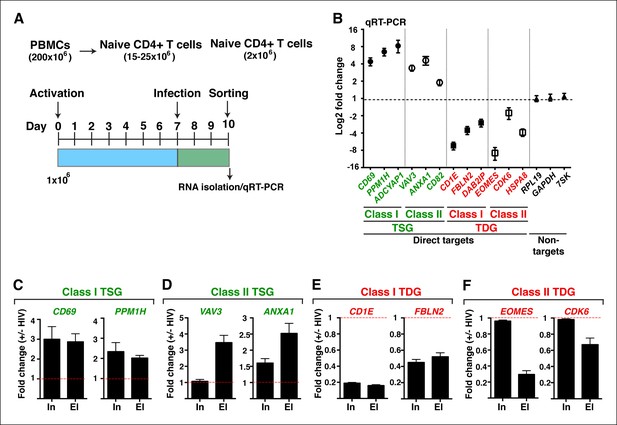
HIV infection of central memory CD4+ T cells triggers deregulation of TSG and TDG detected in the genome-wide approaches.
(A) Scheme of the pipeline used to generate primary central memory T cells (TCM) and infect with replication competent HIV to identify differentially expressed genes. (B) qRT-PCR analysis on the indicated class I and II TSG, TDG and non-target genes (mean ± SEM; n = 3). Cells from panel (A) were used to isolate total RNA and the expression of initiating (In) and elongating (El) transcripts for class I TSG (C), class II TSG (D), class I TDG (E) and class II TDG (F) was measured by qRT-PCR, normalized to RPL19, and plotted as fold RNA change: HIV infection/mock infection (mean ± SEM; n = 3). HIV, human immunodeficiency virus; qRT-PCR, quantitative real time polymerase chain reaction; TDG, Tat downregulated genes; TSG, Tat stimulated genes; SEM, standard error of the mean.
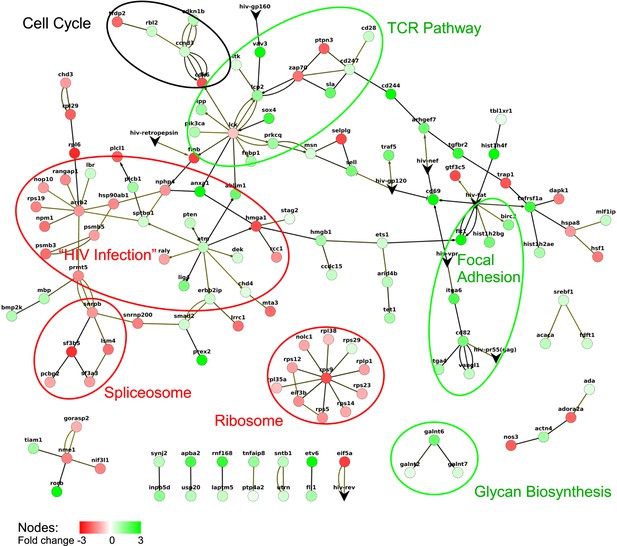
Response network of TSG and TDG.
A response network was constructed based on RNA-seq data using a p-value cut off p = 0.01. The network consists of 138 nodes and 161 edges. Multiple edges between nodes indicate multiple evidence from different protein-protein interaction datasets ([Human Protein Reference Database] [Goel et al., 2011]; IntAct [Kerrien et al., 2012]; NetworKin, BHMRSS, CORUM, Hynet [Konig et al., 2010] and NCBIs HIV-1 interaction databases). Green and red nodes denote TSG and TDG, respectively. Edges are directional according to the respective databases. Edges with circles as ‘arrow tips’ denote phosphorylation reactions, ‘diamond shaped tips’ refer to general activation, and ‘arrows’ describe other reactions. Non-directional edges indicate binding. HIV, human immunodeficiency virus; RNA-seq, RNA sequencing; TDG, Tat downregulated genes; TSG, Tat stimulated genes.
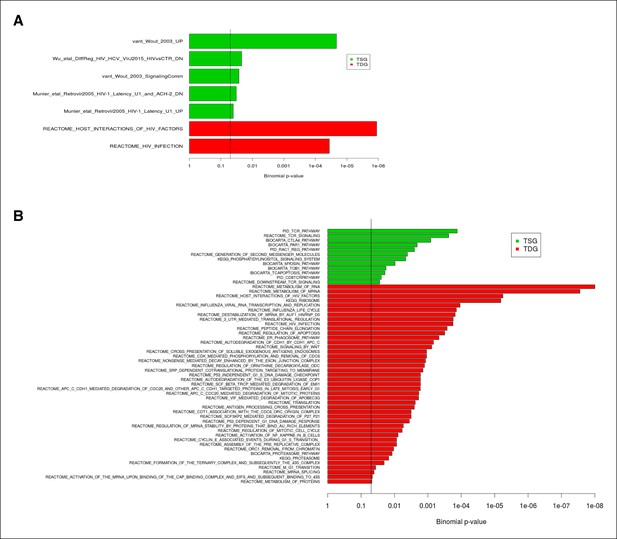
Enrichment of TSG and TDG in publicly available datasets of differentially expressed genes identified in HIV infection and replication experiments.
(A) The enrichment analysis with 48 publicly available differentially expressed gene datasets identified in HIV and replication experiments and 14 HIV relevant pathways from MSigDB are shown. (B) The functional annotation by the MSigDB canonical pathways (set c2.cp) is depicted. Twelve significantly TSG enriched gene sets and 43 significantly TDG enriched gene-sets have been identified. P-values are Bonferroni adjusted for multiple testing. The vertical line indicates FDR = 0.05. FDR, false discovery rate; HIV, human immunodeficiency virus; MSigDB, Molecular Signatures Database; TDG, Tat downregulated genes; TSG, Tat stimulated genes.
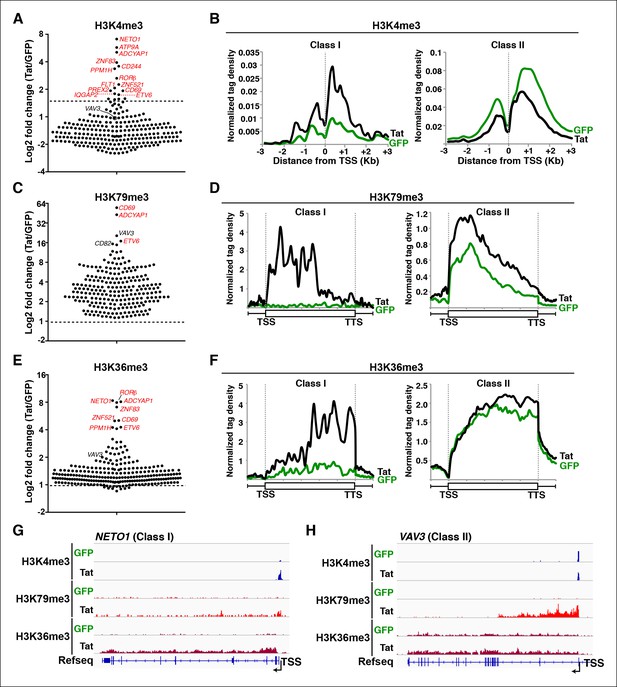
Global analysis of chromatin signatures reveals that Tat activates the transcription initiation and elongation steps.
(A) Dot plots of H3K4me3 log2 fold change in the region encompassing ±3 Kb from the TSS of all TSG. Genes are divided into two clusters: class I and II based on increased (>1.5-fold change) or no change/decreased H3K4me3 levels in the presence of Tat. Selected TSG examples are indicated in red. (B) Metagene plots centered on TSS showing H3K4me3 occupancy profiles at both class I (n = 17) and class II (n = 43) TSG in the presence of Tat or GFP. (C) Dot plots of H3K79me3 log2 fold change in the region from TSS to TTS (see Materials and methods). (D) Metagene analysis showing average H3K79me3 ChIP-seq signals at both class I and II TSG in the presence of Tat or GFP. Units are mean tags per million ChIP-seq reads per bin across the transcribed region of each gene with 2 kb upstream and downstream flanking regions. (E) Dot plots of H3K36me3 log2 fold change in the region from TSS to TTS. (F) Metagene plots showing average H3K36me3 ChIP-seq signals at both class I and cII TSG in the presence of Tat or GFP. (G) Genome browser views showing ChIP-seq signal at a class I TSG (NETO1) in the GFP and Tat cell lines. (H) Genome browser views showing ChIP-seq signal at a class II TSG (VAV3) in the GFP and Tat cell lines. This figure is associated with Figure 2—figure supplements 1–5. ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; TSG, Tat stimulated genes; TSS, transcription start site; TTS, transcription termination site.
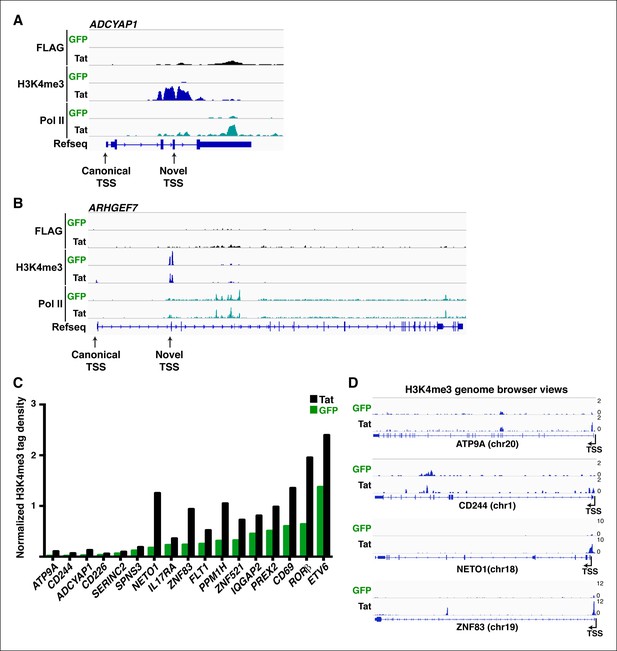
Tat specifies TSS selection and synthesis of alternate isoforms.
(A) Genome browser views of FLAG, H3K4me3 and Pol II ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the ADCYAP1 locus. The position of the canonical and Tat-induced ‘novel’ TSS is indicated with arrows. (B) Genome browser views of FLAG, H3K4me3 and Pol II ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the ARHGEF7 locus. The position of the canonical and Tat-induced ‘novel’ TSS is indicated with arrows. (C) Normalized H3K4me3 tag density (-/+ 3 Kb respective to the TSS) for class I TSG in the Tat (black) and GFP (green) cell lines. (D) Genome browser views of H3K4me3 ChIP-seq tracks in the GFP and Tat cell lines for four class I TSG from panel (C). ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; Pol, polymerase; TSG, Tat stimulated genes.
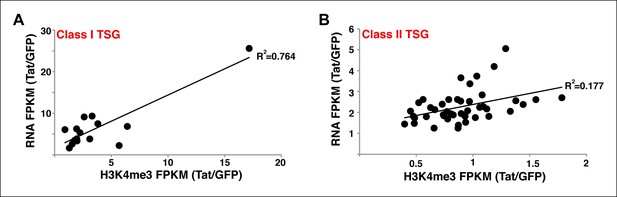
Correlation between gene expression levels and H3K4me3 density surrounding the TSS at class I and II TSG.
(A) Correlation plot between normalized H3K4me3 tag density (-/+ 3 kb respective to the TSS) and total gene expression levels (based on RNA-seq FPKM) at class I TSG in the presence and absence of Tat (Tat/GFP). (B) Correlation plot between normalized H3K4me3 tag density (± 3 kb respective to the TSS) and total gene expression levels (based on RNA-seq) at class II TSG in the presence and absence of Tat (Tat/GFP). GFP, green fluorescent protein; RNA-seq, RNA sequencing; TSG, Tat stimulated genes; TSS, transcription start site.
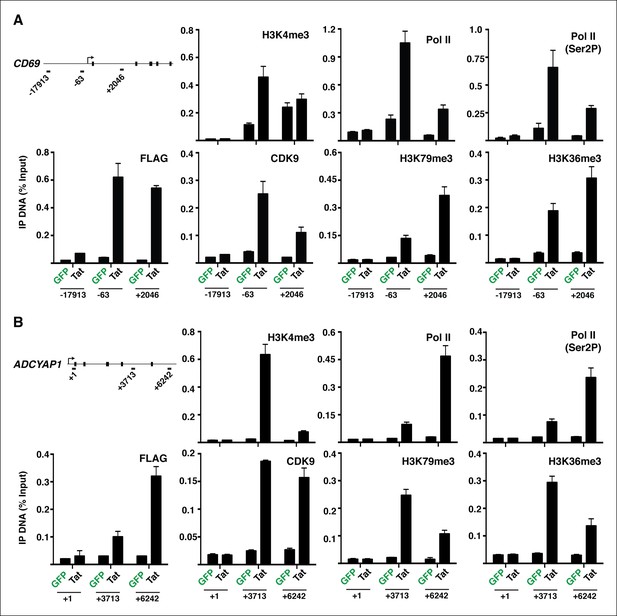
Evidence that Tat increases Pol II and P-TEFb recruitment, and chromatin marks coinciding with transcription initiation and elongation at class I TSG.
(A) Jurkat-GFP and -Tat cell lines were used in ChIP assays to analyze the occupancy of GFP and Tat (FLAG), H3K4me3, H3K79me3, H3K36me3, Pol II (total), Pol II (Ser2P-CTD form) and P-TEFb (CDK9) at the CD69 locus with the three indicated amplicons. The numbers indicate the position of the amplicons respective to the TSS. (B) Jurkat-GFP and -Tat cell lines were used in ChIP assays to analyze the occupancy of GFP and Tat (FLAG), H3K4me3, H3K79me3, H3K36me3, Pol II (total), Pol II (Ser2P-CTD form) and P-TEFb (CDK9) at the ADCYAP1 locus with the three indicated amplicons. The numbers indicate the position of the amplicons respective to the TSS. For both panels, IP DNA (% Input) values represent the average of three independent experiments (mean ± SEM; n = 3). ChIP, chromatin immunoprecipitation; GFP, green fluorescent protein; Pol, polymerase; P-TEFb, positive transcription elongation factor b; SEM, standard error of the mean; TSS, transcription start site.
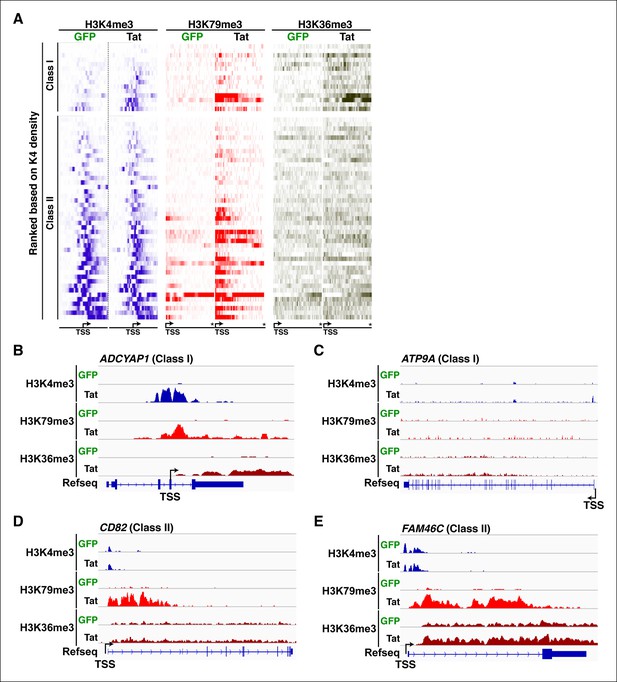
Transcription initiation correlates with increased elongation chromatin markers.
(A) Heatmap representation of ChIP-seq binding for H3K4me3 (blue), H3K79me3 (red), and H3K36me3 (brown) at class I and class II TSG, rank ordered from lowest to most H3K4me3 density increase from the GFP to the Tat cell line. The asterisk denotes the position of the TTS. (B) Genome browser views of H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the ADCYAP1 locus (class I TSG). The position of the TSS is indicated with an arrow. (C) Genome browser views of H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the ATP9A locus (class I TSG). (D) Genome browsers of H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the CD82 locus (class II TSG). (E) Genome browser views of H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the FAM46C locus (class II TSG). ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; TSG, Tat stimulated genes; TSS, transcription start site; TTS, transcription termination site.
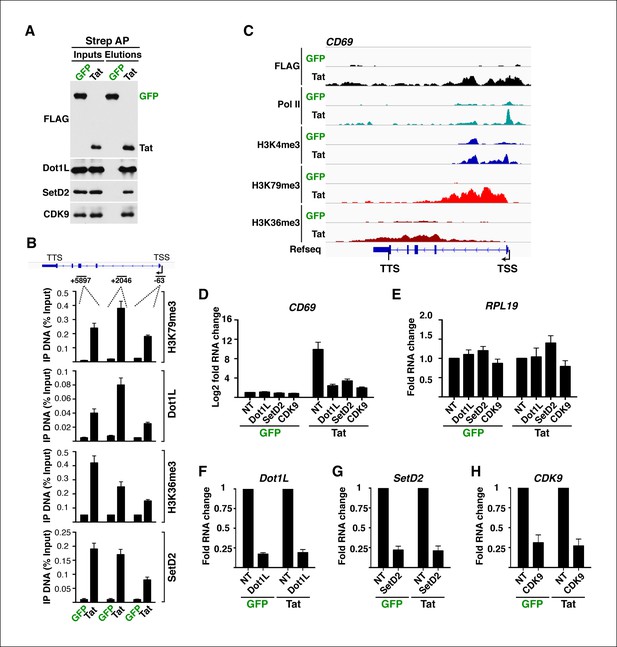
Tat recruits chromatin-modifying enzymes and elongation factors at selected target genes to promote transcription elongation.
(A) Strep-tagged GFP and Tat were affinity purified from nuclear preparations of the Jurkat cell lines (1 × 109 total cells) and interacting partners were analyzed by western blot with the indicated antibodies. CDK9 was used as a positive protein interacting control. (B) ChIP assay to analyze the distribution of histone marks (H3K79me3 and H3K36me3) and chromatin-modifying enzymes (Dot1L and SetD2) at the CD69 locus of the GFP (green) and Tat (black) cell lines. The position of the amplicons used in ChIP-qPCR is shown with the schematic of the locus. Values represent the average of three independent experiments (mean ± SEM; n = 3). (C) Genome browsers of FLAG, Pol II, H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the CD69 locus. The position of the TSS is indicated with an arrow. (D) Knockdown of Dot1L, SetD2 and CDK9 impairs Tat-mediated CD69 transcription activation. qRT-PCR of CD69 in the Jurkat-GFP and -Tat cell lines expressing non-target (NT), Dot1L, SetD2 and CDK9 shRNAs (mean ± SEM; n = 3). (E) Knockdown of Dot1L, SetD2 and CDK9 does not alter RNA steady state levels of RPL19. qRT-PCR of RPL19 in the Jurkat-GFP and -Tat cell lines expressing non-target (NT), Dot1L, SetD2 and CDK9 shRNAs (mean ± SEM; n = 3). (F,G,H) qRT-PCR validation of Dot1L, SetD2 and CDK9 knockdown in the Jurkat-GFP and -Tat cell lines using gene-specific primers and normalized to RPL19 (mean ± SEM; n = 3). ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; NT, non-target; qRT-PCR, quantitative real-time polymerase chain reaction; SEM, standard error of the mean; shRNA, small hairpin RNA; TSG, Tat stimulated genes.
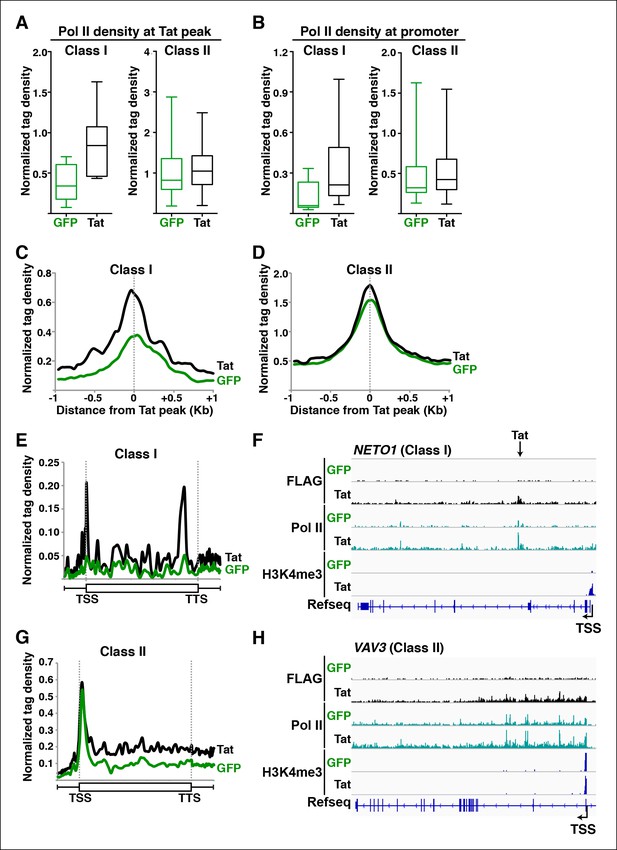
Tat promotes Pol II recruitment and pause release at two distinct TSG classes.
(A) Tat binding promotes Pol II recruitment at class I but not class II TSG. Box plots of normalized Pol II tag density at the Tat peak of class I (n = 17) or II (n = 43) TSG in the GFP and Tat cell lines. (B) Tat binding at class I TSG induces Pol II occupancy at promoters. Box plots of normalized Pol II tag density at the promoter of class I or II TSG in the GFP and Tat cell lines. (C) Pol II normalized tag density relative to the Tat peak at class I TSG. (D) Pol II normalized tag density relative to the Tat peak at class II TSG. (E) Pol II distribution at class I TSG (Metagene plots) in the Tat and GFP cell lines. (F) Genome browser views of ChIP-seq data in the Tat and GFP cell lines at a class I TSG (NETO1). The arrow indicates the position of the FLAG peak called in the Tat cell lines by MACS. (G) Pol II distribution at class II TSG (Metagene plots) in the Tat and GFP cell lines. (H) Genome browser views of ChIP-seq data in the Tat and GFP cell lines at a class II TSG (VAV3). This figure is associated with Figure 3—figure supplements 1,2. ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; MACS, model-based analysis of ChIP-seq; Pol, polymerase; TSG, Tat stimulated genes.
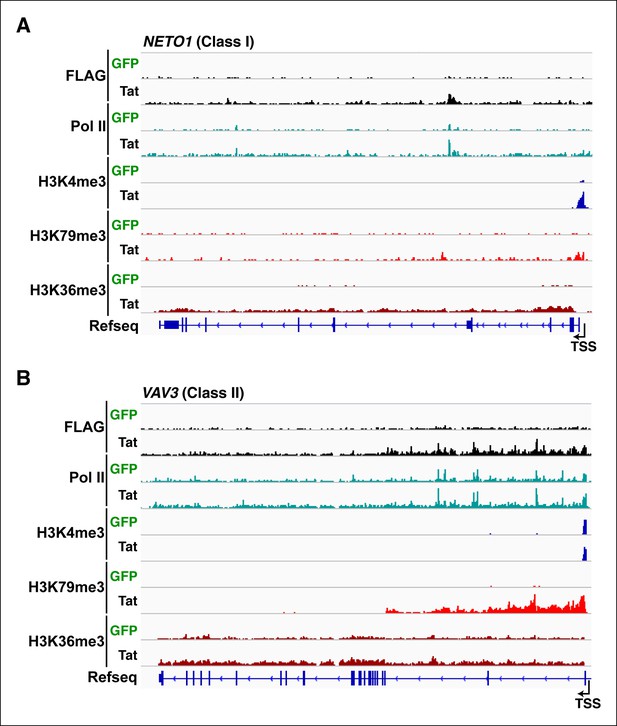
Tat recruitment induces Pol II and chromatin signatures controlling transcription initiation or elongation at different gene classes.
(A) Genome browser views of FLAG, Pol II, H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the NETO1 locus (class I TSG). The position of the TSS is indicated with an arrow. (B) Genome browsers of FLAG, Pol II, H3K4me3, H379me3 and H3K36me3 ChIP-seq tracks in the GFP and Tat cell lines along with the Refseq track for the VAV3 locus (class II TSG). The position of the TSS is indicated with an arrow. GFP, green fluorescent protein; Pol, polymerase; TSG, Tat stimulated genes; TSS, transcription start site.
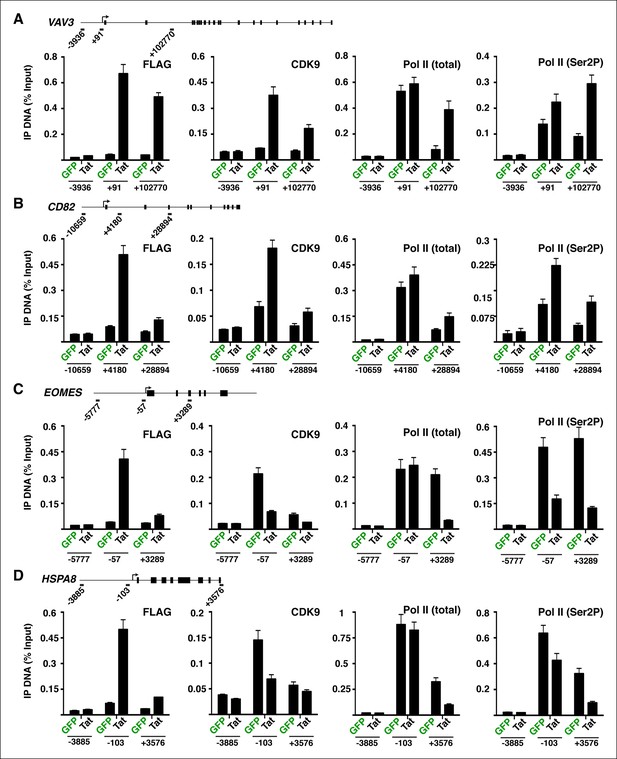
Tat controls P-TEFb and Pol II recruitment at class II TSG and class II TDG.
(A–B) Tat promotes P-TEFb recruitment and Pol II elongation at class II TSG. (C–D) Tat precludes P-TEFb recruitment to block Pol II elongation at class II TDG. Jurkat-GFP and -Tat cell lines were used in ChIP assays to analyze the occupancy of GFP and Tat (FLAG), P-TEFb and Pol II at the indicated target genes using the indicated amplicons. Values represent the average of three independent experiments (mean ± SEM; n = 3). ChIP, chromatin immunoprecipitation; GFP, green fluorescent protein; Pol, polymerase; P-TEFb, positive transcription elongation factor b; SEM, standard error of the mean; TDG, Tat down-regulated genes.
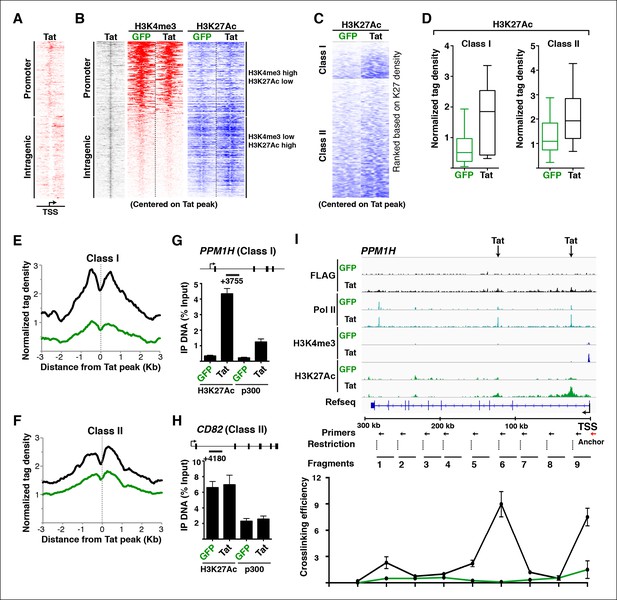
Tat induces transcription initiation from distal sites by inducing gene looping.
(A) Heatmap representation of Tat distribution at TSG promoter or intragenic sites. (B) Heatmap representation of H3K4me3 and H3K27Ac tag density centered on the Tat peak at both TSG promoter and intragenic sites in the GFP and Tat cell lines. Promoter sites are marked with high H3K4me3 and low H3K27Ac levels, while intragenic sites are marked by low H3K4me3 and high H3K27Ac levels. (C) Heatmap representation of H3K27Ac at class I and class II TSG centered on the Tat peak in the GFP and Tat cell lines. Genes are ranked based on H3K27Ac density. (D) Tat binding sharply increases H3K27Ac levels at class I TSG. Box plots showing normalized H3K27Ac density at class I and II TSG in the GFP and Tat cell lines. (E) Metagene analysis of H3K27Ac levels surrounding Tat peaks at class I TSG in the GFP and Tat cell lines. (F) Metagene analysis of H3K27Ac levels surrounding Tat peaks at class II TSG in the GFP and Tat cell lines. (G) ChIP of H3K27Ac and p300 recruitment at an intragenic Tat site at the PPM1H gene. (H) ChIP of H3K27Ac and p300 recruitment at an intragenic Tat site at the CD82 gene. (I) Top, genome browser views of ChIP-seq at the PPM1H locus in the GFP and Tat cell lines. The arrows indicate the position of two intragenic Tat binding sites. Bottom, 3C assay showing the relative crosslinking efficiency at the PPM1H locus in the GFP and Tat cell lines. The position of the primers used in qPCR assays, restriction sites and fragment generated after digestion are indicated. This figure is associated with Figure 4—figure supplements 1–4. 3C, chromosome conformation capture; ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; RNA-seq, RNA sequencing; TSG, Tat stimulated genes.
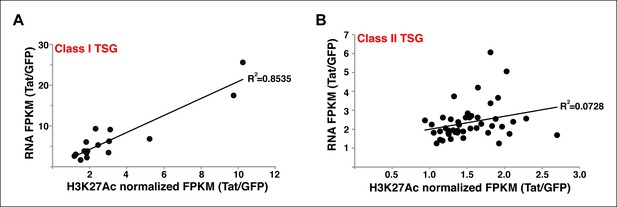
Stimulation of gene expression by Tat correlates with increased H3K27Ac density at the Tat peak.
(A) Correlation plot between normalized H3K27Ac (-/+ 1 kb respective to the Tat peak) and total gene expression levels (based on RNA-seq FPKM) at class I TSG in the presence and absence of Tat (Tat/GFP). (B) Correlation plot between normalized H3K27Ac (-/+ 1 kb respective to the Tat peak) and total gene expression levels (based on RNA-seq FPKM) at class II TSG in the presence and absence of Tat (Tat/GFP). FPKM, Fragments Per Kilobase of transcript per Million mapped reads; GFP, green fluorescent protein; RNA-seq, RNA sequencing; TSG, Tat stimulated genes.
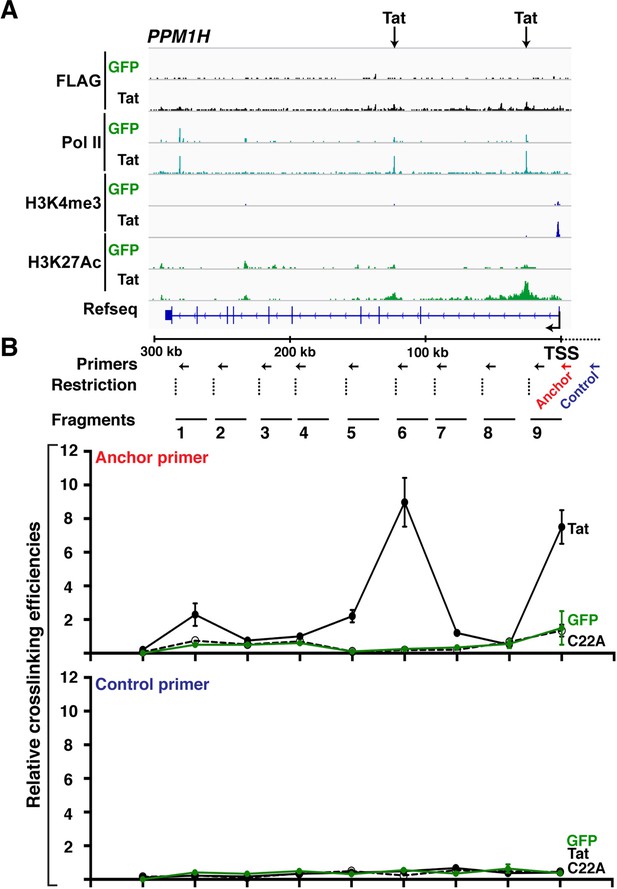
Wild-type Tat but not the C22A non-functional mutant induces gene looping between the promoter and intragenic sites.
(A) Genome browser views of ChIP-seq at the PPM1H locus in the GFP and Tat cell lines. The arrows indicate the position of two intragenic Tat binding sites. (B) 3C assay showing the relative crosslinking efficiency between a promoter anchor primer (top) or an upstream control primer (bottom) and several distal sites at the PPM1H locus in the GFP and Tat cell lines. The position of the primers used in qPCR assays, restriction sites and fragment generated after digestion are indicated (mean ± SEM; n = 3). 3C, chromosome conformation capture; ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; qPCR, quantitative polymerase chain reaction; SEM, standard error of the mean.
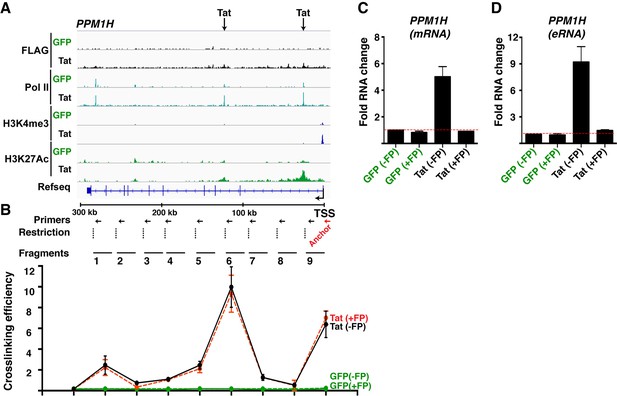
Tat-mediated gene looping does not require transcription activity at enhancers.
(A) Genome browser views of ChIP-seq at the PPM1H locus in the GFP and Tat cell lines. The arrows indicate the position of two intragenic Tat binding sites. (B) 3C assay showing the relative crosslinking efficiency between a promoter anchor primer and intragenic sites at the PPM1H locus in the GFP and Tat cell lines treated with DMSO (± flavopiridol [FP]). The position of the primers used in qPCR assays, restriction sites and fragment generated after digestion are indicated. (C) The GFP and Tat cell lines were treated with DMSO (± flavopiridol [FP]), and RNA isolated to measure levels of PPM1H mRNAs. (D) The GFP and Tat cell lines were treated with DMSO (-FP) or FP (+FP), and RNA isolated to measure levels of the intragenic eRNA (mean ± SEM; n = 3). 3C, chromosome conformation capture; ChIP-seq, chromatin immunoprecipitation sequencing; DMSO, dimethyl sulfoxide; eRNA, enhancer-derived RNA; GFP, green fluorescent protein; qPCR, quantitative polymerase chain reaction; SEM, standard error of the mean.
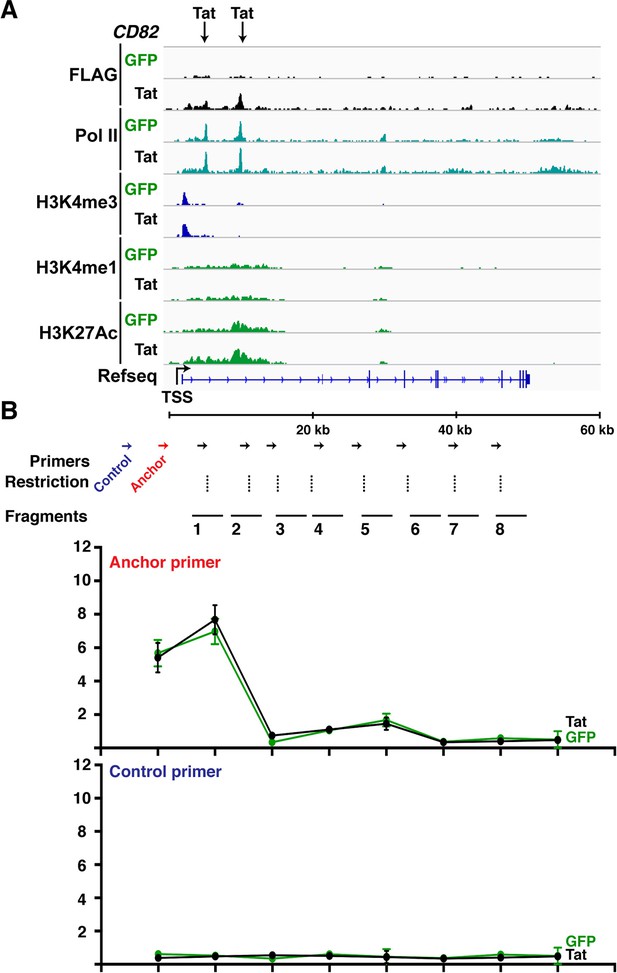
Class II TSG contain stable gene loops between promoter-enhancer that remain unaltered in response of Tat.
(A) Genome browser views of ChIP-seq at the CD82 locus in the GFP and Tat cell lines. The arrows indicate the position of two intragenic Tat binding sites. (B) 3C assay showing the relative crosslinking efficiency at the CD82 locus in the GFP and Tat cell lines using a promoter anchor primer (top) or an upstream control primer (bottom). The position of the primers used in qPCR assays, restriction sites and fragment generated after digestion are indicated (mean ± SEM; n = 3). 3C, chromosome conformation capture; ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; qPCR, quantitative polymerase chain reaction; SEM, standard error of the mean.
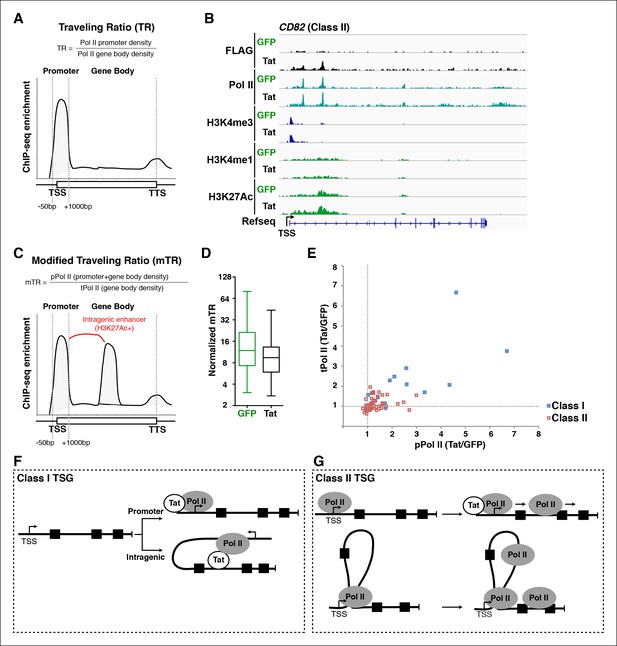
A modified traveling ratio algorithm reveals Tat roles in transcription elongation.
(A) TR algorithm used to calculate levels of promoter-proximal paused and elongation Pol II as previously described (Rahl et al., 2010; Zeitlinger et al., 2007). (B) Genome browser views of ChIP-seq data at the CD82 locus in the GFP and Tat cell lines. (C) mTR algorithm that enables the identification of intragenically paused Pol II at sites of high H3K27Ac levels. (D) mTR decreases at TSG in the Tat cell line. Box plots showing mTR calculations at TSG as indicated in panel (C). (E) Dual-axis plot of the ratio of paused Pol II (pPol II) and traveling Pol II (tPol II) at class I (n = 17) and II (n = 43) TSG in the presence and absence of Tat (Tat/GFP). (F) Model of Tat-mediated Pol II recruitment and gene looping at class I TSG. Tat binds at promoter and/or intragenic regions (in some cases inducing gene looping), and recruits Pol II to promote transcription initiation from both promoter-proximal and promoter-distal sites. (G) Model of Tat-mediated Pol II elongation at class II TSG. Class II TSG are already activated in the absence of Tat, and Tat binds to these genes to primarily stimulate transcription elongation. These genes contain promoter-bound Pol II. This figure is associated with Figure 5—figure supplement 1. GFP, green fluorescent protein; Pol, polymerase; mTR, modified TR; TR, traveling ratio; TSG, Tat stimulated genes.
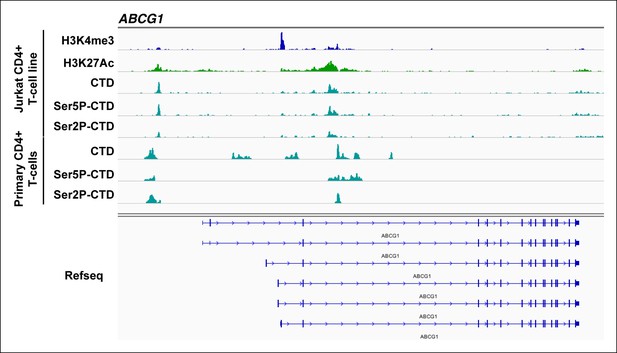
Intragenically paused Pol II is detected in CD4+ T cell lines and primary cells at sites with high H3K27Ac and low H3K4me3.
Genome browser views of H3K4me3, H3K27Ac and different Pol II forms (total CTD, Ser5P-CTD and Ser2P-CTD) in the Jurkat-GFP cell line and primary CD4+ T cells along with the Refseq track for the ABCG1 locus (class II TSG). The position of the different Pol II forms is indicated. GFP, green fluorescent protein; TSG, Tat stimulated genes.
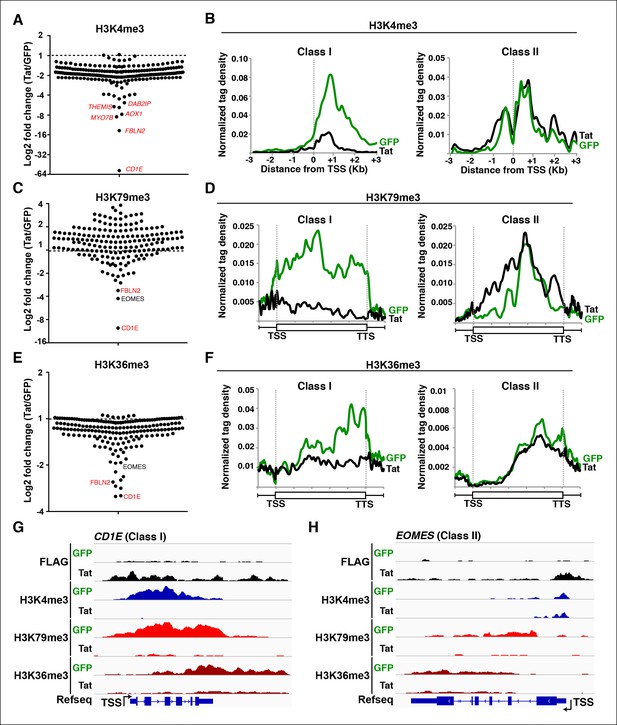
Global analysis of chromatin signatures reveals the basis of transcriptional repression by Tat.
(A) Dot plots of H3K4me3 log2 fold change in the region encompassing -3/+3 Kb from the TSS of all TDG. TDG are divided into two classes: class I (n = 11) and II (n = 14) based on decreased or unchanged levels of H3K4me3 in the presence of Tat, respectively. Selected TDG are indicated in red. (B) Metagene plots centered on TSS showing H3K4me3 occupancy profiles at both class I and class II TDG in the presence of Tat or GFP. (C) Dot plots of H3K79me3 log2 fold change from TSS to TTS. (D) Metagene plots showing average H3K79me3 ChIP-seq signals at both class I and class II TDG in the presence of Tat or GFP (see Materials and methods). (E) Dot plots of H3K36me3 log2 fold change in the region from TSS to TTS. (F) Metagene plots showing average H3K36me3 ChIP-seq signals at both class I and class II TDG in the presence of Tat or GFP. (G) Genome browser views showing ChIP-seq signal at a class I TDG (CD1E) in the GFP and Tat cell lines. (H) Genome browser views showing ChIP-seq signal at a class II TDG (EOMES) in the GFP and Tat cell lines. This figure is associated with Figure 6—figure supplement 1. ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; TDG, Tat downregulated genes; TSS, transcription start site; TTS, transcription termination site.
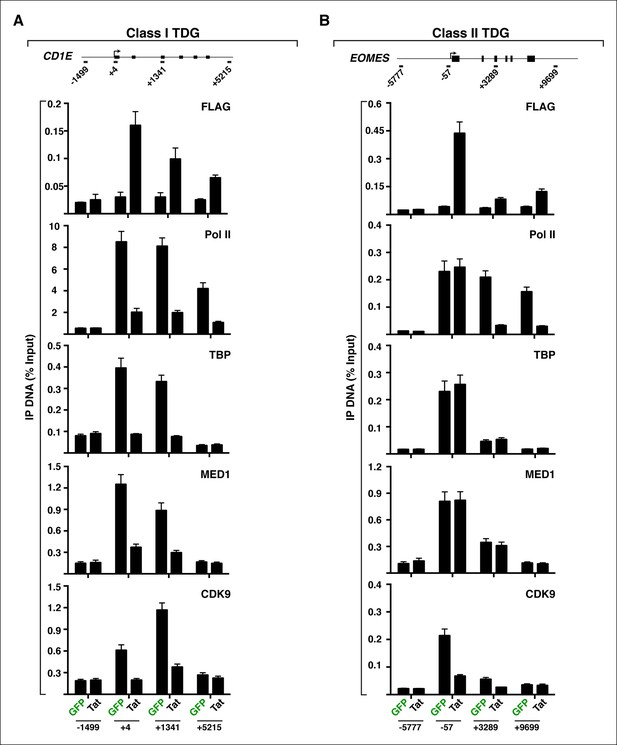
Different modes of Tat repression at class I and II TDG.
(A) Tat blocks transcription initiation of CD1E by interfering with assembly of the pre-initiation complex at the promoter. (B) Tat prevents elongation of EOMES by precluding P-TEFb loading at the promoter. ChIP-qPCR experiments with the indicated antibodies, in the GFP and Tat cell lines. The position of the amplicons used in ChIP-qPCR and their distance to the TSS (arrow) is indicated. Values represent the average of three independent experiments. The SEM is less than 10% and not shown for simplicity. ChIP-qPCR, chromatin immunoprecipitation-quantitative polymerase chain reaction; GFP, green fluorescent protein; SEM, standard error of the mean; TDG, Tat downregulated genes.
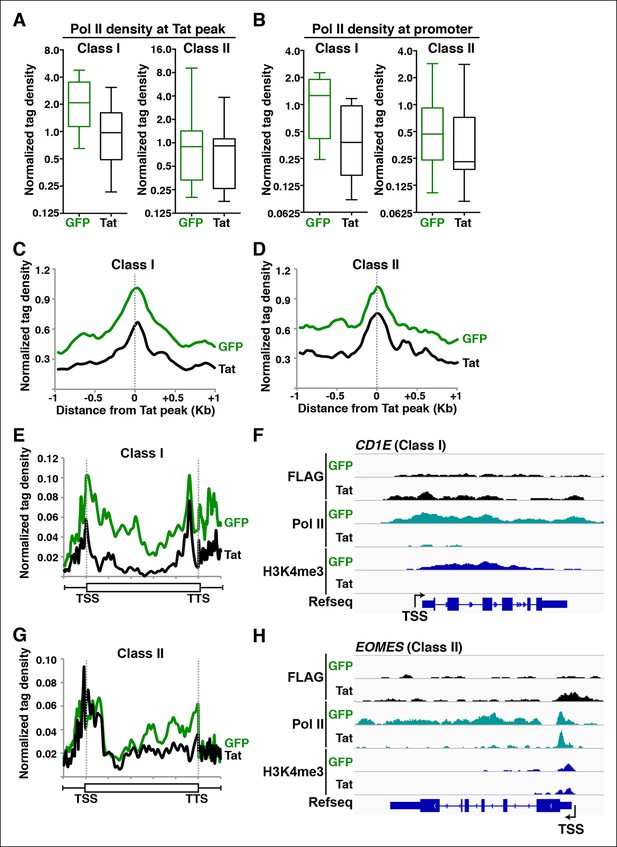
Tat blocks Pol II recruitment and pause release to repress gene transcription at different gene classes.
(A) Tat binding blocks Pol II recruitment primarily at class I TDG. Box plots of normalized Pol II tag density at the Tat peak of class I or II TDG in the GFP and Tat cell lines. (B) Tat binding at class I TDG blocks Pol II density at promoters. Box plots of normalized Pol II tag density at the promoter of class I (n = 11) or class II (n = 14) TDG in the GFP and Tat cell lines. (C) Normalized Pol II tag density relative to the Tat peak at class I TDG. (D) Normalized Pol II tag density relative to the Tat peak at class II TDG. (E) Pol II distribution at class I TDG (Metagene plots) in the Tat and GFP cell lines. (F) Genome browser views of ChIP-seq data at a class I TDG (CD1E) in the Tat and GFP cell lines. (G) Pol II distribution at class II TDG (Metagene plots) in the Tat and GFP. (H) Genome browser views of ChIP-seq data at a class II TDG (EOMES) in the Tat and GFP cell lines. This figure is associated with Figure 7—figure supplement 1. ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; Pol, polymerase; TDG, Tat downregulated genes.
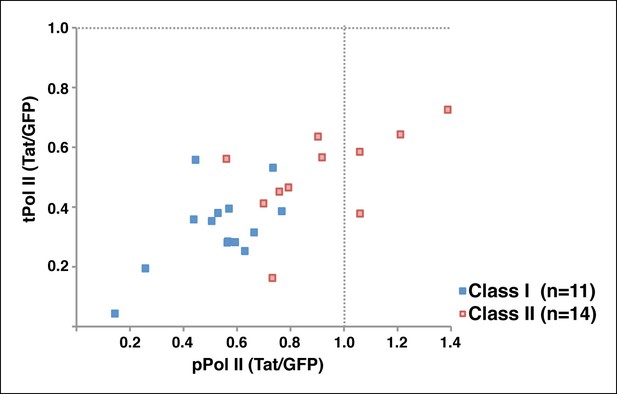
mTR reveals that Tat blocks both Pol II recruitment and pause release at TDG.
Calculations of the ratio of paused Pol II (pPol II) and traveling Pol II (tPol II) at class I and class II TDG in the presence and absence of Tat (Tat/GFP). GFP, green fluorescent protein; mTR, modified traveling ratio; Pol, polymerase; TDG, Tat downregulated genes.
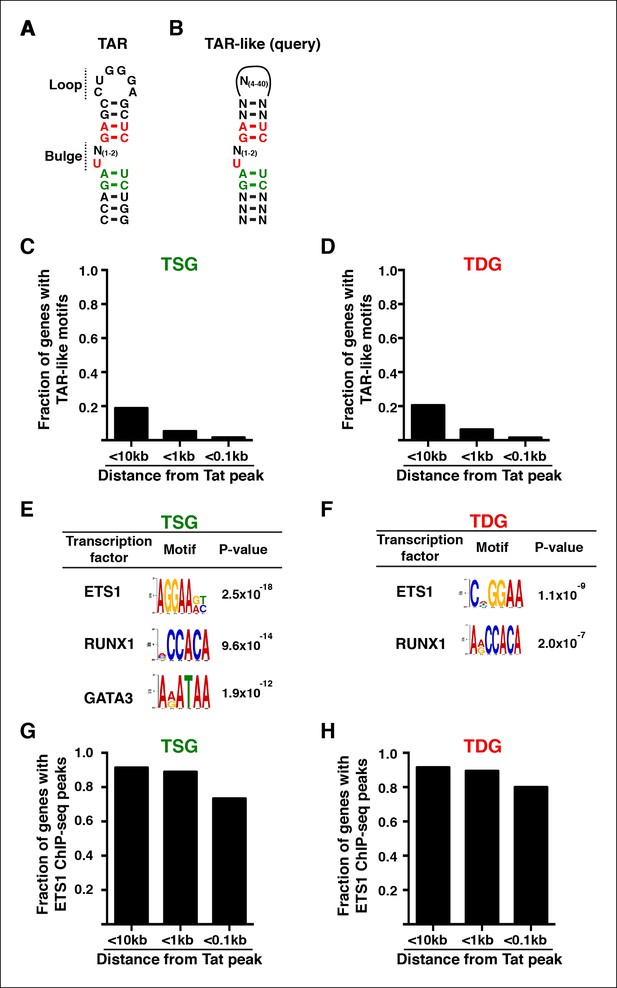
Enrichment of master transcriptional regulators, but not TAR-like, motifs on Tat sites at the direct target genes.
(A) TAR sequence and scheme of the secondary structure. (B) Schematic of the sequence query used to search for TAR-like motifs within the direct Tat target genes. (C) TAR-like motifs are not found at or significantly near Tat peaks in TSG. Fraction of TSG containing TAR-like motifs at <10 kb, <1 kb or <0.1 kb from the Tat peak. (D) TAR-like motifs are not found at or significantly near Tat peaks in TDG. Fraction of TDG containing TAR-like motifs at <10 kb, <1 kb or <0.1 kb from the Tat peak. (E) MEME analysis of 200-bp windows surrounding Tat peaks in TSG reveals high-confidence motifs related to transcription factors ETS1 and RUNX1. (F) MEME analysis of 200-bp windows surrounding Tat peaks in TDG reveal different motifs for ETS1 and RUNX1. (G) Comparison of ETS1 ChIP-seq peak locations in Jurkat, as determined by Hollenhorst et al. (Hollenhorst et al., 2009), to Tat peak locations reveal that 73% of TSG contain an ETS1 peak within 100-bp of a Tat peak. (H) Comparison of ETS1 ChIP-seq peak locations to Tat peak locations reveal that 80% of TDG contain an ETS1 peak within 100-bp of a Tat peak. This figure is associated with Figure 8—figure supplement 1. ChIP-seq, chromatin immunoprecipitation sequencing; TAR, trans-activating RNA; TDG, Tat downregulated genes; TSG, Tat stimulated genes.
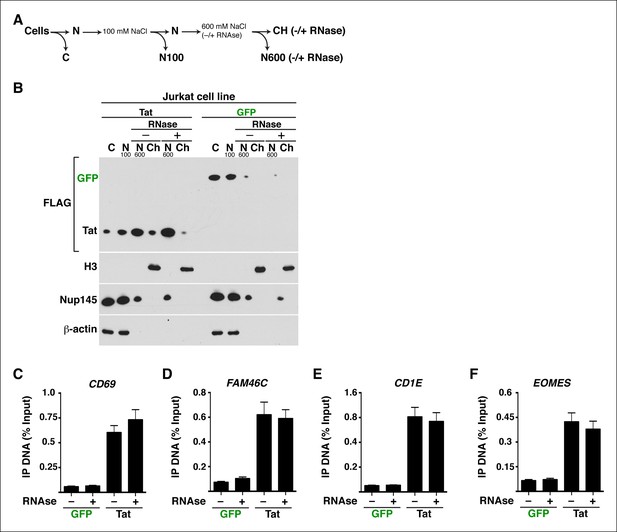
The interaction between Tat and host cell chromatin appears to be primarily dictated by protein–protein interactions.
(A) Fractionation scheme of Jurkat-GFP and Jurkat-Tat cells by increased salt extraction in the absence (–) or presence (+) of RNase A. Ch denotes the chromatin fraction. (B) Western blots of the samples prepared as in panel (A) with the indicated antibodies. The efficiency of RNase treatment was verified by electrophoresis of the purified RNA in an agarose gel stained with ethidium bromide. (C) ChIP assays to analyze the occupancy of GFP and Tat (FLAG) at the CD69 promoter (-63 amplicon) in the absence (–) and presence (+) of RNase. (D) ChIP assays to analyze the occupancy of GFP and Tat (FLAG) at the FAM46C promoter (-182 amplicon) in the absence (–) and presence (+) of RNase. (E) ChIP assays to analyze the occupancy of GFP and Tat (FLAG) at the CD1E promoter (+4 amplicon) in the absence (–) and presence (+) of RNase. (F) ChIP assays to analyze the occupancy of GFP and Tat (FLAG) at the EOMES promoter (-57 amplicon) in the absence (–) and presence (+) of RNase (mean ± SEM; n = 3). ChIP, chromatin immunoprecipitation; GFP, green fluorescent protein; SEM, standard error of the mean.
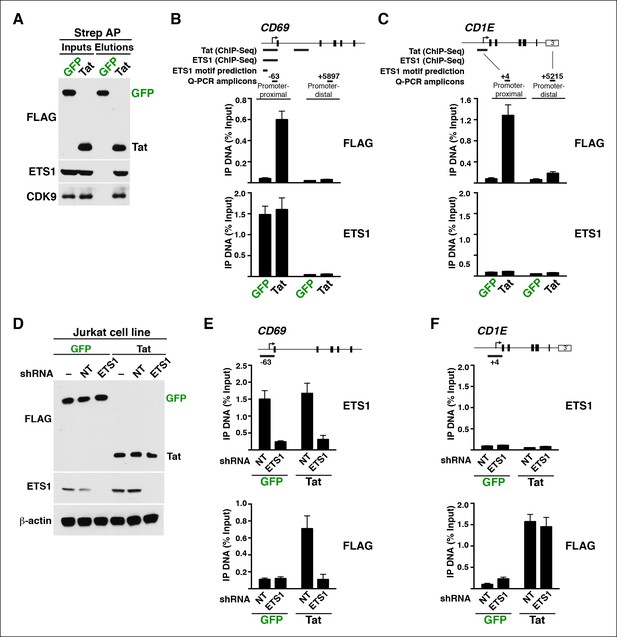
Tat is recruited to its target genes through interaction with the master transcriptional regulator ETS1.
(A) Western blots showing interactions between Tat and ETS1. CDK9 was used as a positive control in the interaction. Strep-tagged Tat and GFP were AP from the Jurkat cell lines using Strep beads, and analyzed by western blot using the indicated antibodies. (B) Tat is recruited to target genes marked by ETS1. ChIP assays showing that Tat and ETS1 co-occupy the promoter-proximal but not promoter-distal region of CD69, in agreement with the location of Tat and ETS1 peaks found by ChIP-seq and the presence of ETS1 motifs (as revealed by enrichment analysis). (C) Tat is also recruited to target genes using ETS1-independent mechanisms. ChIP assays showing that Tat but not ETS1 occupies the promoter-proximal region of the CD1E target gene, in agreement with the Tat and ETS1 ChIP-seq dataset and motif prediction analysis. (D) Generation of Jurkat-GFP and -Tat cell lines expressing non-target (NT) or ETS1 shRNAs. (–) denotes the parental untransduced Jurkat cell lines. Protein lysates were analyzed by western blot using the indicated antibodies to verify for the RNAi efficiency. (E) ETS1 knockdown impairs Tat recruitment to the CD69 promoter. ChIP-qPCR assays showing the density of ETS1 or FLAG at the CD69 promoter in the Jurkat-GFP or -Tat cell lines transduced with the NT or ETS1 shRNAs. (F) ETS1 knockdown does not abolish Tat recruitment to the CD1E promoter. ChIP-qPCR assays showing the density of ETS1 or FLAG at the CD1E promoter in the Jurkat-GFP or -Tat cell lines transduced with the NT or ETS1 shRNAs. This figure is associated with Figure 9—figure supplement 1. AP, affinity purified; ChIP-seq, chromatin immunoprecipitation sequencing; GFP, green fluorescent protein; NT, non-target; qPCR, quantitative polymerase chain reaction; shRNA, small hairpin RNA.
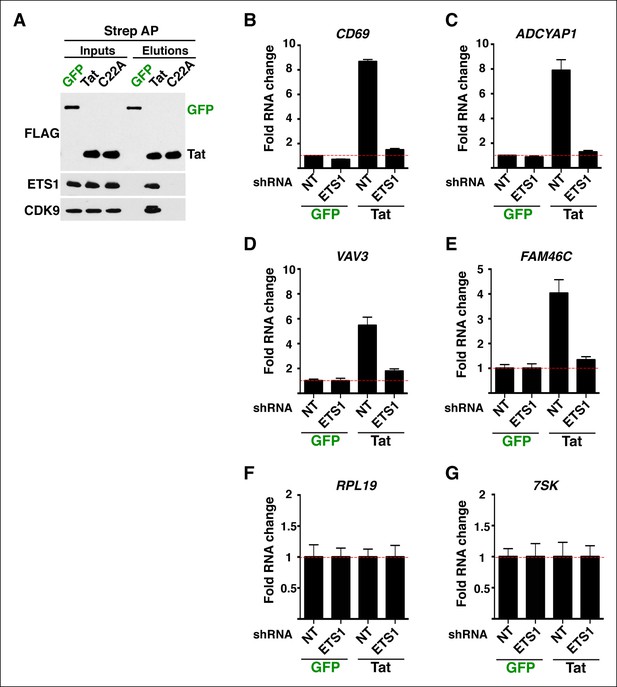
The C22A non-functional Tat mutant fails to bind ETS1 and evidence that ETS1 is critical for transcription activation of Tat target genes.
(A) Western blots showing interactions between Tat (but not GFP or the C22A non-functional Tat mutant) and ETS1. CDK9 was used as a positive control in the interaction. Strep-tagged GFP, Tat, and C22A were AP from the Jurkat-GFP and -Tat cell lines using Strep beads, and analyzed by western blot using the indicated antibodies. (B) qRT-PCR analysis of CD69 expression in the indicated cell lines. (C) qRT-PCR analysis of ADCYAP1 expression in the indicated cell lines. (D) qRT-PCR analysis of VAV3 expression in the indicated cell lines. (E) qRT-PCR analysis of FAM46C expression in the indicated cell lines. (F) qRT-PCR analysis of RPL19 expression in the indicated cell lines. (G) qRT-PCR analysis of 7SK expression in the indicated cell lines. Expression of the indicated genes (panels B–G) was normalized to ACTB. Values represent the average of three independent experiments (mean ± SEM; n = 3). AP, affinity purified; GFP, green fluorescent protein; qRT-PCR, quantitative real-time polymerase chain reaction; SEM, standard error of the mean.
Tables
Genome-wide distribution of Tat in CD4+ T cells.
| Genomic domain | Peaks | Percent | Genome fraction | ChIP (p-value) | |
|---|---|---|---|---|---|
| Total | 6117 | 100 | |||
| Intergenic | 2141 | 35 | 52.7% | ||
| Intragenic | Promoter (–1 kb–TSS) | 1469 | 24 | 1.1% | 3.4 × 10-323 |
| Exon | 128 | 2.1 | 1.9% | 1.4 × 10-3 | |
| Intron | 2227 | 36.4 | 42.4% | 5.1 × 10-4 | |
| 5’ UTR | 73 | 1.2 | 0.4% | 5.3 × 10-193 | |
| 3’ UTR | 79 | 1.3 | 1.5% | 1.4 × 10-2 | |
-
ChIP-seq, chromatin immunoprecipitation sequencing; TSS, transcription start site; UTR, untranslated region.
Genome-wide distribution and function of histone modifications.
| Histone modification | Location | Function |
|---|---|---|
| H3K4me3 | Promoters | Transcription activation |
| H3K79me3 | Gene bodies | Transcription activation |
| H3K36me3 | Gene bodies | Transcription activation |
| H3K4me1 | Enhancers | Does not demarcate active status, but location |
| H3K27Ac | Enhancers | Transcription activation |
| H3K9me3 | Ubiquitous | Transcription repression |
Statistics of the TAR signature selection.
| TSG | TDG | Total | |
|---|---|---|---|
| Targets | 247 | 191 | 438 |
| Genomic sequences (with duplicates) | 1089 | 797 | 1886 |
| Isolated target sequences (with duplicates) | 34739 | 16820 | 51559 |
| With correct secondary structure | 493 | 253 | 746 |
| Corresponding genes | 146 | 86 | 232 |
Additional files
-
Supplementary file 1
Enrichment of DE genes in publicly available HIV infection datasets and functional annotation of canonical pathways at the direct Tat target genes.
- https://doi.org/10.7554/eLife.08955.041
-
Source code 1
Code used to calculate mTR. The mTR algorithm accepts bedgraph files as input and calculates cumulative promoter-proximal paused Pol II occupancy relative to cumulative Pol II occupancy in the gene body after identifying and removing intragenically paused Pol II at sites of high H3K27Ac levels.
- https://doi.org/10.7554/eLife.08955.042





