The anti-tubercular drug delamanid as a potential oral treatment for visceral leishmaniasis
Figures
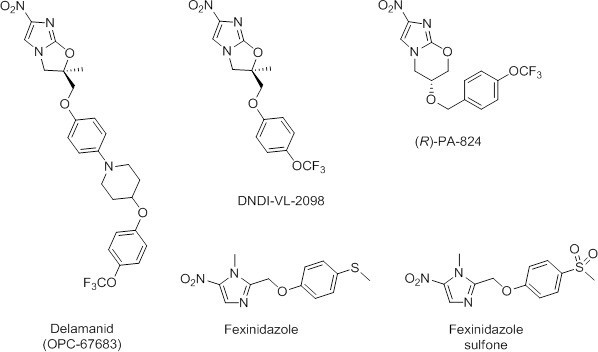
Chemical structures of delamanid ((R)-OPC-67683) and the known anti-leishmanial nitroimidazoles DNDI-VL-2098, (R)-PA-824, fexinidazole and fexinidazole sulfone.
Synthetic schemes for the synthesis of delamanid and analogues are described in Figure 1—figure supplements 1–2.
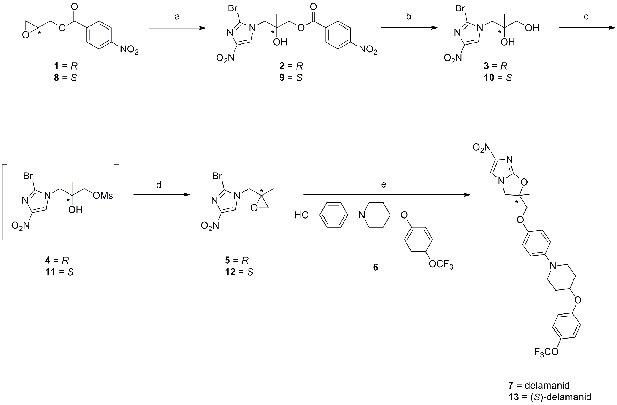
Synthetic route towards delamanid (7) and (S)-delamanid (13).
Reagents and conditions; a) 2-bromo-4-nitro-1H-imidazole, DIPEA, EtOAc, 65°C, 20 hr; b) K2CO3, MeOH, room temp., 16 hr; c) MsCl, pyridine, CH2Cl2, 0°C→room temp., 16 hr; d) DBU, EtOAc, room temp., 16 hr; e) 6, NaH, DMF, 0→50°C, 1.5–4 hr.
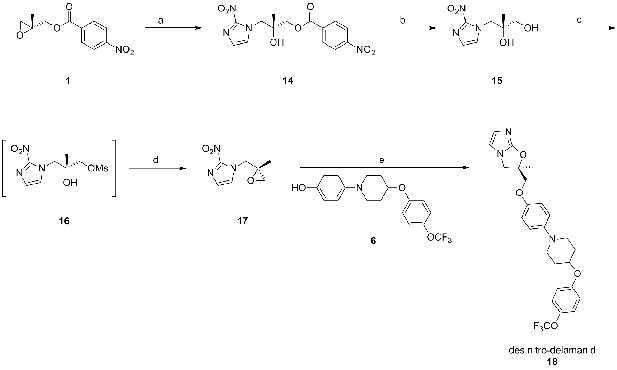
Synthetic route towards des-nitro-delamanid (18).
Reagents and conditions; a) DIPEA, 2-nitroimidazole, EtOAc/MeCN, 77°C, 44 hr, 72%; K2CO3, MeOH, room temp., 20 hr, 83%; MsCl, pyridine, CH2Cl2, 0°C→room temp., 16 hr; DBU, EtOAc, room temp., 16 hr, 15% over 2 steps; NaH, DMF, 0→50°C, 5 h, 30%.
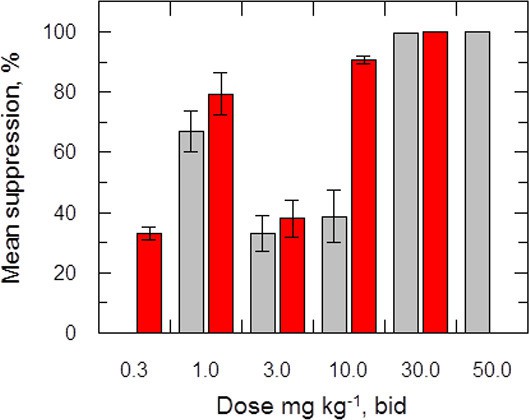
Effects of drug treatment on the parasite burden of mice infected with L. donovani.
Groups of mice (five per group) infected with L. donovani (strain LV9) were dosed with drug vehicle (orally), miltefosine (orally) or delamanid (twice daily, orally) on day 7 post-infection and for a total of 5 or 10 days. Two days after the final dose, animals were humanely euthanized and parasite burdens were determined microscopically by examining Giemsa-stained liver smears. Grey bars, 5-day treatment; red bars, 10 day treatment. This graph shows the combined data from six individual animal studies; n = 9, or 10 for 1, 3 and 10 mg/kg dosing; for all other experiments n = 5. These data are available in tabular form in Figure 2—source data 1.
-
Figure 2—source data 1
Efficacy and PK/PD data from all experiments.
- https://doi.org/10.7554/eLife.09744.008
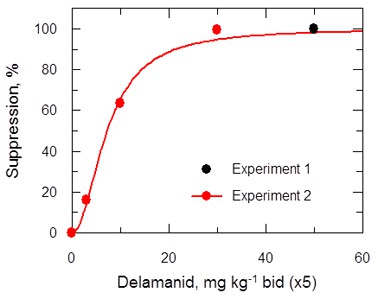
ED50 determination for delamanid in a mouse model of VL.
Results are based on dosing in the range 3 to 50 mg kg-1 b.i.d. oral for 5 days.
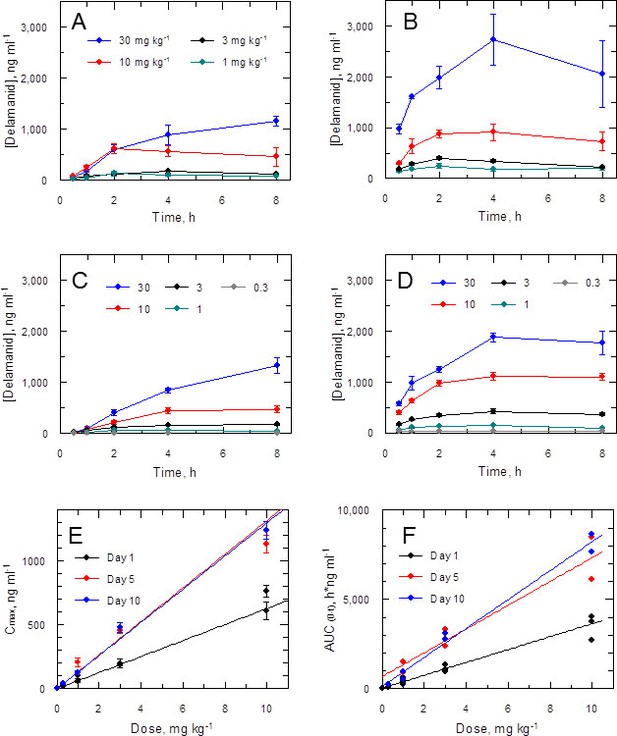
Pharmacokinetic behaviour of delamanid in infected mice.
(A and B) show the blood levels of delamanid following the first oral dose on day 1 (A) and the penultimate oral dose on day 5 (B) for 1, 3, 10 and 30 mg kg-1 b.i.d. (teal, black, red and blue symbols, respectively). Error bars are SEM (n = 3 for 30 mg kg-1, n = 6 all other doses). (C and D) show the blood levels of delamanid from a single VL PK/PD study following the first oral dose on day 1 (C) and the penultimate oral dose on day 10 (D) for 0.3, 1, 3, 10 and 30 mg kg-1 b.i.d. (grey, teal, black, red and blue symbols, respectively). Error bars are SEM (n = 5). (E and F) show the relationship between dose with Cmax or AUC(0-8 h), respectively, after the first oral dose on day 1 (black), or the penultimate dose on day 5 (red) or day 10 (blue). Error bars in (E) are SEM (n ranges from 3–8 depending upon day and dose – see Figure 2 – source data1). Lines in (E) and (F) are best fits by linear regression.
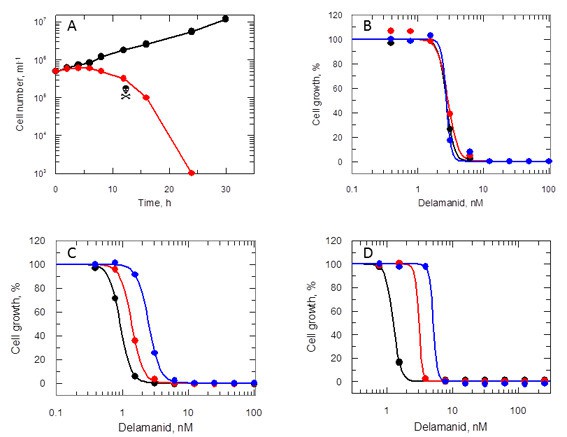
Effects of delamanid on L. donovani promastigotes.
(A) Delamanid causes rapid cell killing. Promastigotes were exposed to delamanid (10 times EC50) and samples removed at intervals to determine cell density and cell viability. Black symbols: no inhibitor; red symbols: plus drug;  the point of irreversible drug toxicity. (B) Drug sensitivity is independent of exposure beyond 24 hr. Black, red and blue symbols are EC50 curves determined after 24, 48 and 72 hr, respectively. (C) Drug sensitivity is cell-density dependent. Black, red and blue symbols are EC50 curves determined after 72 hr, with initial seeding densities of 103, 104 and 105 cells ml-1, respectively. (D) Drug sensitivity is serum dependent. Black, red and blue symbols are EC50 curves determined after 72 hr in the presence of 5, 10 and 20% FCS, respectively.
the point of irreversible drug toxicity. (B) Drug sensitivity is independent of exposure beyond 24 hr. Black, red and blue symbols are EC50 curves determined after 24, 48 and 72 hr, respectively. (C) Drug sensitivity is cell-density dependent. Black, red and blue symbols are EC50 curves determined after 72 hr, with initial seeding densities of 103, 104 and 105 cells ml-1, respectively. (D) Drug sensitivity is serum dependent. Black, red and blue symbols are EC50 curves determined after 72 hr in the presence of 5, 10 and 20% FCS, respectively.
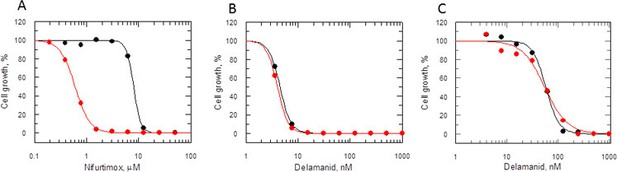
Mode of action of delamanid is distinct from nifurtimox.
(A, B and C) Susceptibility to nifurtimox (A) is increased in NTR-overexpressing promastigotes (red symbols), but not to delamanid (B) compared to WT cells (black symbols). Data are the mean of triplicate cultures from a single experiment. (C) The susceptibility of delamanid is not increased in NTR-overexpressing amastigotes in macrophages (red symbols) compared to WT (black symbols). Data are the mean of duplicate cultures from a single experiment.
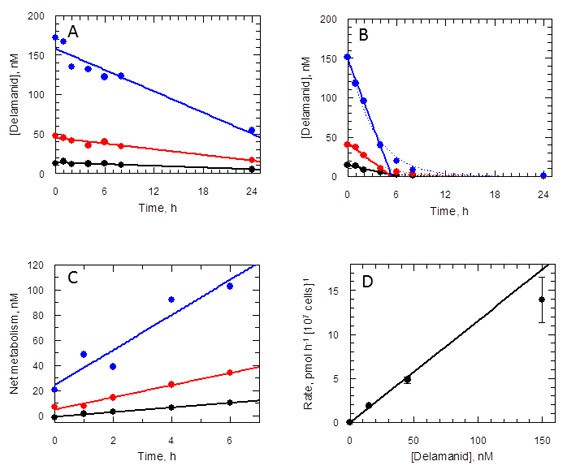
Delamanid metabolism in L. donovani promastigotes.
(A) Medium plus delamanid alone and (B) cells incubated in medium plus delamanid. Delamanid concentrations added are 15, 45 and 150 nM (black, red and blue, respectively). The lines represent best fits by linear regression for all data points in (A) and 0 to 5 hr in (B). The dotted line in (B) is the best fit by non-linear regression to a single exponential decay. (C) Net metabolism of delamanid by cells was obtained by subtraction of (A) from (B). Data fitted by linear regression gave correlation coefficients of 0.996, 0.991 and 0.951 for delamanid concentrations of 15, 45 and 150 nM, respectively. (D) Rates of delamanid metabolism obtained from (C) are linear up to 150 nM (correlation coefficient 0.996, explicit errors used in fit).
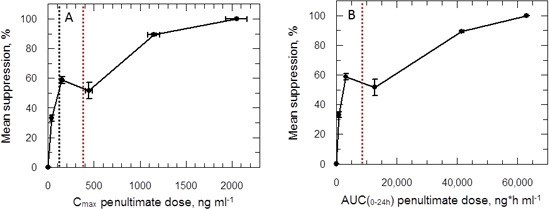
PK/PD relationships in mice.
(A and B) Mean suppression of parasite burden as a function of Cmax for the penultimate dose (panel A) and extrapolated AUC(0–24 hr) for the last day of the 10-day treatment regimen (panel B). The black dotted line in (A) is the EC90 value obtained for infected macrophages after 72 hr exposure (120 ng ml-1). The red dotted line in (A) and (B) represents the mean delamanid Cmax (375–400 ng ml-1) and mean AUC(0–24 hr) (7000–8000 h*ng ml-1) obtained in 144 TB patients after 14 days treatment with 100 mg, oral, once daily from day 14–56 (Sasahara et al., 2015). The data in this graph were derived from a single in vivo study; related aggregated data from previous studies is shown in Figure 7—figure supplement 1.
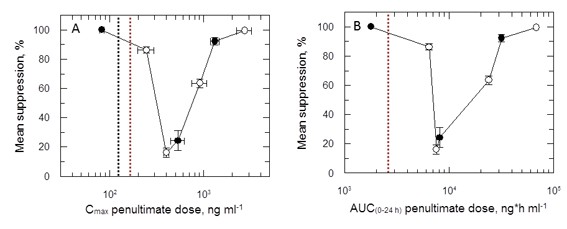
(A and B) Mean suppression of parasite burden as a function of Cmax for the penultimate dose and extrapolated AUC(0–24 hr) for the last day of the 5- and 10-day treatment regimens, respectively.
Open and closed symbols represent data combined from the 5- and 10-day studies, respectively. This graph represents the aggregated data from 3 separate in vivo studies. The black dotted line in (A) is the EC90 value obtained for infected macrophages after 72-hr exposure. The red dotted line in (A) and (B) represents the mean delamanid Cmax (375–400 ng ml-1) and mean AUC(0–24 hr) (7000–8000 h*ng ml-1) obtained in TB patients after 14 days treatment with 100 mg, oral, once daily (Sasahara et al., 2015).
Tables
Activity of delamanid against laboratory and clinical isolates of L. donovani in vitro. EC90 values are calculated from the EC50, Hill slopes and the molecular weight of delamanid.
| Species | Developmental stage | EC50, nM (Hill slope) | EC90, nM | EC90, ng ml-1 |
|---|---|---|---|---|
| Leishmania donovani (LdBOB) | Promastigote | 15.5 ± 0.07 (8.4) | 20.2 | 10.8 |
| Leishmania donovani (LdBOB) | Amastigote (axenic) | 5.4 ± 0.05 (5.3) | 8.2 | 4.4 |
| Leishmania donovani (LV9) | Amastigote (in macrophage) | 86.5 ± 1.7 (2.3) | 225 | 120 |
| Leishmania donovani (DD8) | Amastigote (in macrophage) | 298 ± 13 (2.7) | 672 | 359 |
| Leishmania donovani (BHU1) | Amastigote (in macrophage) | 230 ± 10 (4.1) | 393 | 210 |
| Leishmania donovani (SUKA001) | Amastigote (in macrophage) | 259 ± 7 (3.6) | 476 | 254 |
| L. infantum (ITMAP263) | Amastigote (in macrophage) | 940 ± 0.05 (3.4) | 1790 | 955 |
| Human (HepG2) | N/A | >5000 | - | - |
Leishmania isolates used in delamanid drug sensitivity studies.
| Species | WHO Code | Laboratory Code | Origin | Year of Isolation | Provided by |
|---|---|---|---|---|---|
| L. donovani | MHOM/SD/62/1S CL2D a | LdBOB | Sudan | 1962 | via Professor Stephen Beverley, Washington University |
| L. donovani | MHOM/ET/67/HU3 | LV9 | Ethiopia | 1967 | via Professor Jennie Blackwell, Cambridge University |
| L. donovani | MHOM/IN/02/BHU1 | BHU1 | India | 2002 | via LSHTM, London, UK |
| L. donovani | MHOM/SU/09/SUKA001 | SUKA001 | Sudan | 2009 | via LSHTM, London, UK |
| L. donovani | MHOM/IN/80/DD8 | DD8 | India | 1980 | via LSHTM, London UK |
| L. infantum | MHOM/MA/67/ITMAP263 | ITMAP263 | Morocco | 1967 | via LSHTM, London, UK |
| L. major | MHOM/IL/81/Friedlin | Friedlin | Israel | 1981 | via LSHTM, London, UK |
-
a Derived from this strain (Goyard et al., 2003)





