Host-derived Lactobacillus plantarum alleviates hyperuricemia by improving gut microbial community and hydrolase-mediated degradation of purine nucleosides
Figures
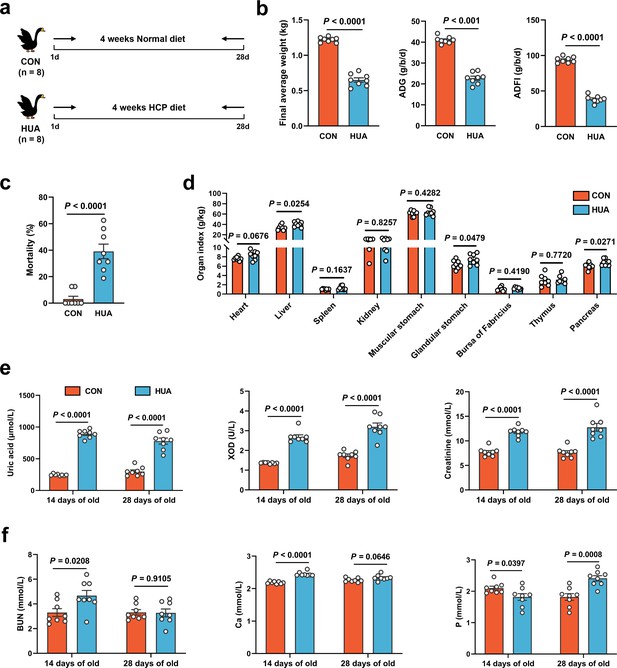
Growth and metabolism differences in diet-dependent hyperuricemia (HUA).
(a) Experimental design. (b) Final average weight, average daily feed intake (ADFI), and average daily gain (ADG) of geese after 28 days of control (CON, n = 8) or HCP diet (HUA, n = 8). (c) Mortality of geese after 28 days of control (CON, n = 8) or HCP diet (HUA, n = 8). (d) The organ index of geese after 28 days of control (CON, n = 8) or HCP diet (HUA, n = 8). (e) The serum uric acid (UA), xanthine oxidase (XOD), and creatinine levels of geese after 14 or 28 days of control (CON, n = 8) or HCP diet (HUA, n = 8). (f) The blood urea nitrogen (BUN), calcium (Ca), and phosphorus (P) levels of geese after 14 or 28 days of control (CON, n = 8) or HCP diet (HUA, n = 8). Statistical significance in (b–f) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m.
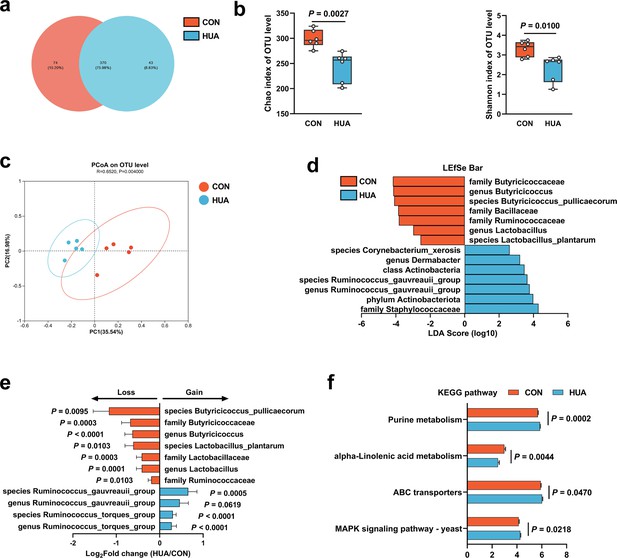
Gut microbiota differences in diet-dependent hyperuricemia (HUA).
(a) Venn plot of bacterial operational taxonomic unit (OTU) in fecal samples of geese fed with control diet (CON, 15.20%) or HCP diet (HUA, 8.83%) after 28 days. (b) Chao and Shannon indices of indicated groups based on alpha diversity analysis (n = 6). (c) Principal component analysis of bacteria on OTU level with 95% confidence regions between CON and HUA groups (n = 5, r = 0.6520, p = 0.0040). (d) Linear discriminative analysis (LDA) score obtained from LDA effect size (LEfSe) analysis of fecal microbiota in CON and HUA groups. Bacterial with Kruskal–Wallis ≤0.05, as well as LDA >2, are reported. (e) The alteration trends of the bacterial relative abundance (n = 5). The x-axis shows the log2 fold change of the bacterial relative abundance in the HUA group compared to the CON group. (f) Summary of major pathway changes based on Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt) prediction analysis of 16S data from fecal samples of CON and HUA groups. Pathways with p ≤ 0.05 by Wilcoxon rank-sum test are reported. Statistical significance in (b) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m.
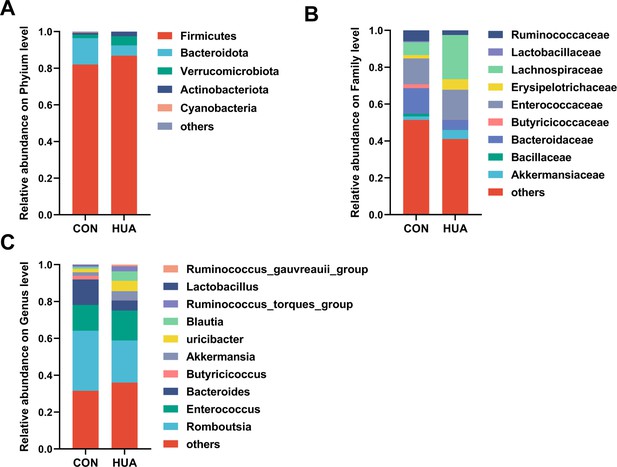
Relative abundance of gut microbes in diet-dependent hyperuricemia (HUA).
Relative abundance of gut microbes at the phylum (A), family (B), and genus (C) levels of goose after 28 days of control (CON, n = 8) or HCP diet (hyperuricemia [HUA], n = 8).
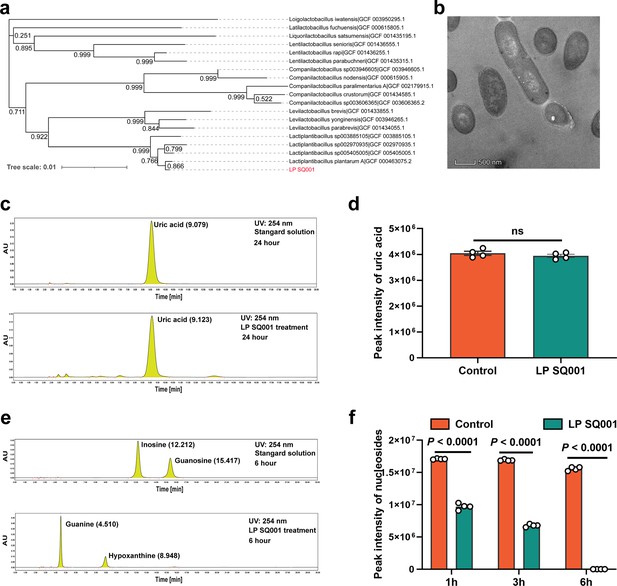
L. plantarum SQ001 can hydrolyze nucleosides in vitro.
(a) The phylogenetic tree of L. plantarum SQ001. 16S ribosomal RNA (rRNA) of L. plantarum SQ001 was extracted to sequence and blast the phylogeny, compared with 18 strains from different origins. Bootstrap values are displayed under the branches. (b) The transmission electron microscope results of the strain L. plantarum SQ001. Magnification of 12,000. (c) Degradation of uric acid by L. plantarum SQ001 from high-performance liquid chromatography (HPLC) analysis. (d) Peak intensity of uric acid by L. plantarum SQ001 after 24 hr treatment (n = 4). (e) Degradation of inosine and guanosine by L. plantarum SQ001 from HPLC analysis. (f) Peak intensity of nucleosides by L. plantarum SQ001 after 1, 3, and 6 hr treatment (inosine and guanosine, n = 4). Statistical significance in (e) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m. LP SQ001, L. plantarum SQ001; LP 23180, L. plantarum 23180.
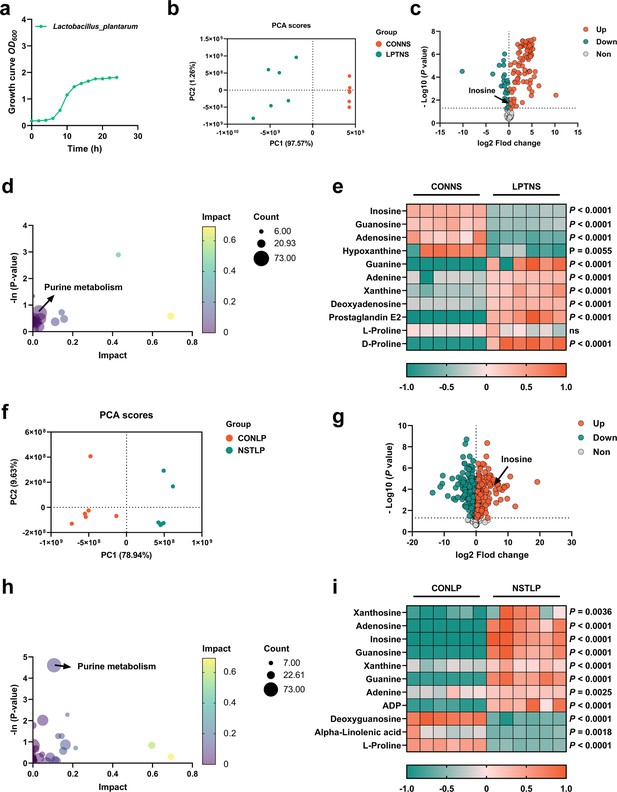
Purine metabolism of L. plantarum SQ001 was activated by nucleosides.
(a) Growth curve of L. plantarum SQ001. (b) Principal component analysis (PCA) score plot of different metabolites collected from nucleoside solution samples with the treatment of LP or CON. All collected data are within the 95% confidence interval. Red represents the CONNS group, green represents the LPTNS group. (c) Volcano plots of changed metabolites for the comparison with LPTNS versus CONNS. (d) Kyoto encyclopedia of genes and genomes (KEGG)-based pathway over-representation analysis of changed metabolites between CONNS and LPTNS groups. The bubble size represents the percentage of significant metabolites contained in each pathway. Bubbles are colored according to the impact. (e) Heatmap of extracellular liquid chromatography–mass spectrometry (LC–MS) data showing marker metabolite changes between CONNS and LPTNS groups (n = 6). Increases in metabolite levels are shown in red, whereas green indicates decreased metabolite. (f) PCA score plot of different metabolites collected from LP samples with the treatment of nucleoside or CON. All collected data are within the 95% confidence interval. Red represents the CONLP group, green represents the NSTLP group. (g) Volcano plots of changed metabolites for the comparison with NSTLP versus CONLP. (h) KEGG-based pathway over-representation analysis of changed metabolites between NSTLP and CONLP groups. The bubble size represents the percentage of significant metabolites contained in each pathway. Bubbles are colored according to the impact. (i) Heatmap of intracellular LC–MS data showing marker metabolite changes between NSTLP and CONLP groups (n = 6). Increases in metabolite levels are shown in red, whereas green indicates decreased metabolite. Statistical significance in (e, i) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m. CONNS: control nucleoside solution; LPTNS: L. plantarum SQ001-treated nucleoside solution; CONLP: control L. plantarum SQ001; NSTLP: nucleoside solution-treated L. plantarum SQ001.
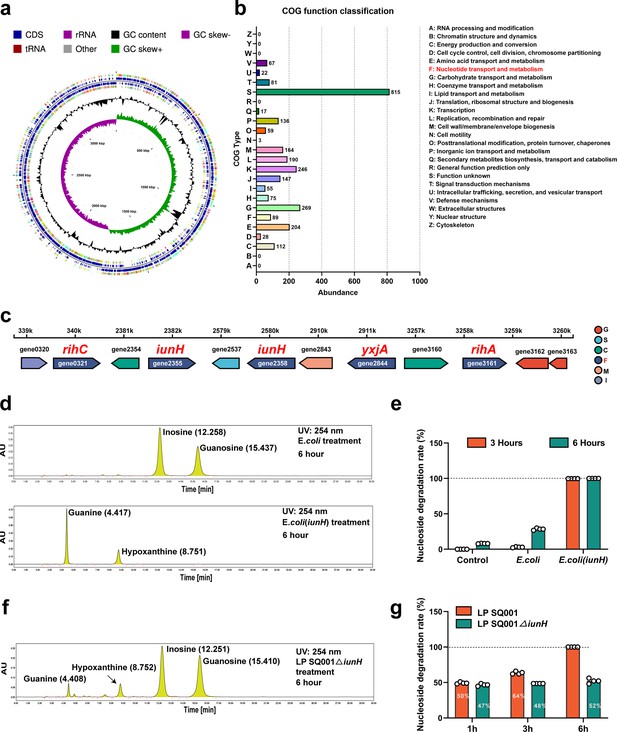
L. plantarum SQ001 degrades nucleosides through nucleoside hydrolase, which has the potential to alleviate hyperuricemia (HUA).
(a) Genome map of L. plantarum SQ001. (b) The clusters of orthologous groups (COG) repertoires of L. plantarum SQ001. (c) Genes involved in nucleotide transporter metabolism are arranged on each scaffold as a linear plot, with each gene labeled for coding direction and color-coded by COG functional classification. (d) Degradation of inosine and guanosine by E. coli and E. coli (iunH) from high-performance liquid chromatography (HPLC) analysis. (e) Nucleoside degradation rate of L. plantarum SQ001 gene iunH after heterologous expression in E. coli. (f) Degradation of inosine and guanosine by L. plantarum SQ001∆iunH from HPLC analysis. (g) Nucleoside degradation rate of L. plantarum SQ001 after knockout of gene iunH. yxjA: nucleoside permease; iunH: nucleoside hydrolase; rihC: ribonucleoside hydrolase; rihA: pyrimidine-specific ribonucleoside hydrolase.
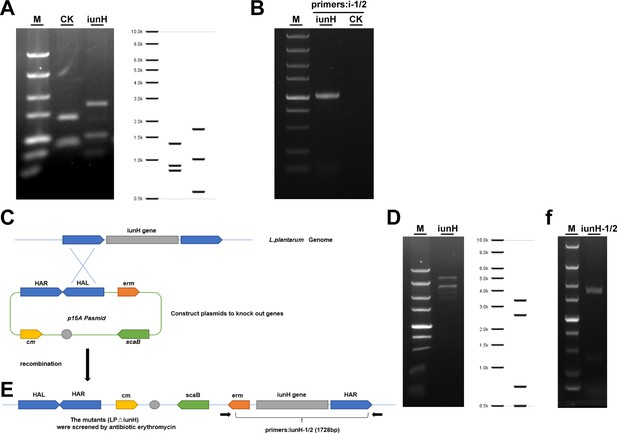
Heterologous expression and knockout validation of the gene iunH.
(A) p15A-cm-Pgenta; p15A-cm-iunH restriction Endonuclease digestion map. M: DL 5000 DNA marker, with stripe sizes ranging from top to bottom from 5, 3, 2, 1.5, 1, 0.75, 0.5, 0.25, and 0.1 kb. (B) PCR verification of the heterologous expression of recombinant plasmids p15A-cm-Pgenta, p15A-cm-iunH. i1: Pgenta-iunH-1, i2: iunH-2, PCR product: 2100 bp (mutants, i1/i2), M: DL5000 DNA ladder, CK: Nissle1917/p15A-cm-Pgenta. (C) Construct plasmids to knock out genes. (D) p15A cm-HA-erm scaB iunH restriction Endonuclease digestion map. M: DL 5000 DNA marker. (E) The recombinant strains were screened for resistance and single colonies were selected for PCR validation to obtain successful recombinant deletion strains. (F) PCR verification of the heterologous expression of recombinant plasmids p15A-cm-HA-erm scaB iunH, iunH-1: check-iunH-1, iunH-2: check-iunH-2, PCR product: 1728 bp (mutants, iunH-1/2), M: DL5000 DNA ladder.
-
Figure 5—figure supplement 1—source data 1
Raw unedited gels for Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/100068/elife-100068-fig5-figsupp1-data1-v1.zip
-
Figure 5—figure supplement 1—source data 2
Uncropped and labeled gels for Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/100068/elife-100068-fig5-figsupp1-data2-v1.zip
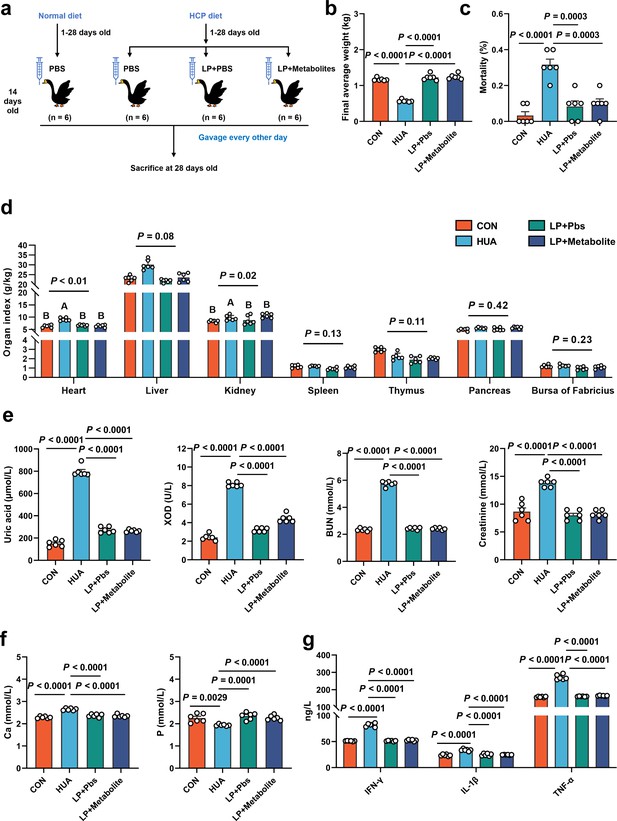
L. plantarum SQ001 and L. plantarum SQ001 with metabolites alleviate HCP diet-induced HUA.
(a) Experimental design. (b) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the final average weight in HCP diet-treated geese (n = 6). (c) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the mortality in HCP diet-treated geese (n = 6). (d) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the organ index in HCP diet-treated geese (n = 6). (e) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the serum uric acid, xanthine oxidase (XOD), blood urea nitrogen (BUN), and creatinine in HCP diet-treated geese (n = 6). (f) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the serum Ca and P levels in HCP diet-treated geese (n = 6). (g) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the serum IFN-γ, IL-1β, and TNF-α levels in HCP diet-treated geese (n = 6). Statistical significance in (b–f) was determined by unpaired two-tailed Student’s t-test. Significant differences in (d) were evaluated by one-way ANOVA with Bonferroni’s multiple comparisons test. Data with error bars represent mean ± s.e.m.
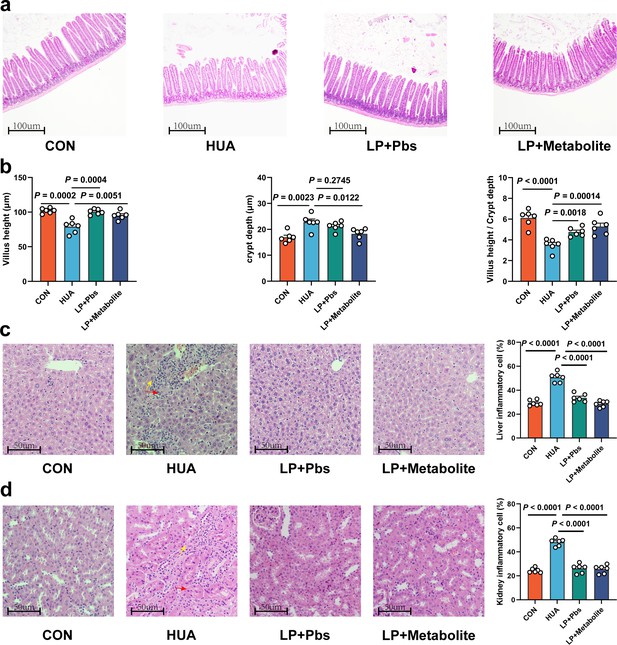
L. plantarum SQ001 and L. plantarum SQ001 with metabolites could alleviate HCP diet-induced damage in the jejunum, liver, and kidneys.
(a) Representative H&E staining images and a histologic score of the jejunum section of geese (n = 6). Scale bar: 100 µm. (b) Villi height, crypt depth, and the value of villi height/crypt depth (n = 6). Six crypts were counted for each section. (c) Representative H&E staining images and inflammatory cell rate of the liver section of geese (n = 6). Scale bar: 50 µm. (d) Representative H&E staining images and inflammatory cell rate of kidney section of geese (n = 6). Scale bar: 50 µm. Statistical significance in b was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m.
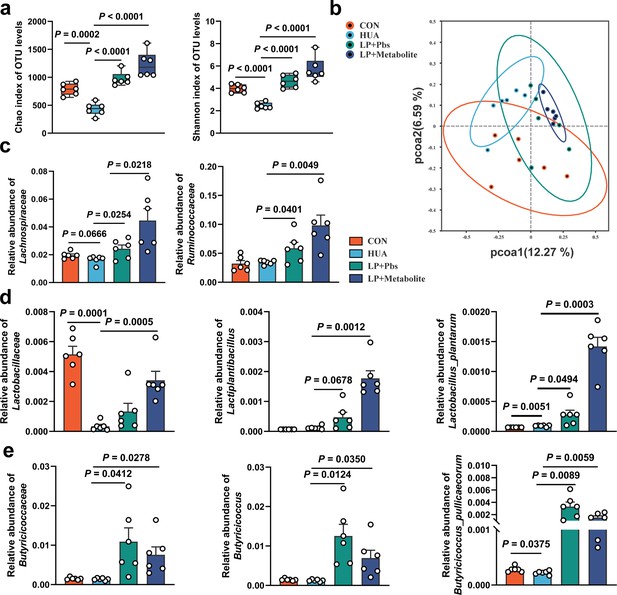
L. plantarum SQ001 and L. plantarum SQ001 with metabolites alleviate HCP diet-induced gut microbial disorder.
(a) Chao and Shannon indices of indicated groups based on alpha diversity analysis (n = 6). (b) Principal component analysis of bacteria with 95% confidence regions of indicated groups (n = 6). (c) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the relative abundance of family Lachnospiraceae and Ruminococcaceae in HCP diet-treated geese (n = 6). (d) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the relative abundance of family Lactobacillaceae, genus Lactiplantibacillus, and species Lactobacillus plantarum in HCP diet-treated geese (n = 6). (e) Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on the relative abundance of family Butyricicoccaceae, genus Butyricicoccus, and species Butyricicoccus pullicaecorum in HCP diet-treated geese (n = 6). Statistical significance in (a, c–e) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m.
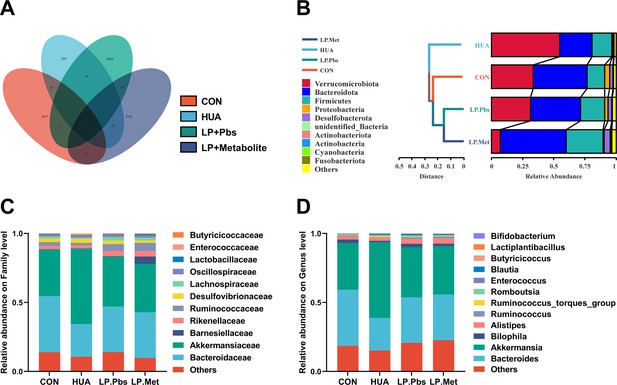
Effect of L. plantarum SQ001 and L. plantarum SQ001 with metabolites on gut microbiota in geese.
(A) Venn plot of bacterial operational taxonomic unit (OTU) in fecal samples of goose with indicated groups. Relative abundance of gut microbes at the phylum (B), family (C), and genus (D) levels of goose with indicated groups.
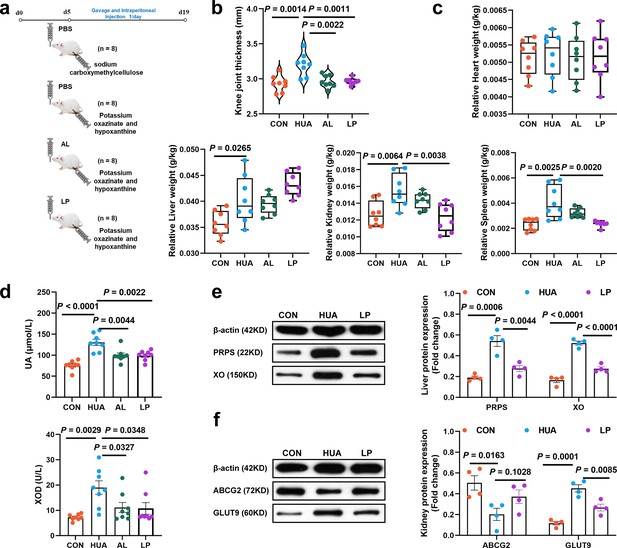
Effect of L. plantarum SQ001 on alleviating HUA and protein expression levels of uric acid (UA) metabolism in mice.
(a) Experimental design. (b) Effect of L. plantarum SQ001 on the knee joint thickness in mice (n = 8). (c) Effect of L. plantarum SQ001 on the organ index (heart, liver, kidney, and spleen) in mice (n = 8). (d) Effect of L. plantarum SQ001 on the serum UA, xanthine oxidase (XOD) in HCP diet-treated geese (n = 8). (e) Representative western blotting images and quantification of proteins (phosphoribosyl pyrophosphate synthetase [PRPS], XO) in the liver tissue between the CON, HUA, and LP groups (n = 4). (f) Representative western blotting images and quantification of proteins (ABCG2, GLUT9) in the kidney tissue between the CON, HUA, and LP groups (n = 4). Statistical significance in (b–f) was determined by unpaired two-tailed Student’s t-test. Data with error bars represent mean ± s.e.m. CON, control group; HUA, hyperuricemia model group; AL, allopurinol group; LP, L. plantarum SQ001 group.
-
Figure 9—source data 1
Raw unedited gels for Figure 9.
- https://cdn.elifesciences.org/articles/100068/elife-100068-fig9-data1-v1.zip
-
Figure 9—source data 2
Uncropped and labeled gels for Figure 9.
- https://cdn.elifesciences.org/articles/100068/elife-100068-fig9-data2-v1.zip
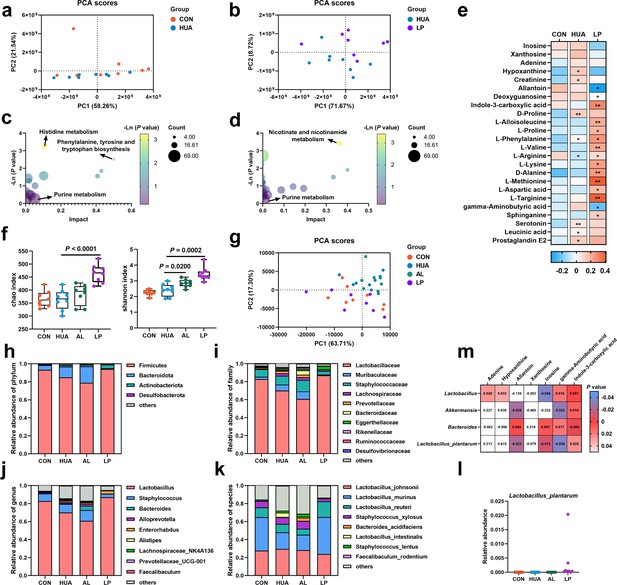
Effect of L. plantarum SQ001 on serum metabolites and gut microbiota in mice.
(a) Principal component analysis of the serum samples. The yellow color represents the CON group (n = 8), while the green color represents HUA group (n = 8). (b) Principal component analysis of the serum samples. The green represents the HUA group (n = 8), while the purple represents LP group (n = 8). (c) Kyoto encyclopedia of genes and genomes (KEGG)-based pathway over-representation analysis of changed metabolites between CON and HUA groups. The bubble size represents the percentage of significant metabolites contained in each pathway. Bubbles are colored according to the impact. (d) KEGG-based pathway over-representation analysis of changed metabolites between HUA and LP groups. The bubble size represents the percentage of significant metabolites contained in each pathway. Bubbles are colored according to the impact. (e) Heatmap of serum metabolites liquid chromatography–mass spectrometry (LC–MS) data showing marker metabolite changes between CON, HUA, and LP groups (n = 8). Increases in metabolite levels are shown in red, whereas blue indicates decreased metabolite. (f) Chao and Shannon indices of indicated groups based on alpha diversity analysis (n = 8). (g) Principal component analysis of bacteria with 95% confidence regions of indicated groups (n = 8). Relative abundance of gut microbes at the phylum (h), family (i), genus (j), and species (k) levels of mice (n = 8). (l) Effect of L. plantarum SQ001 on the relative abundance of species Lactobacillus plantarum in mice (n = 7). (m) Spearman correlation analysis between serum metabolites and gut microbes. Statistical significance in (e, f) was determined by unpaired two-tailed Student’s t-test. ns, no significance. *p < 0.05, **p < 0.01, ***p < 0.001. Data with error bars represent mean ± s.e.m. CON, control group; HUA, hyperuricemia model group; AL, allopurinol group; LP, L. plantarum SQ001 group.
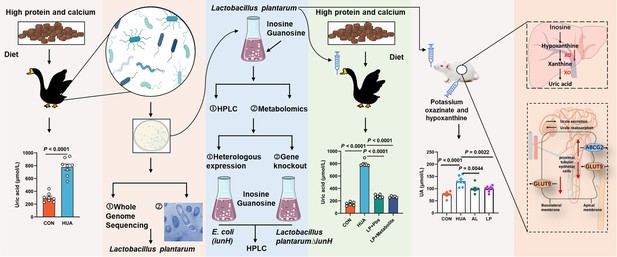
Working hypothesis.
The results suggest that host-derived Lactobacillus plantarum SQ001 hydrolyzes nucleosides through the nucleoside hydrolase iunH, reducing intestinal nucleoside transport, lowering hepatic uric acid production, restoring renal excretion, and ultimately alleviating hyperuricemia (HUA). iunH: Inosine-uridine nucleoside N-ribohydrolase; GLUT9: glucose transporter 9, UA reabsorption transporter; ABCG2: ATP-binding cassette transporter G2, UA excretion transporter.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Lactobacillus plantarum) | SQ001 | Lab stock | CGMCC NO 24791 | |
| Strain, strain background (E. coli) | GB2005 | Lab stock | (HS996, ∆recET, ∆ybcC). The endogenous recET locus and the DLP12 prophage ybcC, which encodes a putative exonuclease similar to the Redα, were deleted | |
| Strain, strain background (E. coli) | GB05-dir | Lab stock | (GB2005, araC-BAD-ETgA) recE, recT, redB3; and recA under BAD promoter was inserted at the ybcC locus | |
| Strain, strain background (E. coli) | Nissle1917/p15A-cm-Pgenta | This study | The Pgenta promoter gene was heterologous expressed in E. coli Nissle1917 | |
| Strain, strain background (E. coli) | Nissle1917/p15A-cm-iunH | This study | The iunH gene was heterologous expressed in E. coli Nissle1917 | |
| Antibody | anti-GLUT9 (Rabbit monoclonal) | Proteintech | Cat#26486-1-AP; RRID: AB_2880532 | 1:1000 |
| Antibody | anti-ABCG2 (Rabbit monoclonal) | Abcam | Cat#ab108312; RRID: AB_10861951 | 1:5000 |
| Antibody | anti-PRRS (Mouse monoclonal) | Bioss | Cat#bs-4504R; RRID: AB_11063921 | 1:1000 |
| Antibody | anti-XO (Mouse monoclonal) | Abcam | Cat#ab109235; RRID: AB_10863199 | 1:5000 |
| Antibody | Anti-β-actin (Mouse monoclonal) | Proteintech | Cat#60009-1-Ig; RRID: AB_2154438 | 1:5000 |
| Antibody | HRP goat anti-mouse IgG (goat polyclonal) | proteintech | Cat#SA00001-1; RRID: AB_2722565 | 1:5000 |
| Antibody | HRP goat anti-rabbit IgG (goat polyclonal) | proteintech | Cat#SA00001-2; RRID: AB_2722564 | 1:6000 |
| Sequence-based reagent | p15A-5 | This paper | PCR primers | AAACTACCGCATTAAAGCTT |
| Sequence-based reagent | p15A-3 | This paper | PCR primers | CTGAACCGACGACCGGGTCG |
| Sequence-based reagent | p15A-Pgenta-5 | This paper | PCR primers | CAGAAATTCGAAAGCAAATTCGACCCGGTCGTCGGTTCAGGAAGGCACGAACCCAGTTG |
| Sequence-based reagent | genta-60kDa-3 | This paper | PCR primers | CGTTGCTGCTCCATAACAT |
| Sequence-based reagent | Pgenta-iunH-5 | This paper | PCR primers | TACGCCGTGGGTCGATGTTTGATGTTATGGAGCAGCAACGGAGGATGTTTTACATGGCA |
| Sequence-based reagent | iunH-3 | This paper | PCR primers | TTTGACAGCTTATCATCGATAAGCTTTAATGCGGTAGTTTCTAATGGGCTTTAAATAAT |
| Sequence-based reagent | erm-5 | This paper | PCR primers | GTTCTATGCTTTCTTTTTGTAGCCGGCTAAACGGATAGTCCCCCAAAATCATCTTGCCTTTGATATTGAGGTATCATTT |
| Sequence-based reagent | erm-3 | This paper | PCR primers | CCTTTTTAATCACAATTCAGAAAATATCATAATATCTCATTTCACTAAATAATAGTGAACTTAGCCGTTAAATATTTTA |
| Sequence-based reagent | sacB-5 | This paper | PCR primers | GTTCACTATTATTTAGTGAA |
| Sequence-based reagent | sacB-3 | This paper | PCR primers | AGTGTGACTCTAGTAGAGAGCGTTCACCGACAAACAACAGTTTGTTAACTGTTAATTGT |
| Sequence-based reagent | Pi-HAR-2 | This paper | PCR primers | GACATTCCCGGCAATCGGAC |
| Sequence-based reagent | Pi-HAR-1 | This paper | PCR primers | AAGGCAAGATGATTTTGGGG |
| Sequence-based reagent | Pi-HAL-2 | This paper | PCR primers | TTTGAACACTCATGTTTAAC |
| Sequence-based reagent | Pi-HAL-1 | This paper | PCR primers | GATGGATGTTCAACTAATTTGTCCGATTGCCGGGAATGTCGCTGATTGATCTGAAAGGA |
| Sequence-based reagent | P15A-PiHAR-2 | This paper | PCR primers | TGTTAGTCATTTTCTTTCTCAACCTCGTCATTGGCACCAAGTTAAACATGAGTGTTCAAAGACGTCGATATCTGGCGAA |
| Sequence-based reagent | check-HA-PiunH-1 | This paper | PCR primers | TTCGCTTGAGGTACAGCGAA |
| Sequence-based reagent | check-HA-PiunH-2 | This paper | PCR primers | GCTGTTAATGGCAGAGGTAG |
| Software | SAS | SAS | RRID: SCR_008567 | Version 9.2 |
Additional files
-
Supplementary file 1
Strains, plasmids, mutants, primers, and antibiotics used in this study.
- https://cdn.elifesciences.org/articles/100068/elife-100068-supp1-v1.docx
-
Reporting standard 1
ARRIVE author checklist.
- https://cdn.elifesciences.org/articles/100068/elife-100068-repstand1-v1.pdf
-
MDAR checklist
- https://cdn.elifesciences.org/articles/100068/elife-100068-mdarchecklist1-v1.docx





