Layer 6 corticocortical neurons are a major route for intra- and interhemispheric feedback
Figures
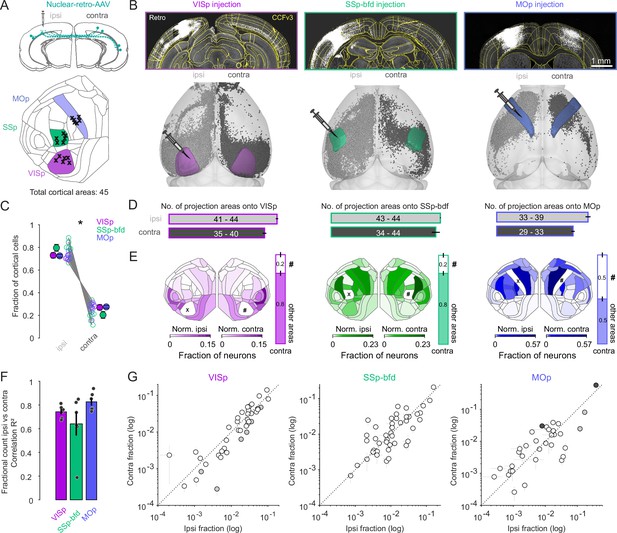
Widespread and bilaterally symmetrical cortical projections onto VISp, SSp-bfd, and MOp.
(A) Retrograde nuclear retro-AAV was injected across all layers of the primary visual cortex (VISp), somatosensory barrel field (SSp-bfd), and the motor cortex (MOp) of mice. The locations of the centroids of the injection bolus is shown on the CCFv3 cortical flat map. Injections performed in the right hemisphere (n = 11) are flipped to their corresponding location in the left hemisphere. (B) Two photon images showing examples of injection sites and labeled nuclei of retro-AAV labeled cells. Visualization of detected projection neurons for three individual mice warped into the 3D rendered space of the two hemispheres. (C) Plot showing the fraction of cortical cells per hemisphere detected for each mouse and pooled according to each injection site (mean ± SEM). Asterisks indicate statistical significance (p < 0.05). (D) Bar graphs indicating the mean number (± SEM) of cortical areas in which labeled neurons were observed. Text indicates the range of areas. (E) Flat maps showing the relative cell counts normalized for each hemisphere excluding both the injection target (X) area its homotopic counterpart (#). Right, bar graphs indicating the fraction of cells in the contralateral hemisphere located in the homotopic versus all other cortical areas (mean ± SEM). Note that areas ILA (5), ENTl (44), and ENTm (45) are not represented in the cortical flat map on the left (see also Figure S1C). (F) Bar graphs showing the average correlation (R2 ± SEM) of the ipsi- versus contralateral fractional count superimposed on the individual R2 values for each experiment. (G) Plot of the ipsi- versus contralateral fractional count for each of the three target areas. Each circle represents the average of six fractional count values (± SEM) for a given area. Open circles indicate instances where the difference of the ipsilateral values are not significant from the contralateral values. Black circles indicate a significant contralateral bias while gray circles indicate an ipsilateral bias. Dashed line indicates the unity line.
-
Figure 1—source data 1
Data corresponding to panels A, B, C, D, E, F, and G.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig1-data1-v1.xlsx
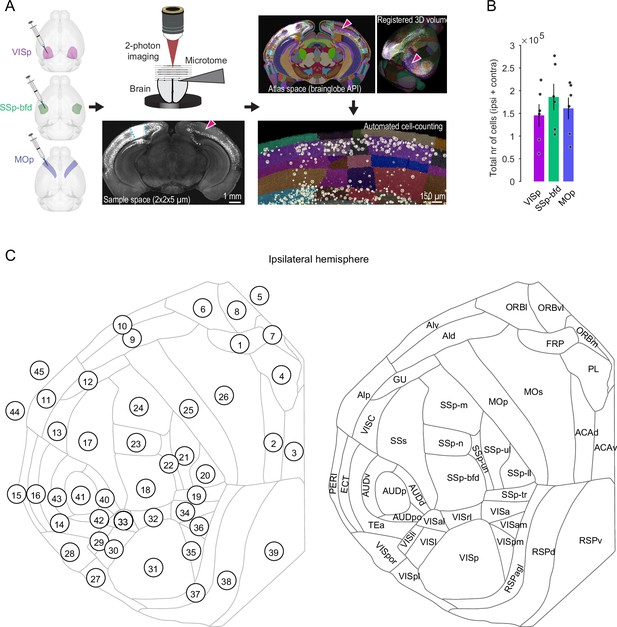
Experimental workflow and cell detection.
(A) AAV-EF1a-H2B-EGFP (nuclear retro-AAV) injection into VISp, SSp-bfd, and MOp. After viral expression, whole brains are cut and then imaged at x–y–z imaging resolution of ~2 × 2 × 5 µm using serial two-photon tomography. Blue dashed line indicates viral injection bolus. Whole brains are mapped onto the common reference atlas (CCFv3) and neurons are automatically detected and counted using the deep-learning based algorithm cellfinder (circles). Arrowheads indicate the contralateral counterpart of the injected area (homotopic area) containing retrogradely labeled nuclei of projection neurons. (B) Bar plot displaying average total number of detected cells (± SEM) in cortex for the three target areas (n = 6 mice, respectively). (C) Cortical flat map displaying all 45 cortical areas and their identification numbers (left) and abbreviations (right). (1) FRP, frontal pole; (2) ACAd, anterior cingulate area, dorsal part; (3) ACAv, anterior cingulate area, ventral part; (4) PL, prelimbic area; (5) ILA, infralimbic area; (6) ORBI, orbital area, lateral part; (7) ORBm, orbital area, medial part; (8) ORBvl, orbital area, ventrolateral part; (9) ALd, agranular insular area, dorsal part; (10) ALv, agranular insular area, ventral part; (11) ALp, agranular insular area, posterior part; (12) GU, gustatory areas; (13) VISC, visceral area; (14) TEa, temporal association area; (15) PERI, perirhinal area; (16) ECT, ectorhinal area; (17) SSs, supplemental somatosensory area; (18) SSp-bfd, primary somatosensory area, barrel field; (19) SSp-tr, primary somatosensory area, trunk; (20) SSp-ll, primary somatosensory area, lower limb; (21) SSp-ul, primary somatosensory area, upper limb; (22) SSp-un, primary somatosensory area, unassigned; (23) SSp-n, primary somatosensory area, nose; (24) SSp-m, primary somatosensory area, mouth; (25) MOp, primary motor area; (26) MOs, secondary motor area; (27) VISpl, posterolateral visual area; (28) VISpor, postrhinal area; (29) VISli, laterointermediate visual area; (30) VISl, lateral visual area; (31) VISp, primary visual area; (32) VISrl, rostrolateral visual area; (33) VISal, anterolateral visual area; (34) VISa, anterior visual area; (35) VISpm, posteromedial visual area; (36) VISam, anteromedial visual area; (37) RSPagl, retrosplenial area, lateral agranular part; (38) RSPd, retrosplenial area, dorsal part; (39) RSPv, retrosplenial area, ventral part; (40) AUDd, dorsal auditory area; (41) AUDp, primary auditory area; (42) AUDpo, posterior auditory area; (43) AUDv, ventral auditory area; (44) ENTl, entorhinal area, lateral part; (45) ENTm, entorhinal area, medial part. Note that areas (5), (44), and (45) are not represented in the cortical flat map on the right.
-
Figure 1—figure supplement 1—source data 1
Data corresponding to panel B.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig1-figsupp1-data1-v1.xlsx
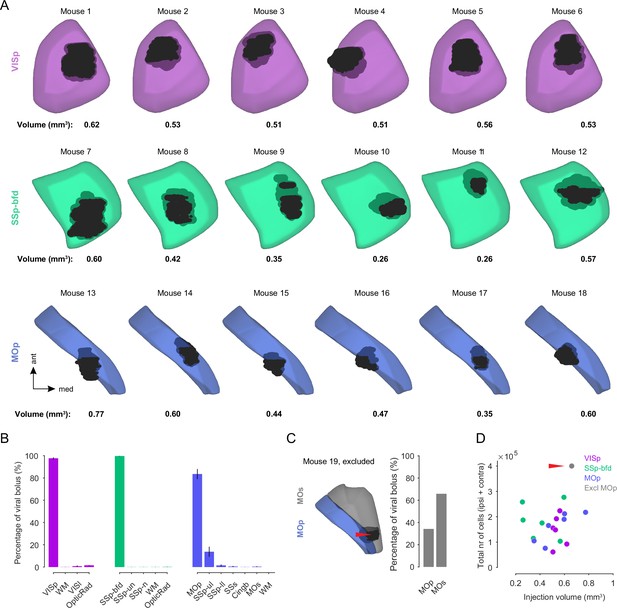
Target area viral expression quantification.
(A) 3D reconstruction of the injection site for each mouse warped into the 3D rendered space of VISp (top), SSp-bfd (middle), and MOp (bottom) of the CCFv3. The value of the volume (in mm3) for each bolus is indicated below. (B) Bar plot displaying the average percentage of the viral bolus (± SEM) located in the target and neighboring areas for VISp, SSp-bfd, and MOp. WM: white matter. (C) Example of a viral injection targeted in MOp which was excluded from the dataset as the majority of the viral bolus was located in the secondary motor cortex (MOs, red arrowhead) in this mouse. (D) Scatter plot displaying the volume per viral injection versus the total cell number detected in the cortex for each mouse for the three target areas. Red arrowhead highlights injection which was primarily located in MOs and was therefore excluded from the dataset.
-
Figure 1—figure supplement 2—source data 1
Data corresponding to panels B, C and D.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig1-figsupp2-data1-v1.xlsx
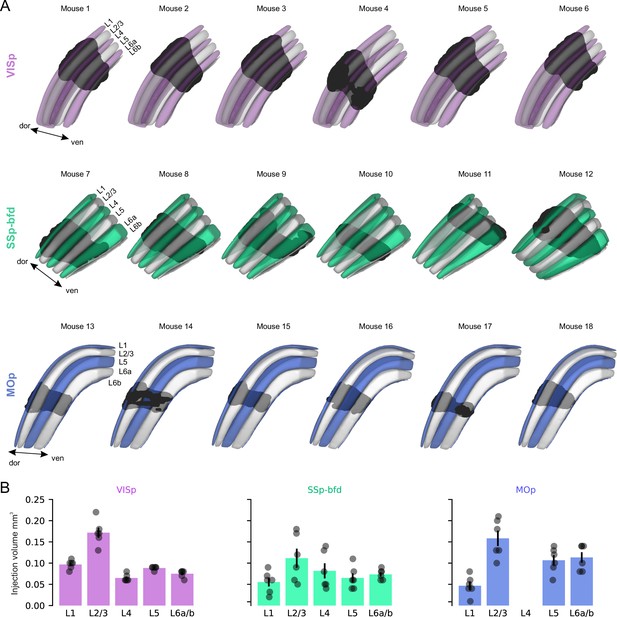
Latero-medial view of target area viral expression.
(A) Latero-medial view of the reconstructed injection site for each mouse warped into the 3D rendered space of VISp (top), SSp-bfd (middle), and MOp (bottom) of the CCFv3 (see also Figure 1—figure supplement 2). (B) Fraction of injection volume per cortical layer for VISp, SSp-bfd, and MOp.
-
Figure 1—figure supplement 3—source data 1
Data corresponding to panel B.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig1-figsupp3-data1-v1.xlsx
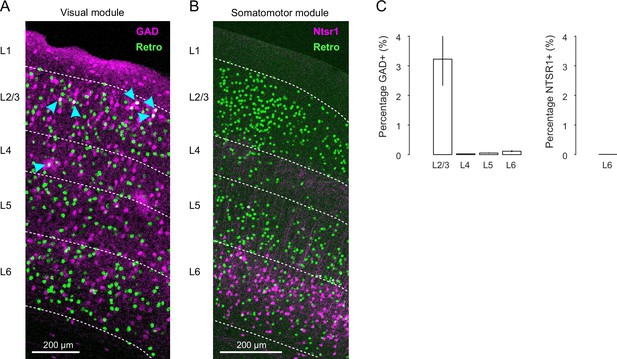
Molecular identity of long-range projection neurons.
(A) Confocal image displaying colocalization of GFP+ retrogradely labeled cells and GAD+ (glutamate decarboxylase) interneurons across cortical layers in an example coronal section of a cortical area of the visual module. Overlap is highlighted with arrowheads. (B) Confocal merged image displaying GFP+ retrogradely labeled cells and NTSR1+ (Neurotensin receptor 1) cells in an example coronal section of a cortical area of the somatomotor module. Note that there is no overlap. (C) (Left) Bar plot displaying the average retrogradely labeled neurons (± SEM) that are GAD+ interneurons in L2/3, L4, L5, and L6 across all cortical areas of both hemispheres. (Right) Bar plot displaying the average retrogradely labeled neurons that are NTSR1+ in L6 across all cortical areas of both hemispheres.
-
Figure 1—figure supplement 4—source data 1
Data corresponding to panel B.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig1-figsupp4-data1-v1.xlsx
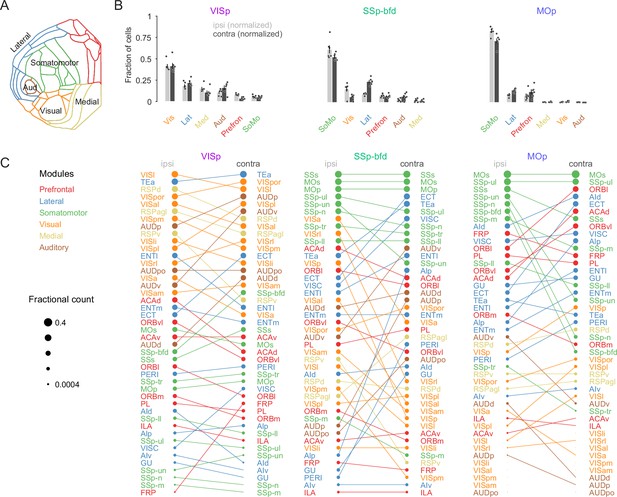
Modular organization of the intra- and interhemispheric cortical projection onto VISp, SSp-bfd, and MOp.
(A) Schematic of the cortical flat map highlighting the six cortical modules and their corresponding areas. (B) Bar graphs of the normalized fractional cell counts for each of the six modules for ipsi- and contralateral hemispheres ranked by decreasing ipsilateral weights. For the contralateral hemisphere the homotopic target region is not included. (C) Rank plot based on the relative weights of connectivity onto the target areas VISp, SSp-bfd, and MOp for all cortical brain areas. The size of the circles indicates relative fractional count on a logarithmic scale. Corresponding areas on the ipsi- and contralateral hemisphere are connected with a line.
-
Figure 2—source data 1
Data corresponding to panels B and C.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig2-data1-v1.xlsx
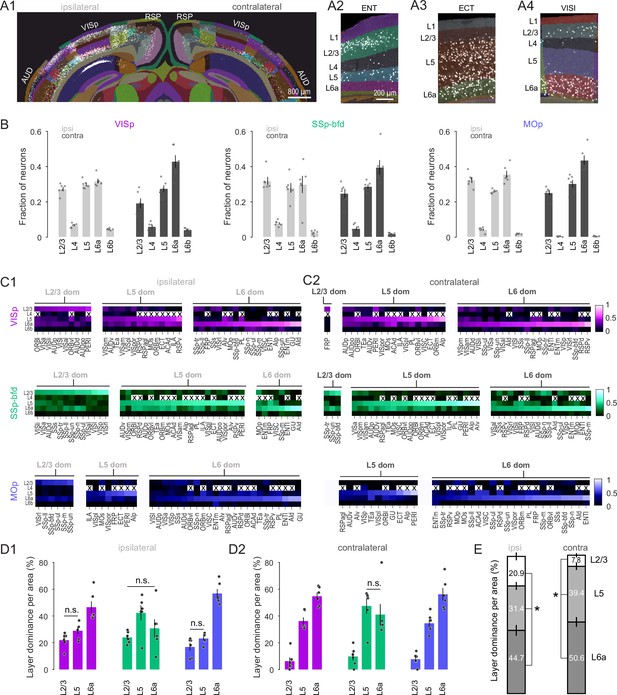
Cortical Layer 6 is a major source of input to VISp, SSp-bfd, and MOp.
(A1) A segmented image of the cortex used to assign the laminar position of detected neurons after injection into VISp. Image of segmented entorhinal (ENT), ectorhinal (ECT), and lateral visual area (VISl) on the contralateral hemisphere showing local dominance by L2/3 (A2), L5 (A3), and L6a (A4), respectively. (B) Bar plot showing average fraction of cells (± SEM, n = 6 mice) in L2/3, L4, L5, L6a, and L6b across the ipsi- and contralateral hemisphere for the three injection targets. (C) Average heatmaps (n = 6 mice) based on the fractional count across laminae for each cortical area separated according to injection target and local dominance for ipsilateral (C1) and contralateral (C2) hemispheres. Dominance quantification is based on average laminar fractional count across animals. Crosses indicate cortical areas that do not contain L4. (D) Bar graphs showing the average percentage of areas (± SEM, n = 6 mice) in which L2/3, L5, and L6 dominate the fractional cell count for ipsilateral (D1) and contralateral (D2) hemispheres. Unless indicated, all comparisons reach statistical significance (n.s. = non-significant). Dominance quantification is based on laminar fractional count per animal. (E) Stacked bar plots showing average laminar dominance for all cortical areas (± SEM) pooled across all injection targets. Asterisks indicate statistical significance (p < 0.05).
-
Figure 3—source data 1
Data corresponding to panels B, C, D and E.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig3-data1-v1.xlsx
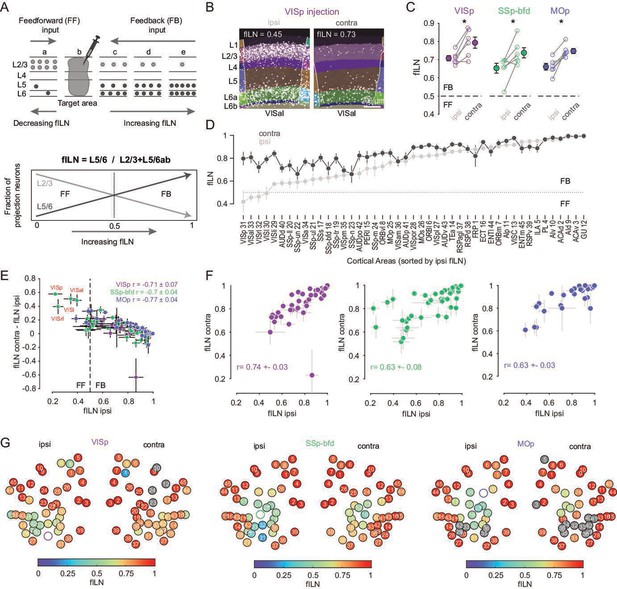
Cortical hierarchy onto VISp, SSp-bfd, and MOp.
(A) Cartoon showing the laminar basis of the anatomical determination of cortical hierarchy (top). As the fraction of supragranular (L2/3) neurons decrease and/or the number of L5 and L6 neurons increase, the fraction of infragranular neurons (fILN) increases reflecting a predominance of feedback input (bottom). (B) Segmented image of the ipsi- and contralateral anterolateral visual area (VISal) projecting onto VISp showing the laminar position of detected cells. (C) Plots showing fILN values for all neurons within each hemisphere (blind to individual cortical areas) for each mouse for the three target injections. Average fILN values (± SEM) are depicted by filled circles. Asterisk indicates statistical significance (p < 0.05). (D) Plot showing the distribution of the average of ipsilateral fILN values (± SEM) for n = 45 areas (pooled across different injection experiments) and ranked from lowest to highest (gray dots). The cortical areas abbreviations and their identification numbers are displayed at the bottom. Note: the number of data points contributing to each mean value ranges from 10 to 18 depending on the number of NaN values for a given area and whether it was excluded due to it being the target injection site. The average fILN values for the same but contralateral areas are also plotted according to their corresponding ipsilateral rank (black dots). (E) Scatter plot showing the correlation between the ipsilateral fILN values and the difference between the contra- and ipsilateral fILN values for the three target injections. Average correlation values (± SEM, n = 6 mice) for each target area are displayed (top right). Highlighted in red are the primary (VISp), rostrolateral (VISrl), lateral (VISl), and anterolateral (VISal) visual areas projecting to SSp-bfd that display low ipsilateral fILN and high contralateral fILN values. (F) Scatter plots showing the correlation between the ipsi- and contralateral fILN values for the three target injections. Average correlation values (± SEM, n = 6 mice) for each target area are displayed. (G) Heatmap plot displaying the ipsi- and contralateral fILN values for each cortical brain area for the three injection targets. Position of the circles correspond approximately to area location. Open circles denote injection areas. Gray circles indicate cortical areas where the number of cells projecting to VISp, SSp-bfd, or MOp did not meet criterion. Numbers 1–45 indicate different brain areas, see also D.
-
Figure 4—source data 1
Data corresponding to panels C, D, E, F, and G.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig4-data1-v1.xlsx
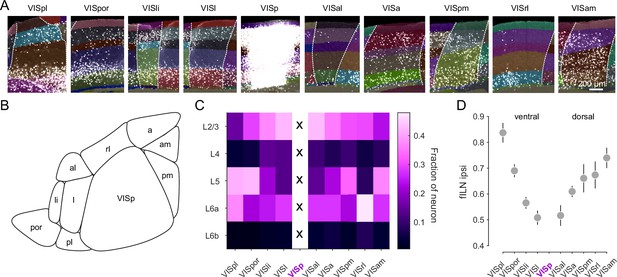
Cortical hierarchy of visual areas based on fILN.
(A) Representative images of retrogradely labeled nuclei in higher visual cortical areas in the ipsilateral hemisphere and the corresponding injection site in VISp. (B) Schematic illustration of the anatomical arrangement of VISp with the surrounding higher visual cortical areas in the left hemisphere. (C) Heatmap showing the layer-specific average projection weights of higher visual cortical areas onto VISp. (D) Rank ordered average fILN values (± SEM) of higher visual cortical areas of the ventral and dorsal visual stream. Injection target highlighted with magenta.
-
Figure 4—figure supplement 1—source data 1
Data corresponding to panels C and D.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig4-figsupp1-data1-v1.xlsx
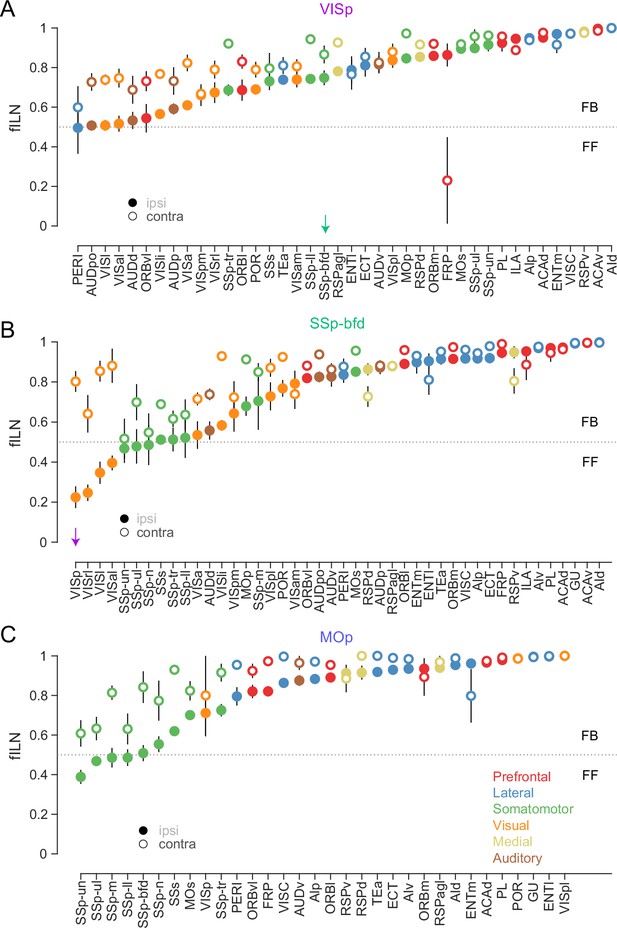
Cortical hierarchy per projection area onto VISp, SSp-bfd, and MOp.
Plot showing the average ipsilateral (closed circle) and contralateral (open circle) fILN values (± SEM) for the individual cortical areas containing both ipsi- and contralateral projections (excluding the injection target area, respectively) for VISp (A), SSp-bfd (B), and MOp (C). Respective fILN values ranked according to the ipsilateral fILN values. Green arrow in (A) highlights ipsi- and contralateral feedback projection from SSp-bfd to VISp. Magenta arrow in (B) highlights ipsilateral feedforward but contralateral feedback projection from VISp to SSp-bfd.
-
Figure 4—figure supplement 2—source data 1
Data corresponding to panels A, B amd C.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig4-figsupp2-data1-v1.xlsx
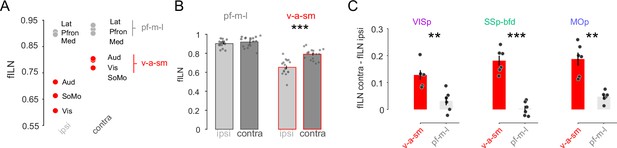
Sensory and motor but not lateral, prefrontal, and medial projections explain the hemispheric differences in cortical hierarchy.
(A) Plot showing the average fILN values for all areas in a given module for n = 18 mice irrespective of target area. Red circles indicate the sensory and motor modules (visual, auditory, and somatomotor; v-a-sm) while gray circles indicate prefrontal, medial, and lateral (pf-m-l) modules. (B) Bar graphs comparing cross hemispheric fILN values (mean ± SEM) for prefrontal, medial, and lateral (pf-m-l) and sensory and motor (v-a-sm) modules. Each circle represents a single animal (n = 18). (C) Bar graphs (mean ± SEM) showing the interhemispheric difference in fILN values between the two modular groupings for each target projectome. Each circle represents a single animal (n = 6, respectively). Asterisks indicate statistical significance (** p < 0.01, *** p<0.001).
-
Figure 5—source data 1
Data corresponding to panels A, B and C.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig5-data1-v1.xlsx
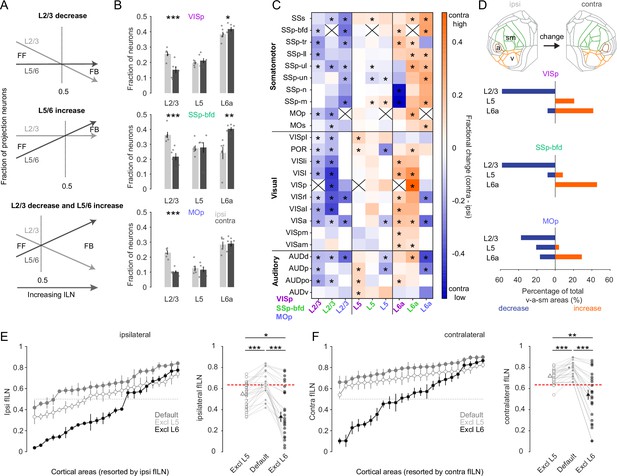
L6 dominates cortical hierarchy in the majority of sensory and motor areas.
(A) Schematic illustrating at least three hypothetical ways in which changes in the relative distribution of L2/3, L5, and L6 neurons can give rise to increases in the fILN and reflect a predominantly feedback network. (B) Bar graphs showing the average (± SEM) normalized fractional count (excluding L4 and 6b) of cells for the sensory and motor modules projecting to each target area from both hemispheres (ipsi, gray; contra, black), (C) Heatmap displaying the fractional change of projections for L2/3, L5, and L6a between the contra- and ipsilateral hemisphere. Displayed are the 24 individual brain areas in the sensory and motor modules for the three target areas VISp, SSp-bfd, and MOp. Asterisks denote areas that significantly change between the ipsi- and the contralateral hemispheres. Crosses indicate excluded homotopic areas. (D) Flat map indicating the three sensory and motor modules (top). Stacked horizontal bar plots displaying the percentage of the total number cortical areas in the sensory and motor modules that significantly decrease (blue) or increase (orange; indicated by the asterisks in C) for the three target areas. (E) (Left) Plots showing the average fILN (± SEM) for each sensory and motor area (pooled across all target areas, n = 18 mice) and ranked according to the ipsilateral values for the default network (gray filled circles), the default network excluding L5 (open circles), and the default network excluding L6 (black filled circles). (Right) Scatter plots showing the average ipsilateral fILN scores for each of the 24 areas of the default network, the default network excluding L5 and the default network excluding L6. Filled triangles represent the mean (± SEM) of the average fILN values. Red line indicates the mean of the ipsi default network. (F) (Left) Plots showing the average fILN (± SEM) for each sensory and motor area (pooled across all target areas, n = 18 mice) and ranked according to the ipsilateral values for the default network (gray filled triangles), the contralateral values for the default network (gray filled circles), the contralateral values for the default network excluding L5 (open circles), and the default network excluding L6 (black filled circles). (Right) Scatter plots showing the average fILN scores for each of the 24 areas of the ipsi and contra default network, the default contra network excluding L5 and the default contra network excluding L6. Filled triangles represent the mean (± SEM) of the average fILN values. Red line indicates the mean of the ipsi default network. Asterisks indicate statistical significance (* p<0.05, ** p < 0.01, *** p<0.001).
-
Figure 6—source data 1
Data corresponding to panels B, C, D, E and F.
- https://cdn.elifesciences.org/articles/100478/elife-100478-fig6-data1-v1.xlsx
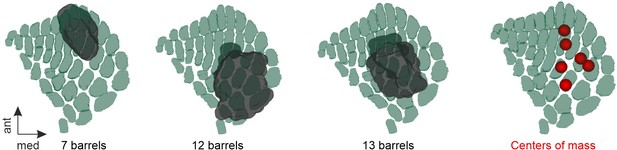
Left: exemplary Injection sites mapped onto the 3D barrel map of SSp-bfd within the Mouse Allen Brain Atlas.
Barrels were reconstructed using a specialized software as described previously (Weiler et al., 2024) Right: Centres of mass of all SSp-bfd injection sites mapped onto the 3D barrel map.
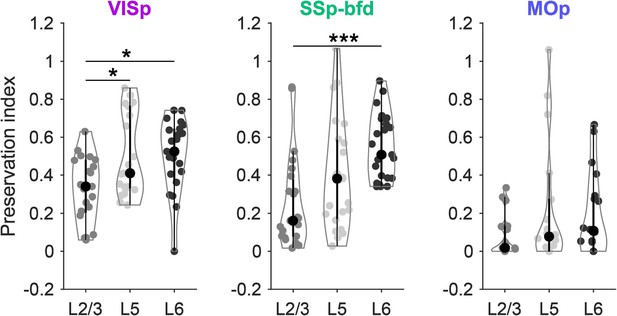
Preservation index for L2/3, L5 and L6 of the 24 sensory-motor areas projecting onto the three target areas VISp, SSp-bfd and MOp.
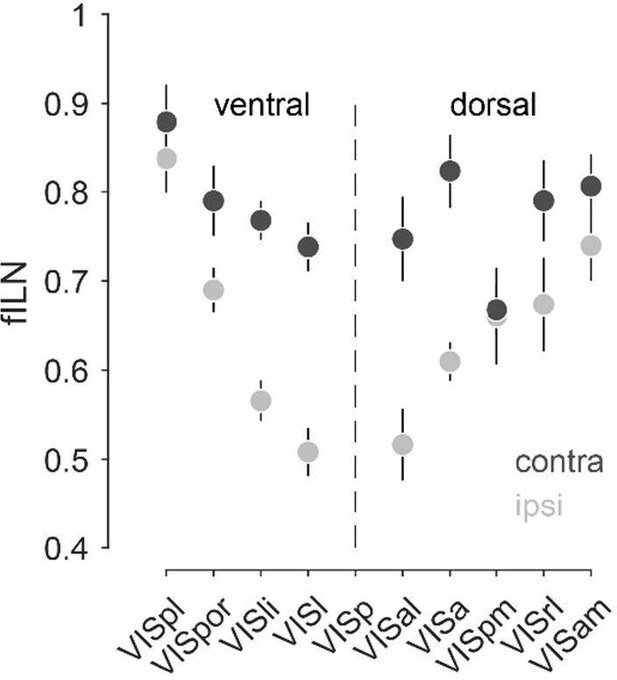
Rank ordered average fILN values (± sem) of higher visual cortical areas of the ventral and dorsal visual stream for the ipsilateral and contralateral hemisphere.
Tables
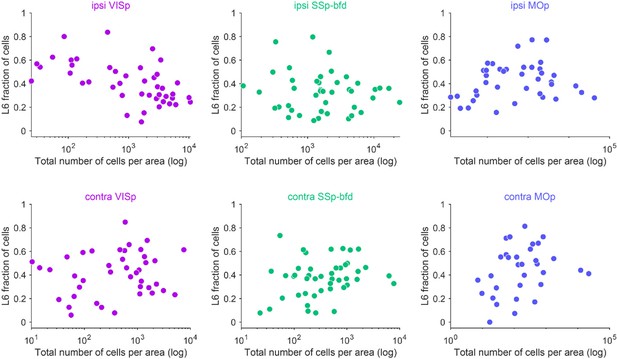
Respective correlations between total numbers of cells and fraction of cells within L6 per cortical brain area for the ipsilateral and contralateral hemisphere for the three target areas (significant correlations highlighted with green).
| VISp | ipsi | |
| Animals | Correlation r Total numbers vs. L6 fraction | |
| 1 | -0.6265 | |
| 2 | -0.6469 | |
| 3 | -0.2314 | |
| 4 | -0.6836 | |
| 5 | -0.6539 | |
| 6 | -0.4186 | |
| contra | ||
| Animals | Total numbers vs. L6 fraction | |
| 1 | 0.196 | |
| 2 | 0.398 | |
| 3 | 0.646 | |
| 4 | 0.1494 | |
| 5 | 0.0579 | |
| 6 | 0.3676 | |
| SSp-bfd | ipsi | |
| Animals | Correlation r Total numbers vs. L6 fraction | |
| 1 | 0.3942 | |
| 2 | 0.0452 | |
| 3 | -0.0533 | |
| 4 | -0.4984 | |
| 5 | -0.234 | |
| 6 | 0.0233 | |
| contra | ||
| Animals | Correlation r Total numbers vs. L6 fraction | |
| 1 | 0.5548 | |
| 2 | 0.2174 | |
| 3 | -0.0788 | |
| 4 | 0.0369 | |
| 5 | 0.0297 | |
| 6 | 0.2001 | |
| MOp | ipsi | |
| Animals | Correlation r Total numbers vs. L6 fraction | |
| 1 | 0.3736 | |
| 2 | 0.4007 | |
| 3 | 0.4911 | |
| 4 | 0.4287 | |
| 5 | 0.1059 | |
| 6 | 0.2606 | |
| contra | ||
| Animals | Correlation r Total numbers vs. L6 fraction | |
| 1 | 0.5828 | |
| 2 | 0.6672 | |
| 3 | 0.6221 | |
| 4 | 0.6355 | |
| 5 | 0.5375 | |
| 6 | 0.639 |