The horizontally-acquired response regulator SsrB drives a Salmonella lifestyle switch by relieving biofilm silencing
Figures
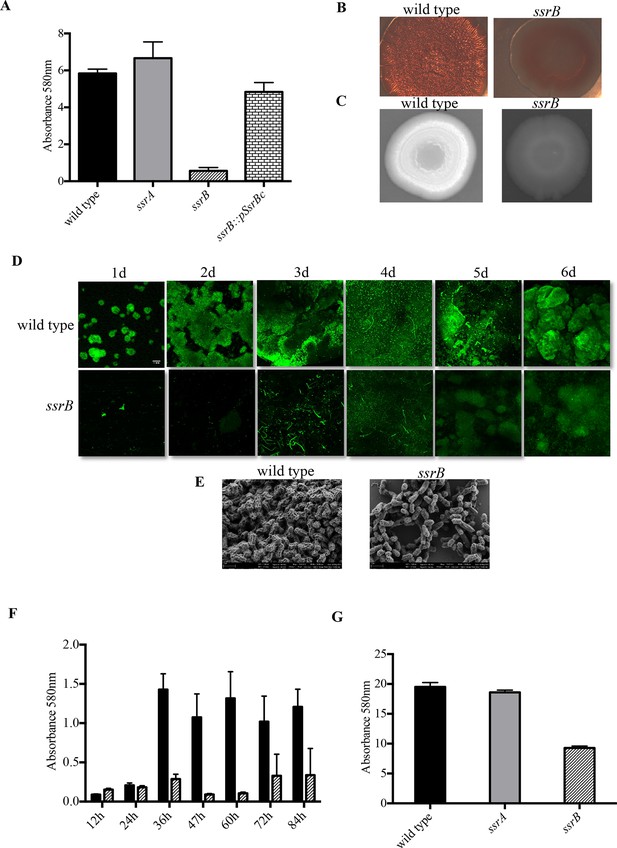
Loss of ssrB but not ssrA decreases Salmonella Typhimuirum biofilms.
(A) The defect in formation of biofilms in the ssrB null was complemented by the overexpression of SsrBc from plasmid pKF104 in trans as measured by crystal violet staining. (B) The typical rdar morphotype of the wild type strain was lost in the ssrB strain as shown on congo red plates. (C) A two day old macrocolony of the ssrB strain is not fluorescent under UV light with Fluorescent Brightener 28. (D) The wild type strain forms thick solid-surface biofilms, while the ssrB strain remains poor for biofilms as monitored for six days by SYTO-9 staining of flow cell biofilms; scale bar = 1 mm. (E) SEM images showing extensive mesh-like network of wild type biofilms and sparse extracellular matrix of the ssrB biofilms; scale bar = 1 µm. (F) The amount of biofilms formed by the wild type strain (solid black bars) increases after 24 hr but the ssrB null (hatched black bars) remains defective up to 84 hr. n = 2, Mean ± SD, p < 0.0001 between wild type and ssrB strains from 36 hr till 84 hr. (G) The amount of cholesterol-attached biofilms formed by the ssrB strain were significantly less than that produced by the wild type. n = 3, Mean ± SD, p < 0.0001. Source data file: Figure 1—source data 1.
-
Figure 1—source data 1
Source data for crystal violet staining in Figure 1A,F and G.
- https://doi.org/10.7554/eLife.10747.004
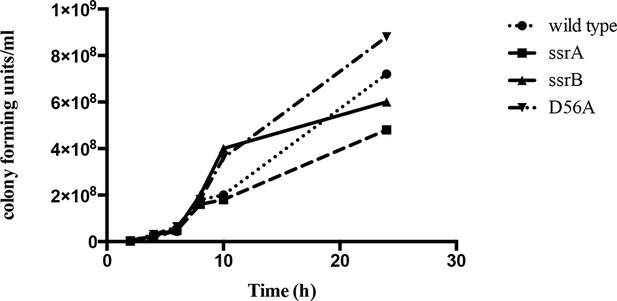
The ssrB mutant is not defective in growth compared to the wild type strain.
Number of colonies formed by the wild type, ssrA, ssrB and D56A strains were the same order of magnitude across all the time points tested. Source data file: Figure 1—figure supplement 1—source data 1.
-
Figure 1—figure supplement 1—source data 1
Growth curves of wild type, ssrA, ssrB and D56A strains.
- https://doi.org/10.7554/eLife.10747.006
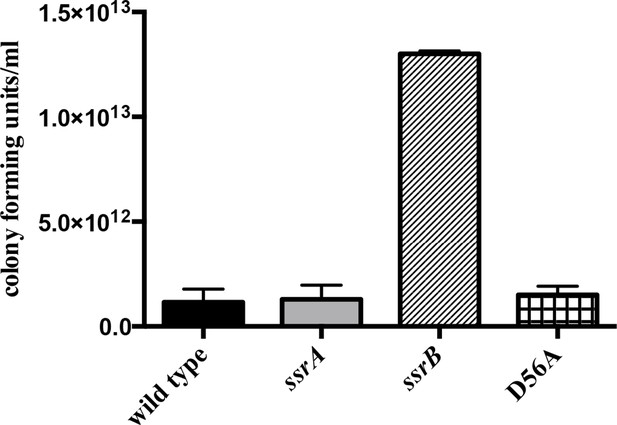
The planktonic sub-population of the ssrB strain was higher by two orders of magnitude compared to the wild type, ssrA and D56A strains at 2 days.
n = 2, Mean ± SD, p < 0.05. Source data file: Figure 1—figure supplement 2—source data 1.
-
Figure 1—figure supplement 2—source data 1
Number of cells in the planktonic sub-population of each strain.
- https://doi.org/10.7554/eLife.10747.008
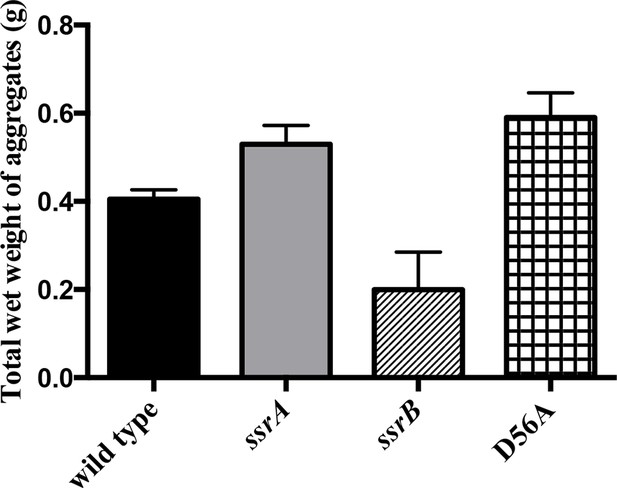
Total wet weight of the adherent sub-population was decreased by at least 50% in the ssrB strain compared to the wild type, ssrA and D56A strains at 2 days.
n = 2, Mean ± SD, p < 0.05 for the ssrB strain versus ssrA/D56A strains and p = 0.08 for the ssrB strain versus wild type. Source data file: Figure 1—figure supplement 3—source data 1.
-
Figure 1—figure supplement 3—source data 1
Total wet weight of the adherent sub-population of each strain.
- https://doi.org/10.7554/eLife.10747.010
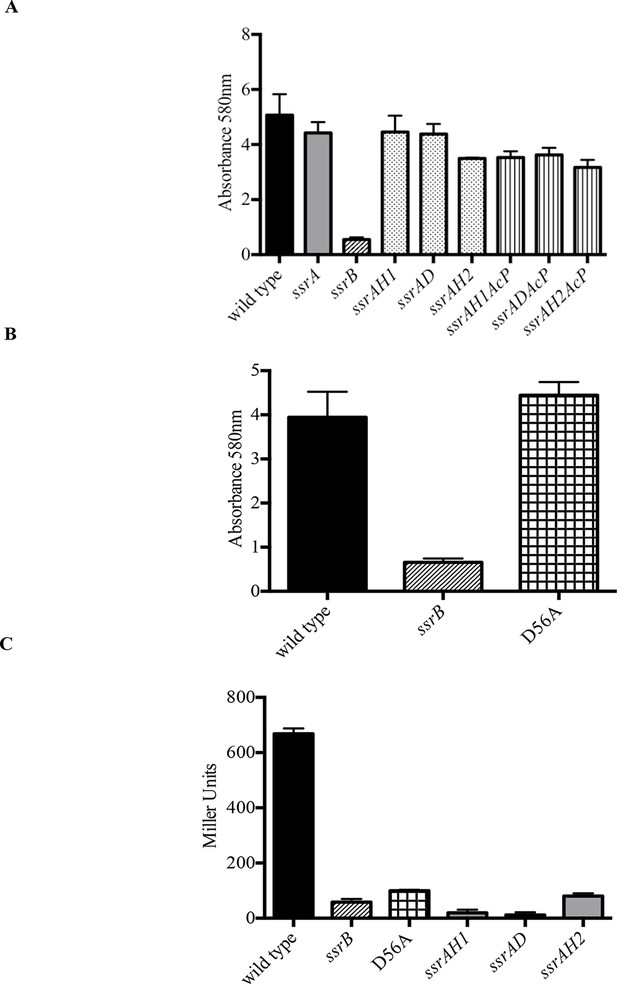
Phosphorylation of SsrB is not required for biofilm formation.
Amount of biofilms formed as measured by crystal violet staining for (A) Strains ssrAH1, ssrAD, ssrAH2, ssrAH1AcP, ssrADAcP, ssrAH2AcP and (B) D56A SsrB shows similar levels to that of the wild type, and higher than the ssrB mutant. Source data file: Figure 2—source data 1. (C) Beta-galactosidase activity of a sifA-lacZ chromosomal fusion was significantly lower in the ssrB null and the D56A SsrB mutant compared to the wild type. n = 3, Mean ± SD, p < 0.0001. Source data file: Figure 2—source data 2.
-
Figure 2—source data 1
Source data for crystal violet staining in Figure 2A and B.
- https://doi.org/10.7554/eLife.10747.012
-
Figure 2—source data 2
Source data for the measurement of beta-galactosidase activity in Figure 2C.
- https://doi.org/10.7554/eLife.10747.013
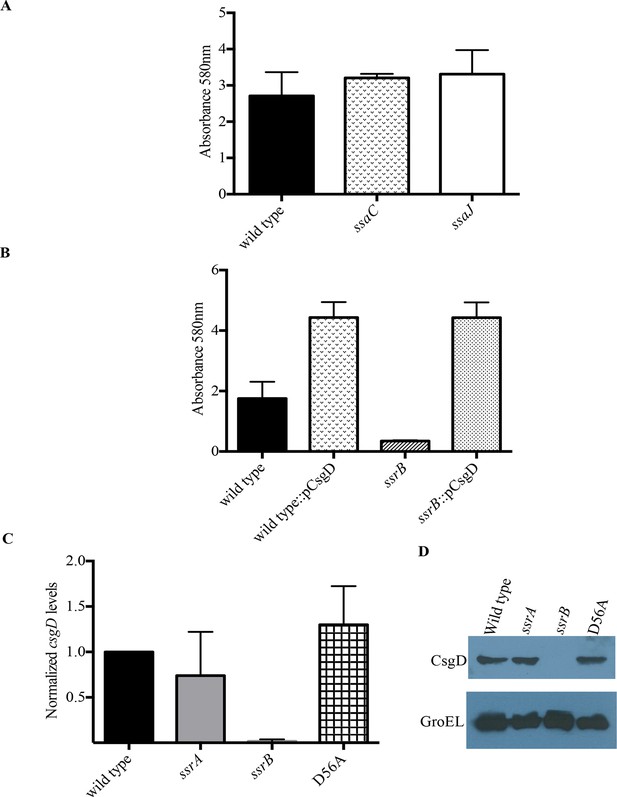
SsrB regulates biofilms by a CsgD-dependent mechanism.
(A) The SPI-2 needle, ssaC and ssaJ mutant strains were not affected in biofilm formation. (B) Over-expression of csgD from a plasmid (pBR328::csgD) in trans rescued biofilm formation in the ssrB mutant, as measured by crystal violet staining, n = 3. Source data file: Figure 3—source data 1. An estimate of csgD expression by (C) Real-time qRT-PCR showed a significant decrease in csgD transcription in the ssrB null, but not in the D56A SsrB and ssrA mutants. rrsA transcript levels were used as control; n = 2, Mean ± SD, p < 0.0001. Source data file: Figure 3—source data 2 and (D) Immunoblot analysis showing the absence of CsgD in the ssrB null strain in two day old biofilms, using GroEL as a loading control.
-
Figure 3—source data 1
Source data for crystal violet staining in Figure 3A and B.
- https://doi.org/10.7554/eLife.10747.015
-
Figure 3—source data 2
Source data for Real-time qRT-PCR in Figure 3C.
- https://doi.org/10.7554/eLife.10747.016
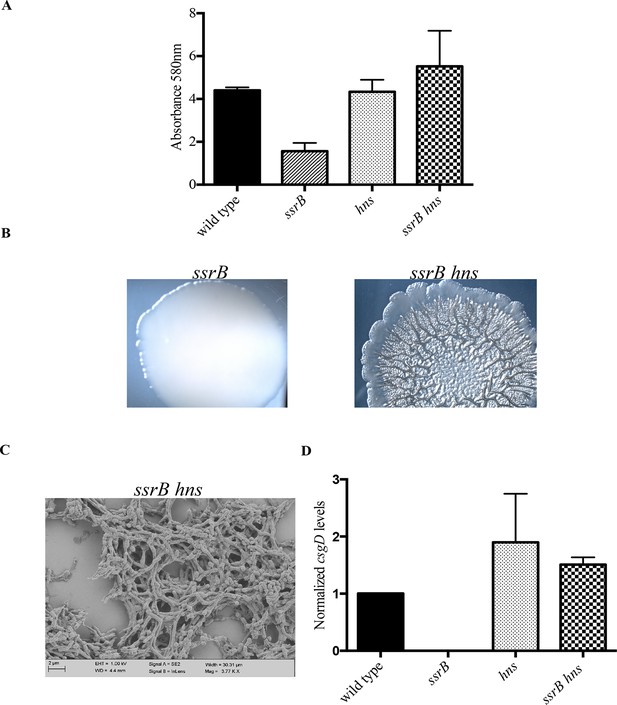
hns deletion rescues biofilm formation in the ssrB mutant as shown by Crystal violet staining.
(A) The amount of biofilms formed is higher in the wild type, hns and ssrB hns strains compared to the ssrB null, n = 3, Mean ± SD, p < 0.0001. Source data file: Figure 4—source data 1. Macrocolony phenotype (B) ssrB hns forms a highly rugose and dry macrocolony, while the ssrB macrocolony was smooth and mucoidy. SEM imaging (C) ssrB hns biofilms were covered by a thick extra-cellular matrix; scale bar = 2 µm. (D) qRT-PCR: csgD levels were restored in the ssrB hns strain and were higher than the wild type (p < 0.03) and the ssrB mutant (p < 0.003) against rrsA transcripts as a control. Note that the normalized csgD levels in the ssrB null were 0.0009, too low for the scale. n = 2, Mean ± SD. Source data file: Source data file: Figure 4—source data 2.
-
Figure 4—source data 1
Source data for crystal violet staining in Figure 4A.
- https://doi.org/10.7554/eLife.10747.018
-
Figure 4—source data 2
Source data for Real-time qRT-PCR in Figure 4D.
- https://doi.org/10.7554/eLife.10747.019
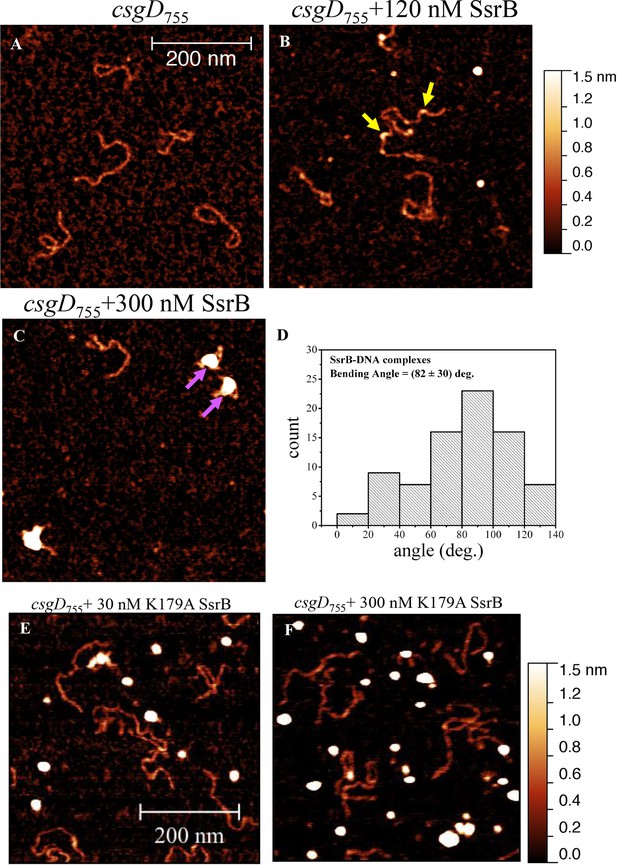
SsrB binds upstream of csgD.
(A) AFM images of the 755 bp csgD regulatory region (csgD755). (B) At 120 nM SsrB, distinct areas of SsrB binding were visualized as sharp bends (yellow arrows). (C) At 300 nM SsrB, areas of condensation (pink arrows) were observed. (D) Binding of SsrB bends the DNA by an average angle of 82º (for the naked DNA angle, refer to Figure 5—figure supplement 2 and for analysis refer to Supplementary method), Scale bar = 200 nm as in (A). (E) and (F) The SsrB mutant, K179A SsrB, which is defective in DNA binding, was unable to bind csgD755 both at 30 nM or 300 nM; Scale bar = 200 nm.
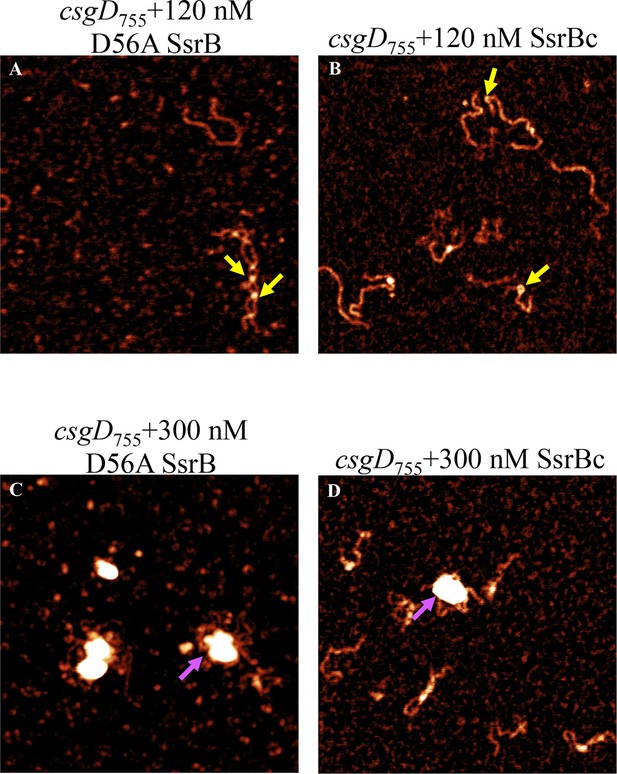
AFM images of the 755 bp csgD regulatory region (csgD755) (A) with 120 nM D56ASsrB and (B) SsrBc.
Distinct areas of binding were visualized as sharp bends (yellow arrows) and as areas of condensation (pink arrows) at 300 nM D56A SsrB (C) and SsrBc (D); Scale bar = 200 nm as in Figure 5A.
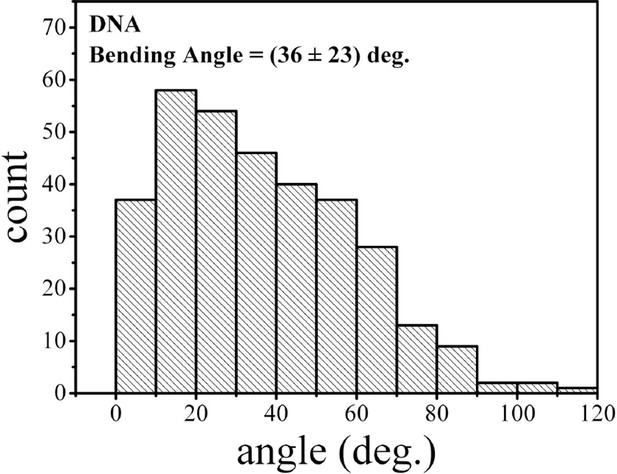
Bending angle of the naked csgD755 fragment.
https://doi.org/10.7554/eLife.10747.022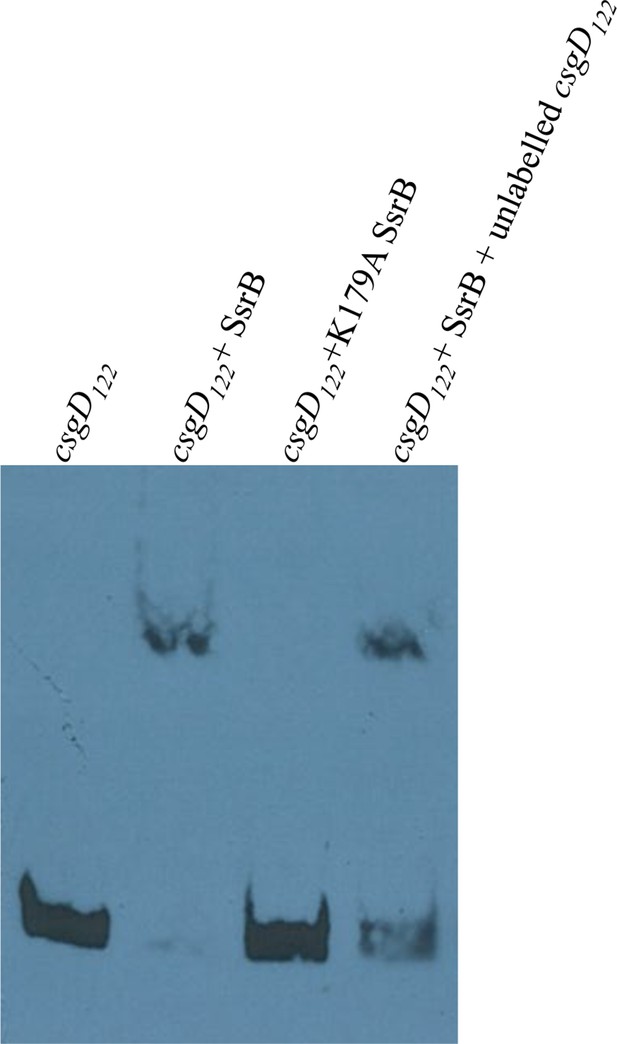
Electrophoretic mobility shift assay with the 122 bp csgD regulatory region, csgD122, showing a DNA-protein complex in the presence of SsrB.
The K179A SsrB mutant did not bind to csgD122. Addition of competitor unlabelled csgD122 fragment decreased the SsrB-DNA complex as apparent by an increase in free, labelled csgD122.
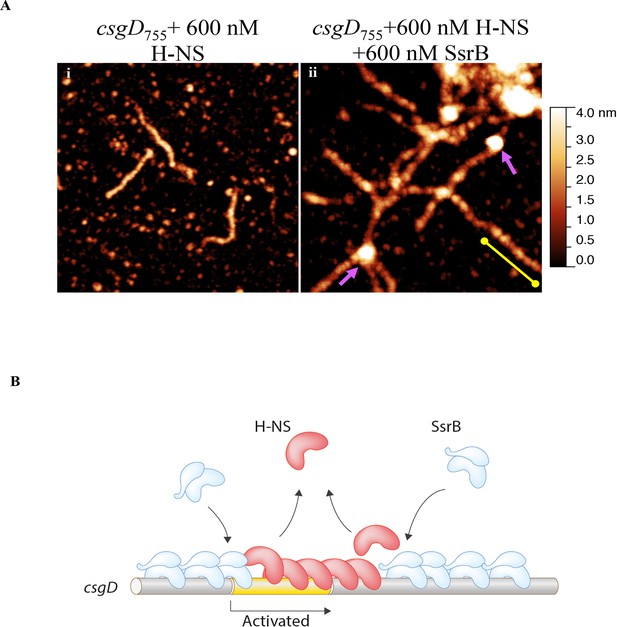
SsrB condenses H-NS bound csgD DNA.
(A) (i) AFM imaging in the presence of 600 nM H-NS shows a straight and rigid filament on csgD755. (ii) Addition of 600 nM SsrB to the H-NS bound csgD DNA resulted in areas of condensation (pink arrows; an ‘SsrB signature’) along with a few areas where the straight H-NS bound conformation persisted (yellow line; an ‘H-NS signature’); Scale bar = 200 nm as in Figure 5A. (B) A model for the mechanism of anti-silencing by SsrB at csgD wherein SsrB likely displaces H-NS from the ends of a stiffened nucleoprotein filament and relieves the blockade on the promoter for RNA polymerase to activate transcription. For details refer to (Winardhi et al., 2015).
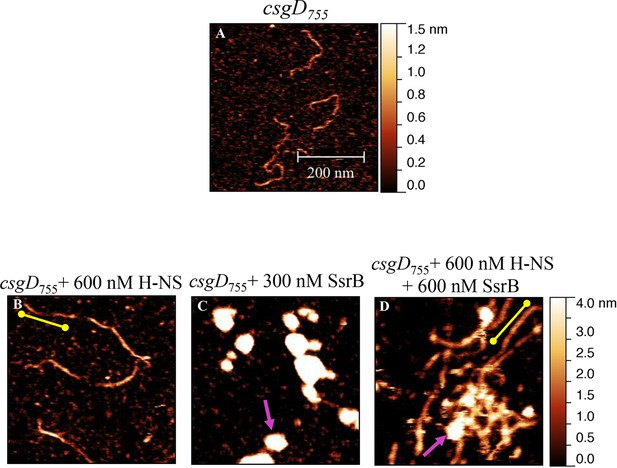
Liquid AFM imaging of (A) the 755 bp csgD regulatory region.
(B) In the presence of 600 nM H-NS, a straight and rigid filament was observed (yellow line). (C) In the presence of 300 nM SsrB, areas of condensation were evident (pink arrow). (D) Addition of 600 nM SsrB to the H-NS bound csgD DNA resulted in areas of condensation (pink arrows; an ‘SsrB signature’) along with a few areas where the straight H-NS bound conformation persisted (yellow line; an ‘H-NS signature’); Scale bar = 200 nm.
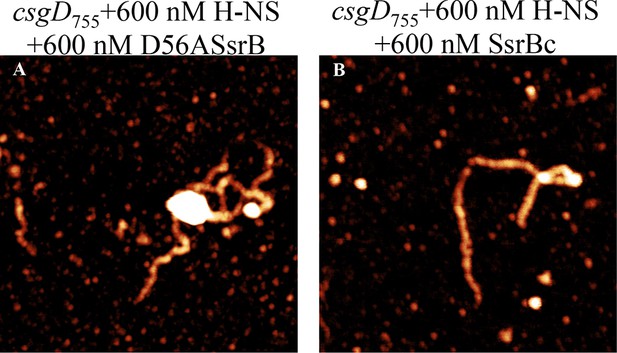
SsrB D56A and SsrBc condense H-NS-bound csgD DNA.
AFM imaging in the presence of H-NS shows areas of condensation upon addition of (A) 600 nM D56ASsrB and (B) 600 nM SsrBc; Scale bar = 200 nm as in Figure 5A.
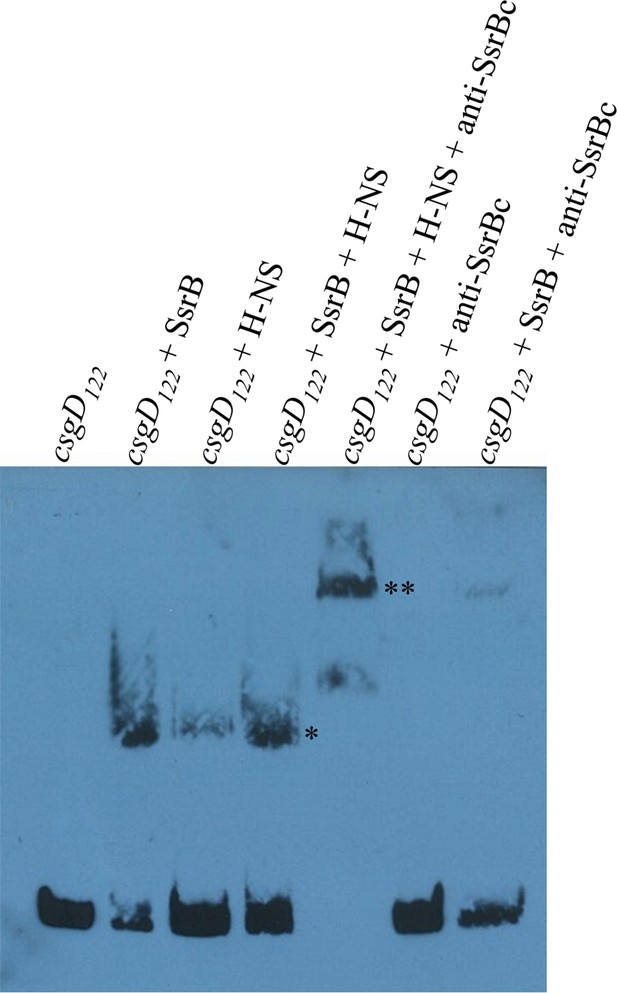
SsrB and H-NS form a complex on csgD.
Electrophoretic mobility shift assay with the 122 bp csgD regulatory region, csgD122 (left to right); in the presence of SsrB and H-NS, the DNA-protein complex (*) is super-shifted in the presence of anti-SsrBc serum (**). A DNA-protein complex is also observed when SsrB and H-NS were present alone. Note the absence of any complex in a control reaction with csgD122 and anti-SsrBc, while anti-SsrBc recognizes the SsrB-csgD122 complex.

The sequence of the 755 bp csgD regulatory region indicating the H-NS binding region according to Gerstel et al. (2003); and the SsrB binding motif as found by Feng et al. (2004).
https://doi.org/10.7554/eLife.10747.028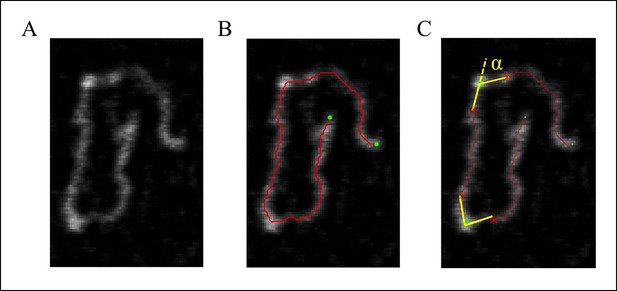
DNA tracing and bending angle measurement.
(A) The original AFM image as processed with the Gwyddion software. (B) A Matlab code was used to trace the DNA in the AFM images. The digitized binary line represents the DNA backbone (red line), with the two end points marked by green dots. (C) For the bending angle measurement, a point along the red line was manually selected as the binding location of SsrB on DNA (green asterisk). The red asterisks indicate two points that are 15 nm upstream and downstream from this protein-binding site. Linear interpolation of points scattered along the green and red asterisk was used to plot a straight line between these points (yellow line). The angle between the yellow lines (α) is determined as the bending angle of SsrB; 81 such measurements were made. The same procedure was followed for the control AFM images of csgD755, 327 such measurements were made.
Tables
List of Bacterial strains and plasmids.
| Strain | Description/Nomenclature | Reference |
|---|---|---|
| wild type | Salmonella enterica serovar Typhimurium strain 14028s | Lab strain collection |
| ssrA | ssrA::TetRA derivative of 14028s | This work |
| ssrB | DW85 | Don Walthers (originally from Stephen Libby) |
| ssaC | ssaC::TetRA derivative of 14028s | Chakraborty et al. (2015) |
| ssaJ | ssaJ::TetRA derivative of 14028s | Hideaki Mizusaki unpublished |
| D56A | D56A SsrB derivative of 14028s | This work |
| ssrAH1 | DW748 | Don Walthers unpublished |
| ssrAD | DW749 | Don Walthers unpublished |
| ssrAH2 | DW750 | This work |
| ssrAH1 sifA-LacZ | Made by transducing sifA-lacZ from DW636 to DW748 | This work |
| ssrAD sifA-lacZ | Made by transducing sifA-lacZ from DW636 to DW749 | This work |
| ssrAH2 sifA-lacZ | Made by transducing sifA-lacZ from DW636 to DW750 | Don Walthers; Lab strain collection |
| DW636 | sifA-lacZ at attP site in 14028s | Don Walthers; Lab strain collection |
| DW637 | ssrB::Km derivative of DW636 | This work/Don Walthers (lab strain collection) |
| ssrAH1 AcP | ackA-pta::Km (from DW142) transduced in DW748 | This work/Don Walthers (lab strain collection) |
| ssrAD AcP | ackA-pta::Km (from DW142) transduced in DW749 | This work/Don Walthers (lab strain collection) |
| ssrAH2 AcP | ackA-pta::Km (from DW142) transduced in DW750 | This work/Don Walthers (lab strain collection) |
| hns | hns::TetRA derivative of DW636 | This work/Don Walthers (lab strain collection) |
| hns ssrB | hns::TetRA derivative of DW637 | This work/Don Walthers (lab strain collection) |
| Plasmid pKF46 | D56A His-SsrB pMpM-A5Ω construct | Feng et al. (2004) |
| Plasmid pKF43 | His-SsrB pMpM-A5Ω construct | Feng et al. (2004) |
| Transformant DW160 | DH5α harboring His-HNS (S. typhimurium) in pMpM-A5Ω | Walthers et al. (2011) |
| Plasmid pKF104 | His-SsrBc pMpM-A5Ω construct | Feng et al. (2004) |
| pBR328::csgD | csgD construct | Prof Iñigo Lasa’s group |
| Plasmid pRC24 | K179A His-SsrB pMpM-A5Ω construct | Carroll et al. (2009) |
List of oligonucleotides.
| Purpose/name | Sequence (5’-3’) |
|---|---|
| Digf (forward 755bp csgD regulatory region) | tgatgaaactccacttttttta |
| Digr (reverse 755bp csgD regulatory region) | tgctgtcaccctggacctggtc |
| ssrA knockout (forward) | atgaatttgctcaatctcaagaatacgctgcaaacatctt ttaagacccactttcacatt |
| ssrA knockout (reverse) | agccgatacggcattttcaatatcagccagcaagaggtcc ctaagcacttgtctcctg |
| csg1 (forward csgD internal) | ggaagatatctcggccggttgc |
| csg2 (reverse csgD internal) | tcagcctagggataatcgtcag |
| rrsA1 (forward rrsA internal) | gcaccggctaactccgtgcc |
| rrsA2 (reverse rrsA internal) | gcagttcccaggttgagcccg |
| PSsrBF (forward for pKF46) | atgaaagaatataagatcttat |
| PSsrBTR (hybrid reverse for pKF46) | ttaatactctaattaacctcattcttcgggcacagttaagtctaagcacttgtctcctg |
| TSsrBF (forward TetRA-ssrB) | acttaactgtgcccgaagaatgaggttaatagagtattaattaagacccactttcacatt |
| TSsrBR (reverse TetRA- after ssrB stop) | catcaaaatatgaccaatgcttaataccatcggacgcccctggctaagcacttgtctcctg |
| Digb (Forward for EMSA) | Biotin- tgatgaaactccacttttttta |
| CsgDigRS (Reverse for EMSA) | aatatttttctctttctggata |
| hns knockout (forward) | gctcaacaaaccaccccaatataagtttggattactacattaagacccactttcacatt |
| hns knockout (reverse) | atcccgccagcggcgggattttaagcatccaggaagtaaactaagcacttgtctcctg |





