Ceapins are a new class of unfolded protein response inhibitors, selectively targeting the ATF6α branch
Figures
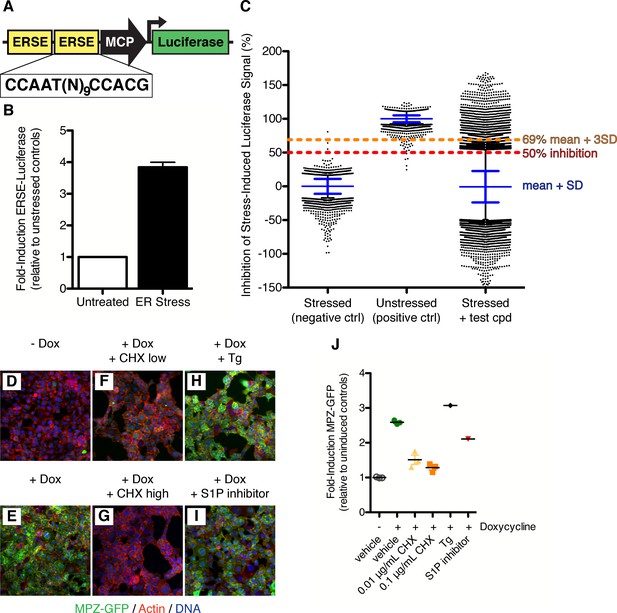
Isolation of small molecule inhibitors of ATF6 mediated transcriptional response induced by ER stress.
(A) Schematic representation of ERSE-luciferase construct used to make screening cell line. Two copies of the ER Stress Response Element (ERSE) were cloned in front of a minimal CMV promoter (MCP) driving expression of luciferase. (B) Luciferase activity is induced 3.84 ± 0.16 fold upon ER stress (100 nM Tg) in ERSE-Luciferase 293T reporter cell line. Mean of three independent experiments with at least duplicate wells is plotted; error bars are standard error of the mean. (C) Primary screen data from ERSE-luciferase transcriptional reporter cell line. Each plate was internally normalized from 0–100% inhibition using stressed and unstressed controls respectively. 106,281 compounds were added in combination with ER stressor and assayed for their ability to inhibit stress-induced production of ERSE-luciferase. Plot shows % inhibition for each control and compound tested - blue lines denote mean and standard deviation of each population, black dots indicate those wells more than two standard deviations away from the mean of the population.1142 compounds scoring more than three standard deviations from the mean (>69% inhibition, orange line) were classified as hits. (D–J). 293 cells expressing doxycycline inducible MPZ-GFP were uninduced (D) or induced with 50 nM doxycycline without (E) or with inhibitors (F–I) for seven hours and then fixed and stained for GFP (green), actin (red) and DNA (blue). Inhibitors tested were the protein synthesis inhibitor cycloheximide at either 0.01 μg/mL (F) or 0.1 μg/mL (G), the ER stressor thapsigargin (100 nM, H) or the S1P inhibitor (50 μM Pf-429242, I). (J) Mean induction of GFP per cell per image was quantified and plotted as fold induction relative to uninduced controls.
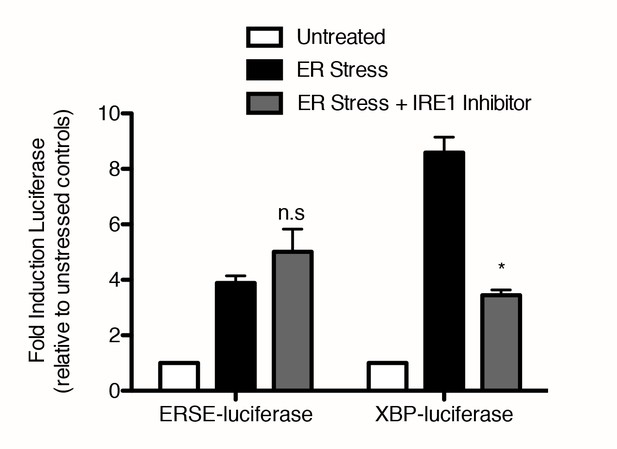
IRE1 inhibitor blocks induction of luciferase activity through XBP-luciferase but not ERSE-luciferase.
293T reporter cell lines expressing either ERSE-luciferase or XBP-luciferase were treated with either vehicle (DMSO) or ER stressor (100 nM Tg) without or with 10 μM IRE1 inhibitor for nine hours. Mean of duplicate experiments each with duplicate wells is plotted, error bars are standard error of the mean. For each cell line, ER stress ± IRE1 inhibitor were compared using two-tailed t-test – n.s non-significant, *p = 0.013.
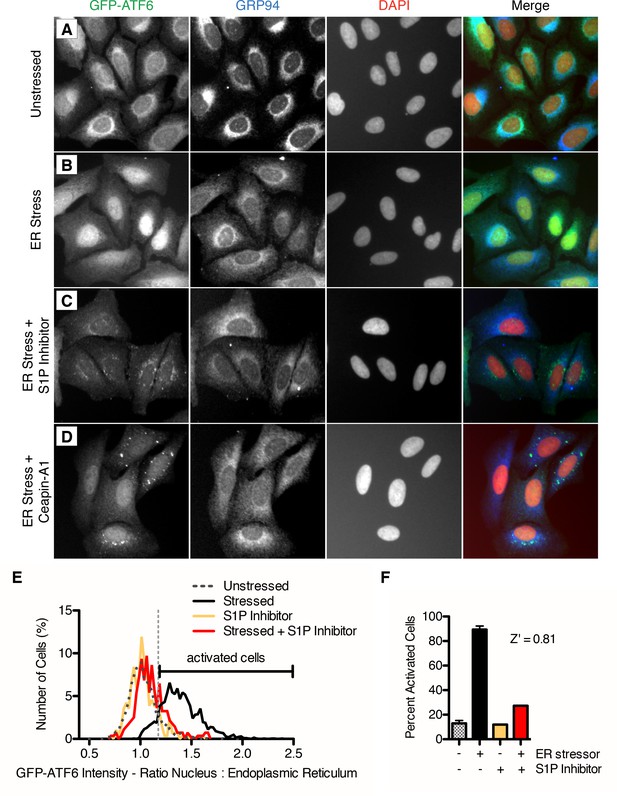
Isolation of small molecule inhibitors of ER stress induced nuclear translocation of ATF6.
(A–D) Nuclear translocation assay in U2-OS GFP-ATF6α cells. U2-OS cells expressing GFP-ATF6α were treated with either vehicle (unstressed, DMSO, A) or ER stressor in the absence (ER stress, 100 nM Tg, B) or presence of S1P inhibitor (20 μM Pf-429242, C) or screen hit (6.6 μM Ceapin-A1, D). After five hours, cells were fixed and stained for GFP (green), GRP94 (ER marker, blue) and DNA (nuclear marker, DAPI, red). (E–F) Quantification of nuclear translocation assay. (E) Single cell ratios of nuclear: ER GFP intensity were calculated for four images per well for each treatment (unstressed and stressed control wells are present seven times per plate) and plotted as histograms. For each plate, the minimum nuclear: ER ratio where the percentage of stressed cells is greater than the percentage unstressed cells is calculated and annotated as the threshold for activation by ER stress (light grey vertical dashed line). (F) For each plate, the percent activation by ER stress is calculated for the control wells (unstressed n = 1904, ER stress n = 2095, unstressed + S1P inhibitor n = 366, stressed + S1P inhibitor n = 330 cells) and used to generate a Z’ score for the plate.
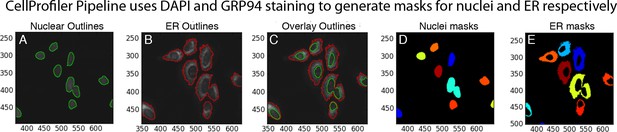

Annotation of nuclear and ER regions used for calculation of the ratio of nuclear to ER GFP-ATF6α signal for each cell.
(A–E) Calculation of nuclear: ER ratio of GFP intensity for nuclear translocation assay using CellProfiler. For each individual cell, nuclear DNA stain (DAPI, A) and ER staining (anti-GRP94, B) were used to generate outlines of the nucleus and ER respectively. These outlines were overlaid on the corresponding GFP-ATF6α image (C) and used to quantify the intensity of GFP signal in the nucleus (D) and ER (E) of each single cell in each image.
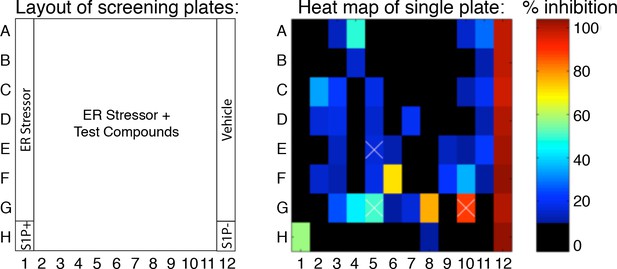
Example of a heat map for a plate from the nuclear translocation assay secondary screen showing percent inhibition of test compounds compared to controls.
Example of the heat map generated for each plate showing the percent inhibition of each compound tested. In this presentation, wells in column 1 and 12 represent stressed and unstressed cells, respectively, with H1 and H12 containing the S1P inhibitor. Columns 2–11 contain stressed cells treated with test compounds. 0% and 100% inhibition are set by the controls - unstressed wells (100%, vehicle, A12-G12) and stressed wells (0%, A1-G1) respectively. S1P inhibitor alone (H12) or in combination with ER stressor (H1) was used as the positive control in each plate. Wells G8, F6, and A4 represent the top three hits on this plate. Wells containing less than 50% the number of cells of ER stressed controls were annotated as toxic with white X signs – wells E5, G5 and G10.
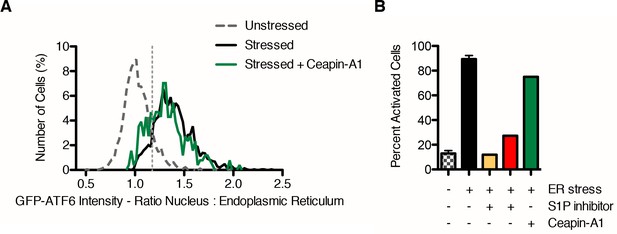
Ceapin-A1 inhibits nuclear translocation of GFP-ATF6.
(A) Single cell ratios of nuclear: ER GFP intensity from four images per well for Ceapin-A1, unstressed and stressed controls from a single plate are plotted as histograms. (B) The percent activation by ER stress is for the control wells (unstressed n = 1904, ER stress n = 2095, unstressed + S1P inhibitor n = 366, stressed + S1P inhibitor n = 330 cells) and for Ceapin-A1 (stressed + Ceapin-A1 n = 256 cells) from a single plate. From three independent plates, nuclear translocation of GFP-ATF6α by Ceapin-A1 was inhibited by 35.07 ± 18.15%. Note that in this assay, compounds were screened at 6.6 μM, which is below the IC50 for the Ceapin-A1 stock in the library plate (8.49 μM).
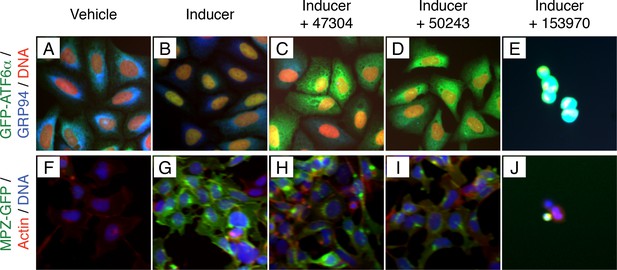
Combining data from high content assays identified non-specific inhibitors of trafficking and toxic compounds.
(A–E) Nuclear translocation assay showing unstressed (A) and ER stressed (B) GFP-ATF6α U2-OS cells. GFP-ATF6α is green, GRP94 is blue and DNA is in red. Two compounds with similar scaffold (47304, C and 50243, D) scored as inhibitors of ATF6 processing. (E) 153970 is an example of a toxic compound. (F–J) Inducible GFP assay showing uninduced (F) and induced (G) MPZ-GFP T-Rex cells. MPZ-GFP is in green, actin is in red, DNA is in blue. Both compounds identified as inhibitors of ATF6 processing also show defects in trafficking of MPZ-GFP from the ER to the plasma membrane (47304, H and 50243, I) and are unlikely to be selective inhibitors of ATF6. (J) 153970 is also toxic to this cell line.
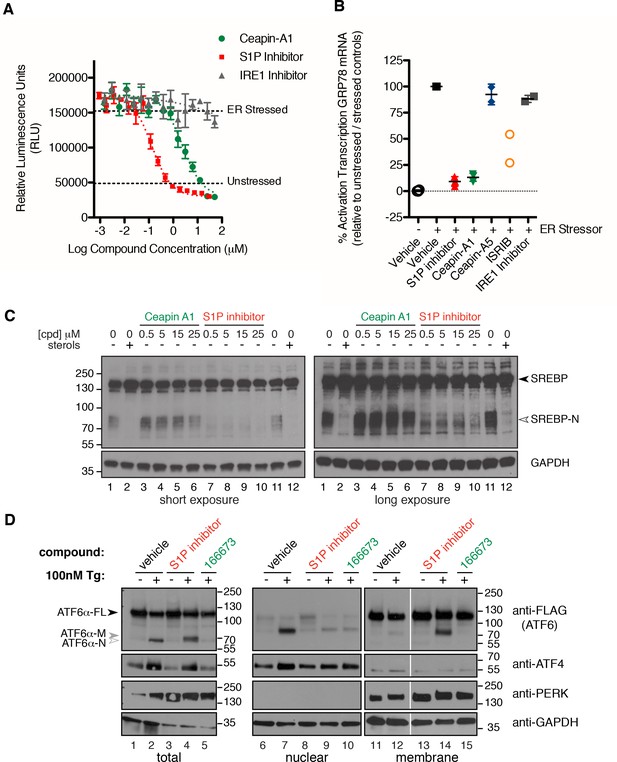
Isolation of Ceapin-A1, a small molecule inhibitor of ATF6 but not SREBP processing.
(A) ERSE-luciferase assay in HEK293T cells. Cells were treated without (DMSO) or with ER stressor (100 nM Tg) in the presence or absence of inhibitors for nine hours. Increasing concentrations of either S1P inhibitor (Pf-429242, red) or Ceapin-A1 (green) but not IRE1 inhibitor (4 μ8C, grey) block ER stress-induced luciferase activity. Plotted is one representative experiment showing mean and standard deviation for each inhibitor concentration (triplicate wells per point). Dashed grey lines indicate the relative luciferase activity of unstressed and stressed controls. (B) ER stress induced upregulation of the endogenous ATF6α target gene GRP78 in U2-OS cells. Cells were treated without (DMSO, open circles) or with ER stress (100 nM Tg, black squares) in the absence or presence of inhibitors for four hours prior to isolation of mRNA. Upregulation of GRP78 mRNA was measured using qPCR. mRNA levels for GRP78 were normalized to GAPDH for each well and then compared to unstressed and stressed controls. ER stress induced GRP78 mRNA induction is inhibited by co-incubation with either S1P inhibitor (2.3 μM Pf-429424, red) or Ceapin-A1 (10 μM, green) but not the inactive Ceapin analog A5 (10 μM, blue). Inhibition of the ISR (200 nM or 400 nM ISRIB, orange) partially inhibits GRP78 induction while inhibition of IRE1 (10 μM 4 μ8C, grey) has only minor effects. Data plotted is the mean percent activation of GRP78 transcription relative to unstressed (0%) and stressed (100%) controls from two or three independent experiments, each with duplicate reactions carried out on duplicate wells. (C) Induction of SREPB processing by lipoprotein depletion in HeLa cells. HeLa cells were grown in lipoprotein deficient media for 16.5 hr prior to addition of either sterols or inhibitors for five hours. One hour prior to lysis proteasome inhibitor (25 μg/mL ALLN) was added to prevent the degradation of the cleaved SREBP-N fragment. Whole cell lysates were analyzed by Western blotting for SREPB1 and GAPDH. Arrowheads denote positions of full-length (SREBP) and cleaved (SREBP-N) variants of SREBP1. Lipoprotein depletion induces cleavage of SREBP (lanes 1, 11) that is inhibited by addition of sterols (10 μg/mL cholesterol, 1 μg/mL 25-hydroxycholesterol, lanes 2, 12) or increasing concentrations of a S1P inhibitor (Pf-429242, lanes 7–10) but not increasing concentrations of Ceapin-A1 (lanes 3–6). Data shown is representative of three independent experiments. (D) Induction of ATF6α processing by ER stress in T-Rex cells expressing FLAG-tagged ATF6α. Arrowheads denote positions of full-length (ATF6α), cleaved membrane-bound (ATF6α-M) and cleaved nuclear (ATF6α-N) variants of ATF6. Cells were treated without (lanes 1,6,11) or with (lanes 2,7,12) ER stressor (100 nM Tg) alone or in combination with either S1P inhibitor (0.75 μM Pf-429242, lanes 3,4,8,9,13,14) or Ceapin-A1 (14.95 μM, lanes 5,10,15) for two hours prior to harvesting. One hour prior to lysis proteasome inhibitor (MG132, 10 μM) was added. Cells were harvested and separated by centrifugation into total, membrane and nuclear fractions and analyzed by Western blot for ATF6α (anti-FLAG), PERK (membrane control), ATF4 (nuclear control) and GAPDH (loading control). Note that totals were run on 10% gels while membrane and nuclear fractions were run on gradient gels to visualize the migration differences between ATF6α-N and ATF6α-M and between PERK and phosphorylated PERK respectively. Data shown is representative of two independent experiments.
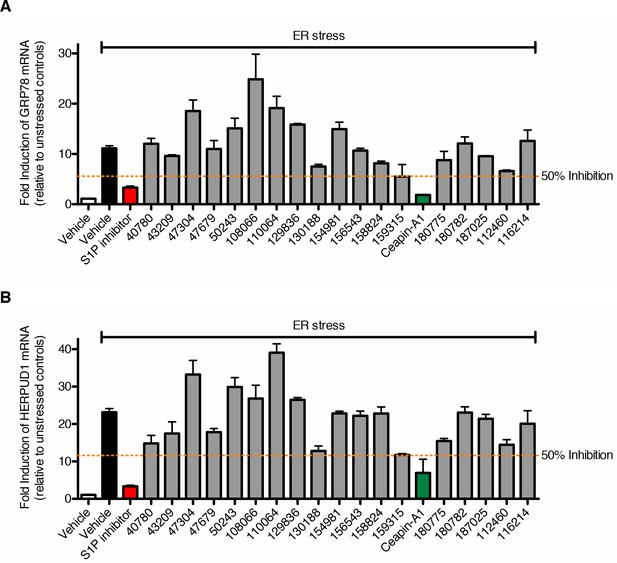
Identification of Ceapin-A1, a small molecule that inhibits ATF6 processing in response to ER stress.
(A–B) qPCR analysis of endogenous ATF6α target genes GRP78 (A) and HERPUD1 (B). U2-OS cells were treated without (DMSO) or with ER stressor (100 nM Tg) alone or in combination with ATF6 processing hits (either 10 x IC50 or 10 μM, depending on solubility) for four hours prior to harvesting and mRNA extraction. Fold induction of each target gene is calculated relative to unstressed controls. Dashed orange lines denote 50% inhibition. Data plotted are means and standard deviations from duplicate reactions from duplicate wells. It is unclear why so many compounds failed at this point having passed through earlier filters. These results validate the effects of the Ceapin-A1 on endogenous gene expression and so based on these results we focused exclusively on the Ceapin series.
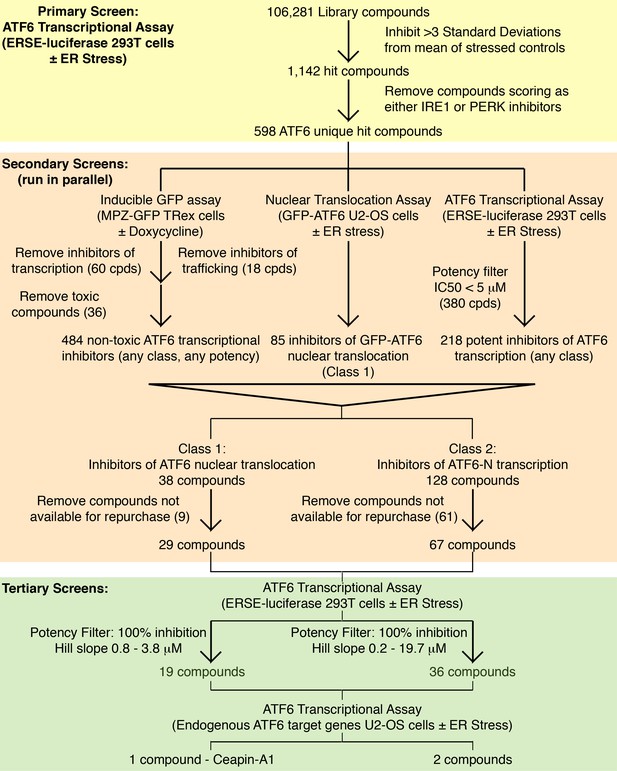
Screening workflow Summary of screening workflow that lead to the identification of Ceapin-A1 consisting of primary (yellow), secondary (orange) and tertiary (green) screens.
The primary screen was run on the entire library, the secondary screens on cherry-picked compounds from the primary screen and the tertiary screens on repurchased material. At each stage compounds were triaged and filtered according to the criteria described.
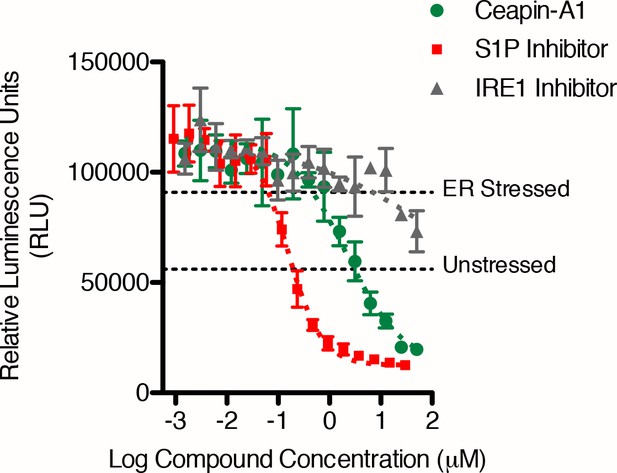
Ceapin-A1 inhibits ER stress induced ERSE-luciferase activity ERSE-luciferase assay in HEK293T cells.
Cells were treated without (DMSO) or with ER stressor (2 μg/mL Tm) in the presence or absence of inhibitors for ten hours. Increasing concentrations of either S1P inhibitor (Pf-429242, red) or Ceapin-A1 (green) but not IRE1 inhibitor (4μ8C, grey) block ER stress-induced luciferase activity. Plotted is a representative experiment showing mean and standard deviation for each inhibitor concentration (triplicate wells per point). Dashed grey lines indicate the relative luciferase activity in unstressed and stressed controls.
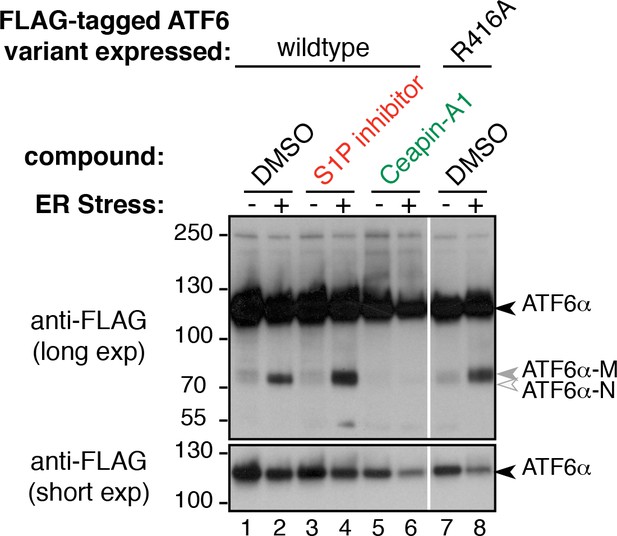
Mutation of S1P cleavage site in ATF6α leads to production of ATF6α-M upon ER stress.
Induction of ATF6α processing by ER stress in T-Rex cells expressing FLAG-tagged ATF6α either wild-type or with S1P cleavage site mutated (R416A). Arrowheads denote positions of full-length (ATF6α-FL), cleaved membrane bound (ATF6α-M) and cleaved nuclear (ATF6α-N) variants of ATF6α. Cells were treated without (lanes 1,7) or with (lanes 2,8) ER stressor (100 nM Tg) alone or in combination with either S1P inhibitor (0.75 μM Pf-429242, lanes 3,4) or Ceapin-A1 (15 μM, lanes 5,6) for two hours prior to harvesting. Cells were harvested and analyzed by Western blot for ATF6 (anti-FLAG). White line indicates where intervening lanes have been removed. Data shown is representative of duplicate experiments.
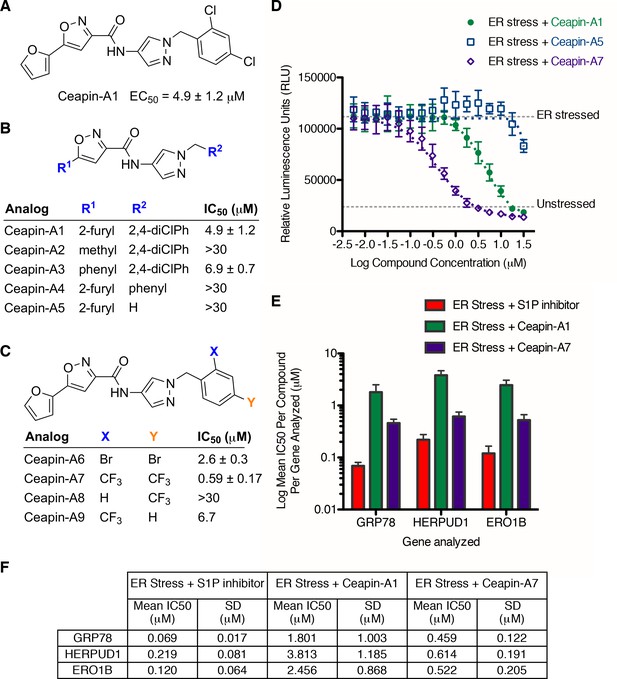
SAR studies of Ceapin-A1 improve potency by an order of magnitude.
(A–C) Summary of structure activity relationship for Ceapin analogs. (A) Chemical structure of the initial screen hit, Ceapin-A1. SAR of rings (B) or substituents on bis-substituted phenyl ring (C) of Ceapin scaffold. IC50 values were obtained using ERSE-luciferase assay in 293T cells where compounds were tested in dose-response in combination with ER stressor (100 nM Tg). IC50 values are from at least four independent experiments for each compound. (D) ERSE-luciferase assay showing improved potency of Ceapin-A7 (purple) and lack of activity of Ceapin-A5 (blue) compared to Ceapin-A1 (green). 293T cells with stably integrated ERSE-luciferase reporter were treated with ER stressor (100 nM Tg) and increasing concentrations of Ceapin analogs for nine hours prior to reading luciferase activity. Data plotted are mean values from a representative experiment with each point done in triplicate, error bars are standard deviation. (E) qPCR analysis of endogenous ATF6α target genes. U2-OS cells were treated without or with ER stressor in the presence of increasing concentrations of Ceapin analogs for four hours prior to harvesting of mRNA for qPCR analysis. mRNA levels for GRP78, HERPUD1 and ERO1B were normalized to GAPDH for each well and then compared to unstressed controls. Data plotted are mean IC50 values calculated from duplicate experiments, each with duplicate qPCR reactions from duplicate wells for S1P inhibitor (Pf-429242, red), Ceapin-A1 (green) and Ceapin-A7 (purple). Error bars are standard deviation. (F) Calculated mean IC50 values and standard deviations from qPCR analysis described above.
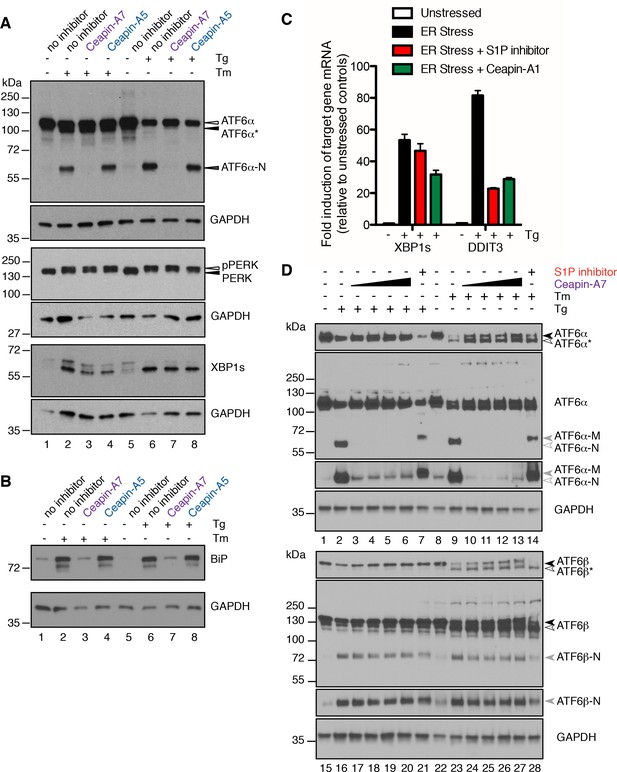
Ceapins are selective inhibitors of the ATF6α branch of the UPR.
(A) U2-OS cells were treated without (DMSO) or with ER stressor (100 nM Tg or 2.5 μg/mL Tm) in the absence or presence of Ceapin analogs (6 μM each) for two hours. One hour prior to lysis, proteasomal inhibitor (10 μM MG132) was added to prevent the degradation of the cleaved ATF6α-N fragment. Cells were harvested and analyzed by Western Blot for ATF6α, PERK, XBP1 and GAPDH (loading control). Arrowheads denote the positions of full-length (ATF6α), unglycosylated full-length (ATF6α*) and cleaved (ATF6α-N) variants of ATF6α and also PERK and phospho-PERK. Data shown is representative of three independent experiments. (B) U2-OS cells were treated without (DMSO) or with ER stressor (100 nM Tg or 2.5 μg/mL Tm) in the absence or presence of Ceapin analogs (6 μM each). After eight hours, cells were harvested and analyzed by Western Blot for BiP and GAPDH (loading control). (C) U2-OS cells were treated without (DMSO) or with ER stressor (100 nM Tg, black) in the absence or presence of ten-fold the IC50 of either S1P inhibitor (Pf-429242, 3.2 μM, red) or Ceapin-A1 (35.7 μM, green). Four hours later cells were harvested and mRNA extracted. mRNA levels for XBP1s or DDIT3 were normalized to GAPDH for each well and then compared to unstressed controls. Data plotted are from duplicate qPCR reactions from duplicate wells, error bars are standard deviation. (D) U2-OS cells were treated without (DMSO) or with ER stressor (100 nM Tg or 2.5 μg/mL Tm) in the absence or presence of increasing concentration of Ceapin-A7 (0.6, 1.89, 6, 18.9 μM) or S1P inhibitor (5 μM Pf-429242) for four and a half hours. One hour prior to lysis, proteasomal inhibitor (10 μM MG132) was added to prevent the degradation of the cleaved ATF6α-N and ATF6β-N fragments. Cells were harvested and analyzed by Western Blot for ATF6α, ATF6β and GAPDH (loading control). Arrowheads denote the positions of full-length (ATF6α, ATF6β), unglycosylated full-length (ATF6α*, ATF6β*), cleaved membrane bound (ATF6α-M) and cleaved (ATF6α-N, ATF6β-N) variants of ATF6α and ATF6β. Data shown is representative of three independent experiments.
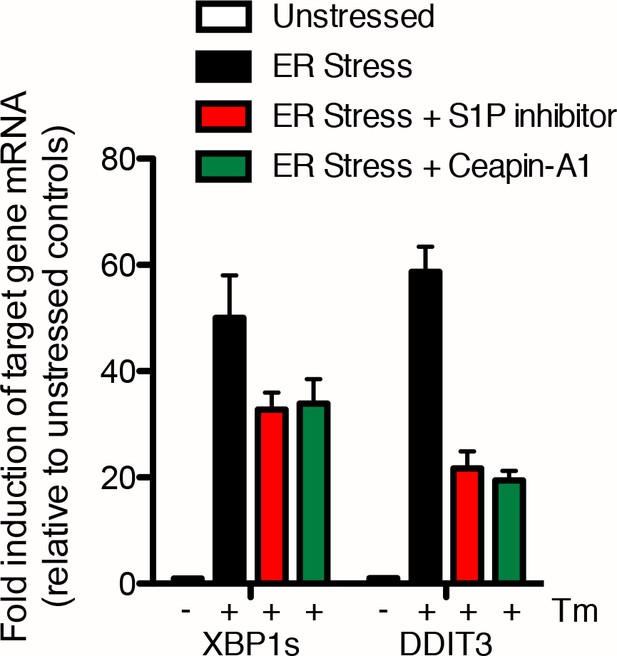
Induction of XBP1s and DDIT3 mRNA is only partially inhibited by either the S1P inhibitor or Ceapin-A1 U2-OS cells were treated without (DMSO) or with ER stressor (2.0 μg/mL Tm, black) in the absence or presence of ten-fold the IC50 of either S1P inhibitor (Pf-429242, 3.2 μM, red) or Ceapin-A1 (35.7 μM, green).
Four hours later cells were harvested and mRNA extracted. mRNA levels for XBP1s or DDIT3 were normalized to GAPDH for each well and then compared to unstressed controls. Data plotted are from duplicate qPCR reactions from duplicate wells, error bars are standard deviation.
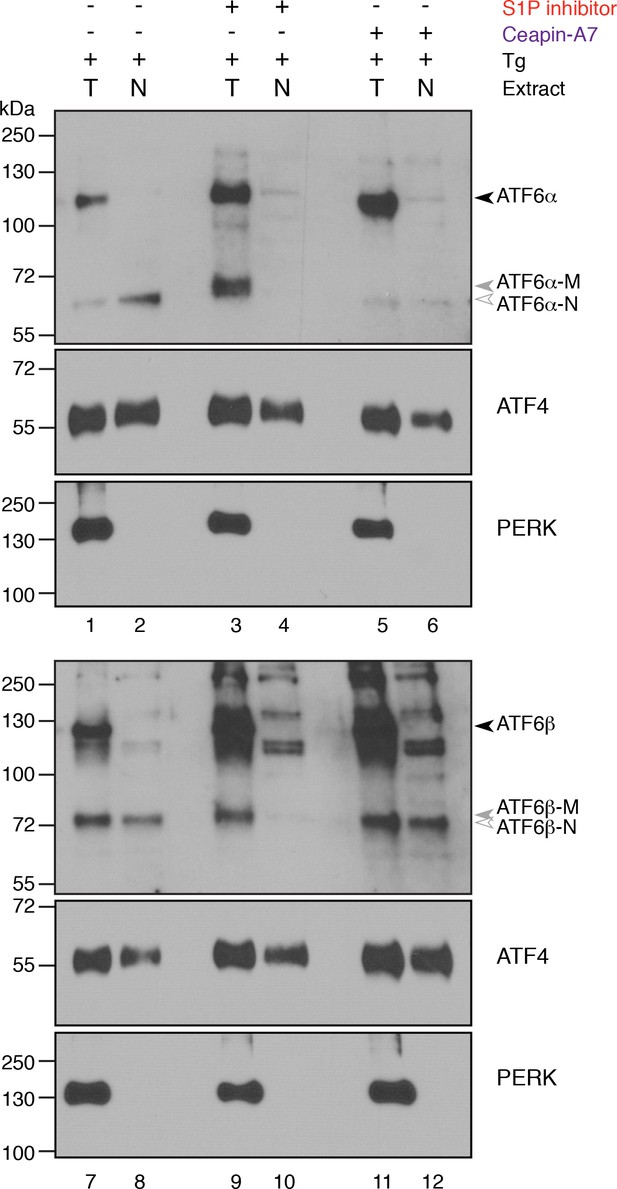
ATF6β-N is generated in ER stressed cells treated with Ceapin-A7 but not ER stressed cells treated with the S1P inhibitor.
U2-OS cells were treated with ER stressor (100 nM Tg) in the absence (lanes 1, 2, 7 and 8) or presence of S1P inhibitor (5 μM Pf-429242, lanes 3, 4, 9 and 10) or Ceapin-A7 (6 μM, lanes 5, 6, 11 and 12) for four hours. One hour prior to lysis, proteasomal inhibitor (10 μM MG132) was added to prevent the degradation of the cleaved ATF6α-N and ATF6β-N fragments. Cells were harvested and separated by centrifugation into total (lanes 1, 3, 5, 7, 9 and 11) and nuclear fractions (lanes 2, 4, 6, 8, 10 and 12) and analyzed by Western blot for ATF6α, ATF6β, PERK and ATF4. Arrowheads denote the positions of full-length (ATF6α, ATF6β), cleaved membrane bound (ATF6α-M) and cleaved (ATF6α-N, ATF6β-N) variants of ATF6α and ATF6β. The transcription factor ATF4 is present in both total and nuclear extracts while membrane proteins PERK, ATF6α and ATF6β are present only in total extracts. In ER stressed cells faster migrating bands corresponding to ATF6α-N (lanes 1 and 2) and ATF6β-N (lanes 7 and 8) are found in both total and nuclear fractions. In ER stressed cells treated with the S1P inhibitor the faster migrating ATF6α and ATF6β bands found in the total extract are not present in the nuclear extract (compare lanes 3 and 4 for ATF6α and lanes 9 and 10 for ATF6β) indicating that neither ATF6α-N nor ATF6β-N were produced and these bands are likely ATF6α-M and ATF6β-M. In ER stressed cells treated with Ceapin-A7 ATF6α is not cleaved and no ATF6α-N is found in the nuclear extract (lanes 5 and 6). In contrast, ATF6β is cleaved and this faster migrating band is found in the nuclear extract indicating it is ATF6β-N (lanes 11 and 12). Data shown is representative of three independent experiments.
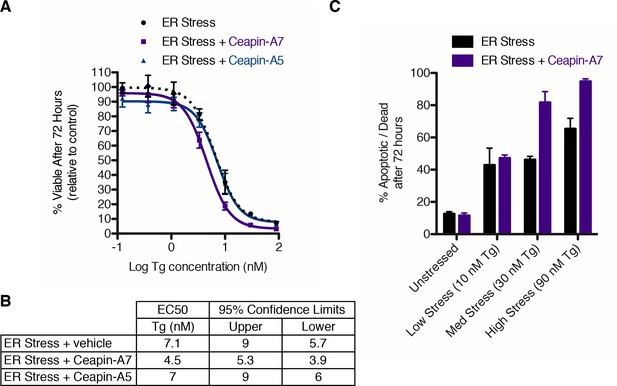
Ceapin-A7 sensitizes cells to ER stress.
(A–B) U2-OS cells were treated with increasing concentrations of ER stressor (Tg) in the absence (black) or presence of six micromolar Ceapin analogs - Ceapin-A5 (inactive, blue) or Ceapin-A7 (purple). (A) After seventy-two hours reducing potential of living cells was assayed to determine cell viability. Data plotted are the means of four independent experiments performed in triplicate, error bars represent the standard error of the mean. (B) EC50 values calculated for ER stressor in the absence or presence of Ceapin analogs showing mean and 95% confidence limits. (C). U2-OS cells were treated with increasing concentrations of ER stressor (Tg) in the absence (black) or presence of 6 μM Ceapin-A7 (purple). To analyze cell death, cells were stained with Annexin V and 7AAD and analyzed by flow cytometry. Data plotted are the means from three independent experiments performed in duplicate; error bars represent standard deviation.
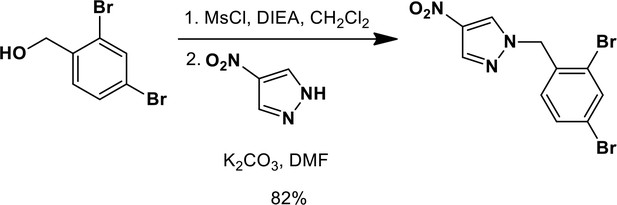
1-{[2,4-Dibromophenyl]methyl}-4-nitro-1H-pyrazole.
https://doi.org/10.7554/eLife.11878.020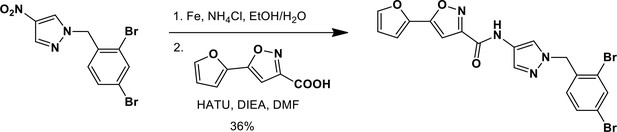
N-(1-{[2,4-Dibromophenyl]methyl}-1H-pyrazol-4-yl)-5-(furan-2-yl)-1,2-oxazole-3-carboxamide (Ceapin-A6).
https://doi.org/10.7554/eLife.11878.021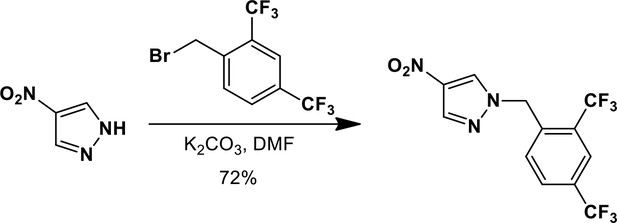
1-{[2,4-Bis(trifluoromethyl)phenyl]methyl}-4-nitro-1H-pyrazole.
https://doi.org/10.7554/eLife.11878.022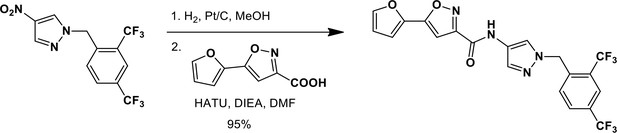
N-(1-{[2,4-bis(trifluoromethyl)phenyl]methyl}-1H-pyrazol-4-yl)-5-(furan-2-yl)-1,2-oxazole-3-carboxamide (Ceapin-A7).
https://doi.org/10.7554/eLife.11878.023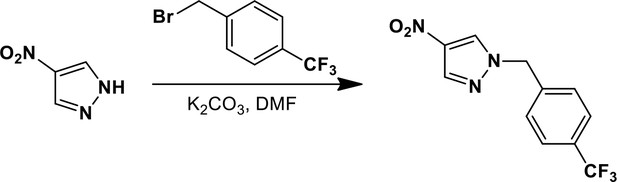
1-{[4-(Trifluoromethyl)phenyl]methyl}-4-nitro-1H-pyrazole.
https://doi.org/10.7554/eLife.11878.024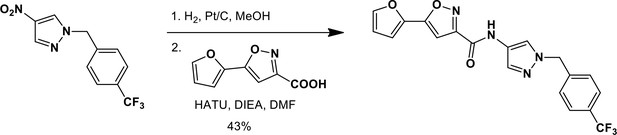
N-(1-{[4-(Trifluoromethyl)phenyl]methyl}-1H-pyrazol-4-yl)-5-(furan-2-yl)-1,2-oxazole-3-carboxamide (Ceapin-A8).
https://doi.org/10.7554/eLife.11878.025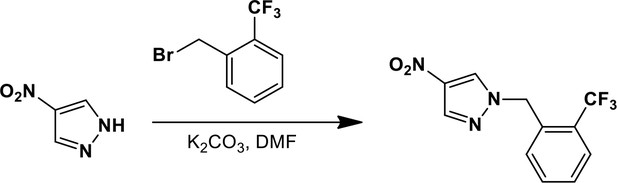
1-{[2-(Trifluoromethyl)phenyl]methyl}-4-nitro-1H-pyrazole.
https://doi.org/10.7554/eLife.11878.026