AXL receptor tyrosine kinase is required for T cell priming and antiviral immunity
Figures
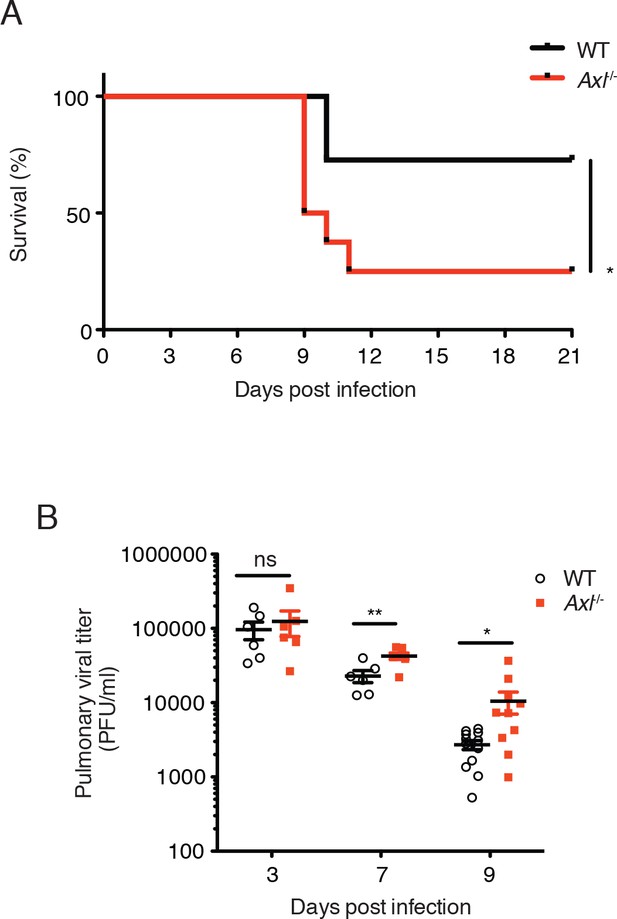
Loss of Axl increases susceptibility to influenza A virus infection in vivo.
(A) Kaplan-Meier survival curves for wild-type (WT) and Axl-/- mice infected with 10 PFU of A/PR8 virus, 8–11 mice of each genotype and representative of 5 independent experiments. (B) Viral titers in the bronchoalveolar lavage (BAL) of WT and Axl-/- mice on days 3, 7, and 9 post infection with 10 PFU of PR8, as determined by qPCR of PR8 polymerase acidic protein (PA) RNA. PFU = plaque forming units. 6–12 mice were used per condition. ns, non-significant; *p<0.05; **p<0.01.
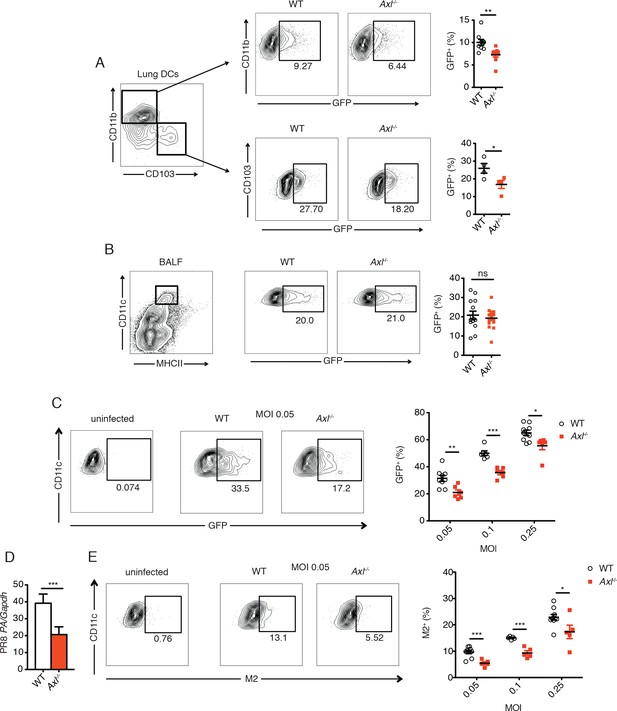
Genetic ablation of Axl confers resistance to IAV infection in dendritic cells in vivo and in vitro.
WT and Axl-/- mice were infected with 3x106 PFU of PR8-GFP for 72 hr and lung DCs were identified by flow cytometry. (A) Top, representative flow cytometry plots (left) and percentage of GFP+CD11c+MHCII+CD11b+ DCs (right) in infected WT and Axl-/- mice. n = 9 for each genotype, representing 3 independent experiments. Bottom, representative plots (left) and percentage of GFP+CD11c+MHCII+CD103+ DCs (right) in infected WT and Axl-/- mice. n = 4 for each genotype, representative of 3 independent experiments. (B) Representative flow cytometry plots (left) and percentage of GFP+ alveolar macrophages (right) in infected WT and Axl-/- mice. 14–16 mice per genotype, 3 independent experiments. (C) WT and Axl-/- BMDCs were infected with PR8-GFP with indicated multiplicities of infection (MOIs) for 12 hr. Representative flow cytometry plots (left) and percentage of GFP+ BMDCs (right) are shown. (D) Abundance of PR8 PA RNA normalized to Gapdh in WT and Axl-/- BMDCs after 12 hr of infection with 0.25 MOI of PR8-GFP, as determined by qPCR. (E) WT and Axl-/- BMDCs were infected as in (C). Representative plots (left) and percentage of IAV M2 ion channel+ BMDCs (right) are shown. For (C) and (E), 5–9 samples were tested in each condition. Data are shown as representative or as the mean ± SEM of at least 4 independent samples per group representative of 4 independent experiments. ns, non-significant; *p<0.05; **p<0.01; ***p<0.001.
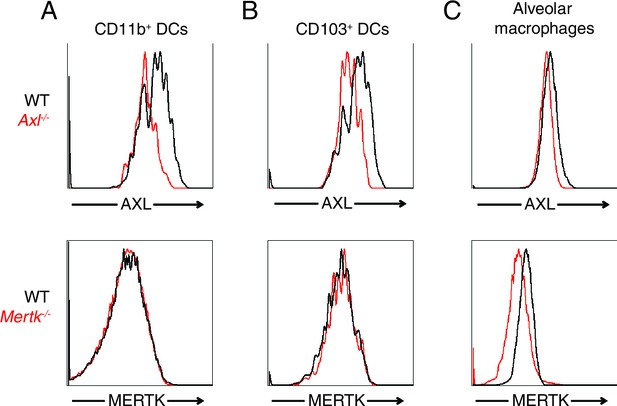
AXL and MERTK expression in naive lung dendritic cells and alveolar macrophages.
Representative histograms showing AXL (top) and MERTK (bottom) expression in (A) CD11c+MHCII+CD11b+CD103- lung DCs, (B) CD11c+MHCII+CD11b-CD103+ lung DCs, and (C) CD11chighMHCIIint alveolar macrophages.
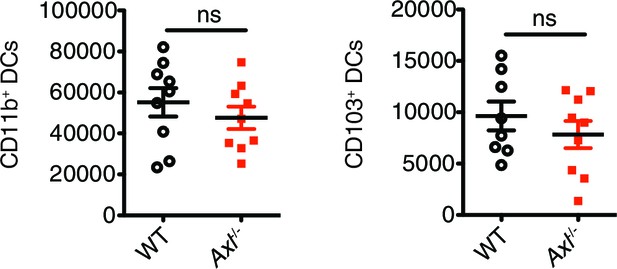
Total number of CD11c+MHCII+CD11b+CD103- and CD11c+MHCII+CD11b-CD103+ cells in the lung 72 hr post infection with 3x106 PFU A/PR8 NS1-GFP.
Data are shown as the mean ± SEM of independent experiments, n = 8–9 of each genotype. ns, non-significant.
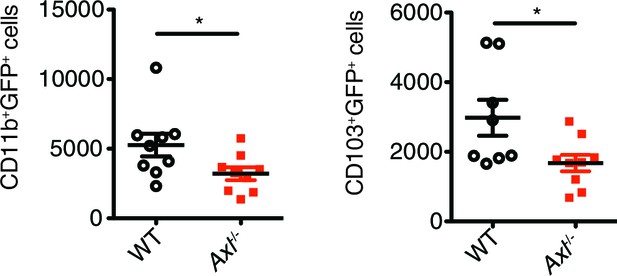
Axl-/- mice have fewer IAV-infected lung DCs than WT mice.
Total number of GFP+CD11c+MHCII+CD11b+CD103- and GFP+CD11c+MHCII+CD11b-CD103+ cells in the lung 72 hr post infection with 3x106 PFU A/PR8 NS1-GFP. Data are shown as the mean ± SEM, n = 8–9 of each genotype. *p<0.05.
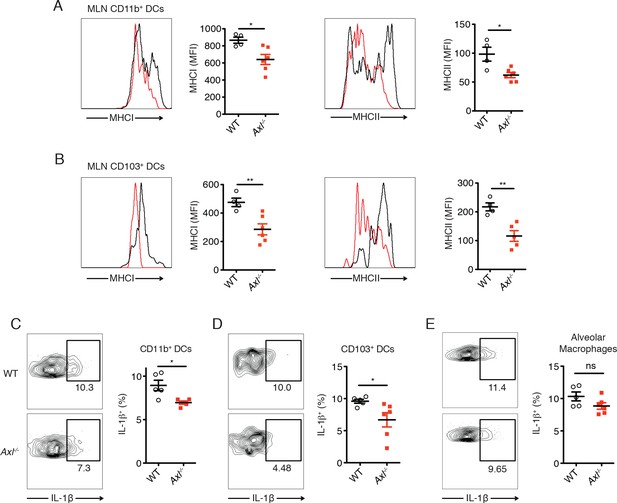
DCs in Axl-/- mice are less activated and produce less IL-1β than WT mice during IAV infection.
(A) Expression of MHC-I and MHC-II molecules on CD11c+MHCII+CD11b+CD103- mediastinal lymph node (MLN) DCs after 72 hr of infection with 3x106 PFU of PR8-GFP IAV as detected by flow cytometry. (B) Expression of MHC-I and MHC-II molecules on CD11c+MHCII+CD11b-CD103+ MLN DCs in mice infected as in (A). (C) Intracellular staining of IL-1β in lung CD11b+ DCs 72 hr post infection with 3x106 PFU of PR8-GFP. (D) Intracellular staining of IL-1β in lung CD103+ DCs infected as in (C). Data are presented as the mean ± SEM of 4–6 mice per condition, representative of 2–4 independent experiments. ns, non-significant; *p<0.05; **p<0.01.
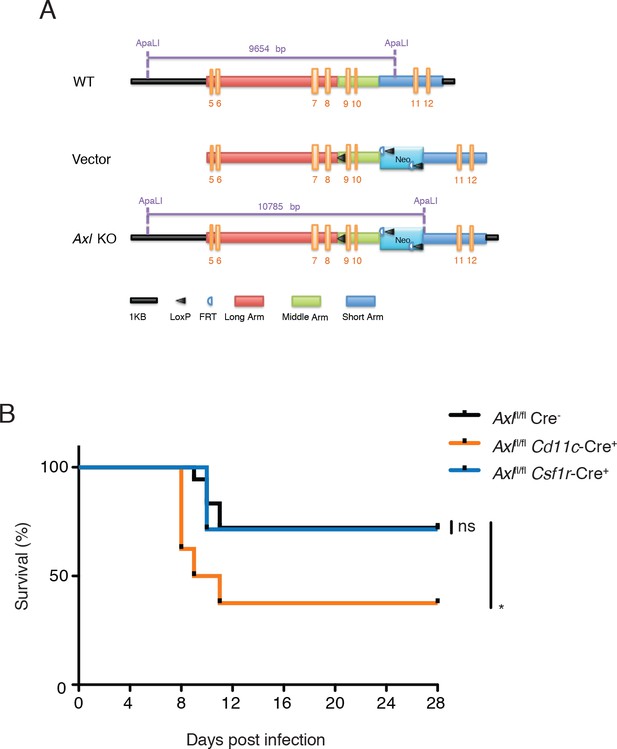
Cd11c-Cre+Axlfl/fl mice but not Csf1r-Cre+Axlfl/fl mice succumb to IAV infection.
(A) Cloning strategy for the generation of Axl-floxed mice. Axlfl/fl mice were subsequently crossed with Cd11c-Cre or Csf1r-Cre mice. (B) Kaplan-Meier survival curves for Cre-Axlfl/fl, Cd11c-Cre+Axlfl/fl, and Csf1r-Cre+Axlfl/fl mice infected with 10 PFU of A/PR8 virus, 7–18 mice per group and representative of 2 independent experiments. ns, non-significant; *p<0.05.
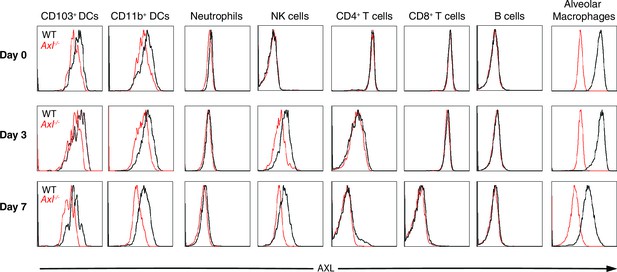
AXL expression by immune cells in the lung during IAV infection.
Representative histograms of AXL expression in CD11c+MHCII+CD11b-CD103+ DCs, CD11c+MHCII+CD11b+CD103- DCs, CD11c-Ly6g+ neutrophils, CD11c-Ly6g-NK1.1+DX5+ NK cells, CD11c-CD4+ T cells, CD11c-CD8+ T cells, CD11c-CD11b-B220+ B cells, and CD11chighMHCIIint alveolar macrophages. Samples were collected from naïve mice or those infected with 10 PFU of PR8 for 3 days or 7 days, as indicated. Histograms are representative of 6–20 mice from 2–5 independent experiments.
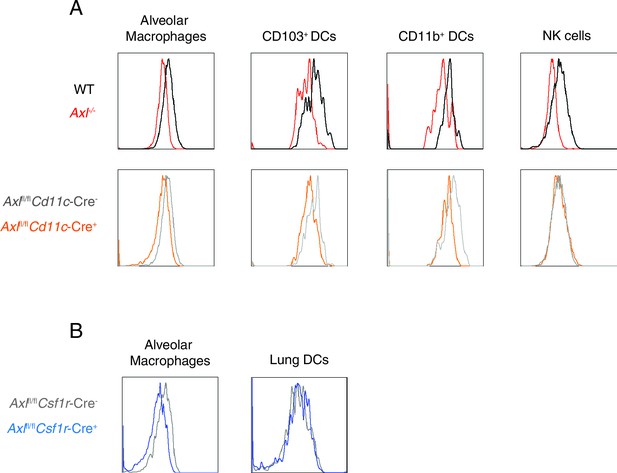
AXL is selectively ablated in Cd11c-Cre+Axlfl/fl and Csf1r-Cre+Axlfl/fl mice.
(A) AXL expression in lung C CD11chigh MHCIIint alveolar macrophages, CD11c+MHCII+CD11b-CD103+ DCs, CD11c+MHCII+CD11b+CD103- DCs, and CD11c-Ly6g-NK1.1+DX5+ NK cells from WT, Axl-/-, Cd11c-Cre-Axlfl/fl and Cd11c-Cre+Axlfl/fl mice. Histograms are representative of 4–5 mice per genotype from 2–5 independent experiments. DC and alveolar macrophage AXL expression is shown from naive mice and NK cell AXL expression is represented from mice infected with 10 PFU of PR8 for 7 days. (B) AXL expression in lung CD45+CD11c+CD115+Siglec F+CD11blo alveolar macrophages and CD45+CD11c+CD115-Siglec F- DCs from Csf1r-Cre-Axlfl/fl and Csf1r-Cre+Axlfl/fl mice. Histograms are representative of 4–5 mice per genotype from 2 independent experiments.
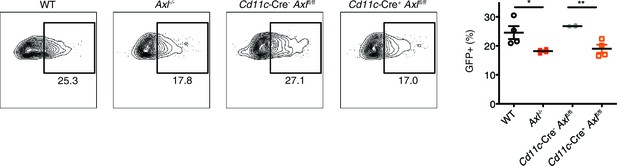
Cd11c-Cre+Axlfl/fl BMDCs are resistant to IAV infection.
WT, Axl-/-, Cd11c-Cre-Axlfl/fl, and Cd11c-Cre+Axlfl/fl BMDCs were infected with 0.05 MOI of PR8-GFP for 12 hr. Representative flow cytometry plots (left) and percentage of GFP+ BMDCs (right) are shown. Data are shown as representative or as the mean ± SEM of 4 independent samples per genotype from 2 independent experiments. *p<0.05; **p<0.01
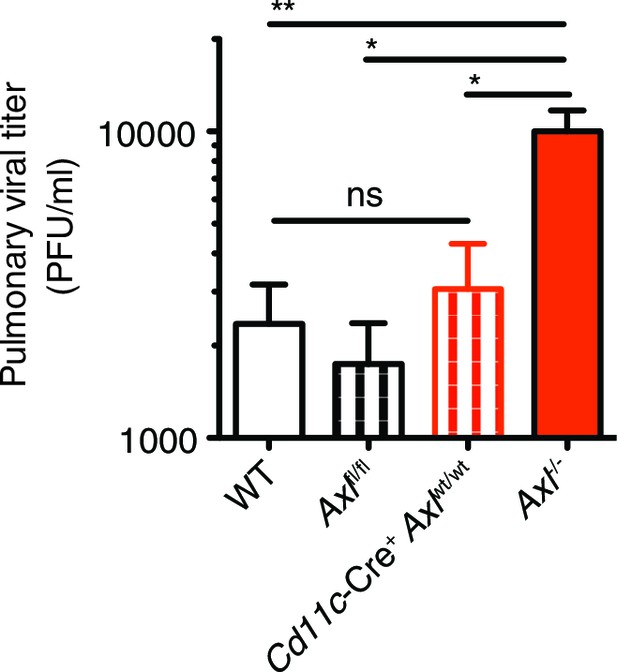
Cre-Axlfl/fl and Cd11c-Cre+Axlwt/wt mice clear IAV infection.
Viral titers in the bronchoalveolar lavage (BAL) of WT, Cre-Axlfl/fl, Cd11c-Cre+Axlwt/wt, and Axl-/- mice 9 days post infection with 10 PFU of PR8. Data represents 5 mice per condition. ns, non-significant; *p<0.05; **p<0.01
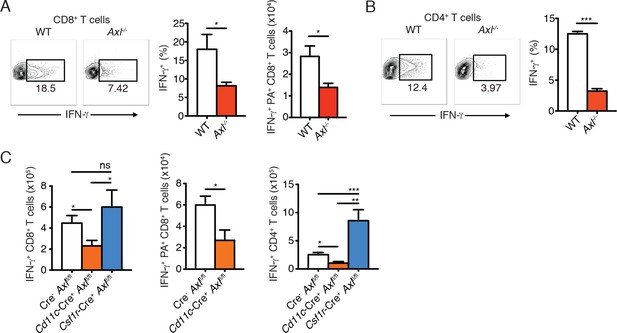
Axl-/- mice and Cd11c-Cre+Axlfl/fl mice mount impaired T cell responses to IAV infection.
(A) Representative plots (left) and percentage (middle) of CD8+IFN-γ+ T cells in the lung of WT and Axl-/- mice after 9 days of infection with 10 PFU of PR8. 4–5 mice per genotype, representative of 4 independent experiments. Right, quantification of IFN-γ-producing H-2Db-restricted CD8+ T cells specific for IAV PA amino acids 224–233 in the lung 9 days post infection with 10 PFU of PR8. 7–8 mice per genotype, 2 independent experiments. (B) Representative plots (left) and percentage (right) of CD4+IFN-γ+ T cells in the lung of WT and Axl-/- mice infected as in (A). (C) Number of CD8+IFN-γ+ T cells (left), CD8+PA+IFN-γ+ T cells (middle), and CD4+IFN-γ+ (right) in the lung 9 days post-infection with 10 PFU of PR8 in Cre-Axlfl/fl, Cd11c-Cre+Axlfl/fl, and Csf1r-Cre+Axlfl/fl mice, as indicated. 5–10 mice per genotype, representative of 2–3 independent experiments. ns, non-significant; *p<0.05; **p<0.01; ***p<0.001
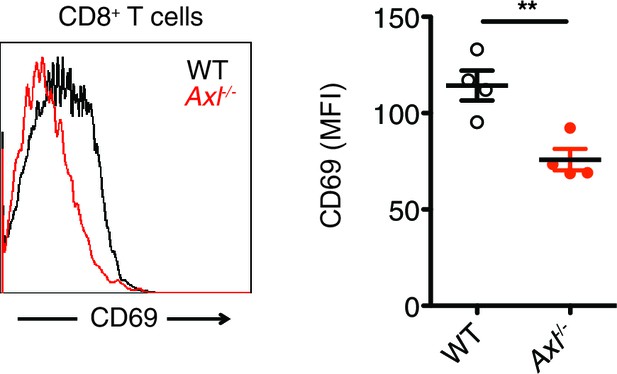
Axl-/- mice display an early defect in CD8+ T cell activation during PR8 infection.
Representative histogram (left) and MFI (right) for CD69 expression on CD8+ T cells in the MLN of WT and Axl-/- mice 3 days post infection with 10 PFU of PR8. n = 4 of each genotype, representative of 2 independent experiments. **p<0.01
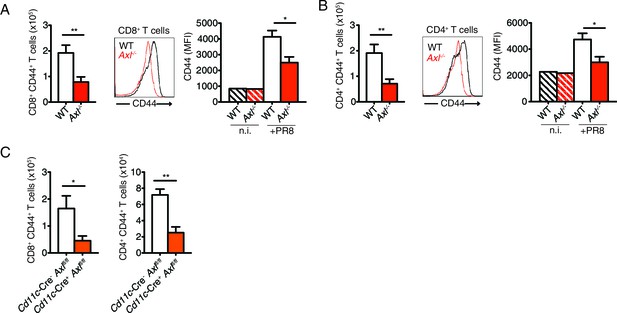
T cells of Axl-/- and Cd11c-Cre+Axlfl/fl mice have reduced CD44 expression during IAV infection.
(A) Number of CD8+CD44+ T cells in the MLN 9 days post-infection with 10 PFU of PR8 (left). n = 5 of each genotype, representative of 4 independent experiments. Representative histogram and MFI (right) for CD44 expression on CD8+ T cells in the lung of WT and Axl-/- mice after 9 days of infection with 10 PFU of PR8. n = 6 of each genotype, representative of 4 independent experiments. (B) Number of CD4+CD44+ T cells in the MLN infected as in (A) (left). n = 5 of each genotype, representative of 4 independent experiments. Representative histogram and MFI (right) for CD44 expression on CD4+ T cells in the lung of WT and Axl-/- mice after 9 days of infection with 10 PFU of PR8. n = 6 of each genotype, representative of 4 independent experiments. (C) Quantification of CD8+CD44+ T cells (left) and CD4+CD44+ T cells (right) in the MLN 9 days post-infection with 10 PFU of PR8 in Cd11c-Cre-Axlfl/fl and Cd11c-Cre+Axlfl/fl mice. 5 mice per genotype, representative of 2 independent experiments. *p<0.05; **p<0.01.
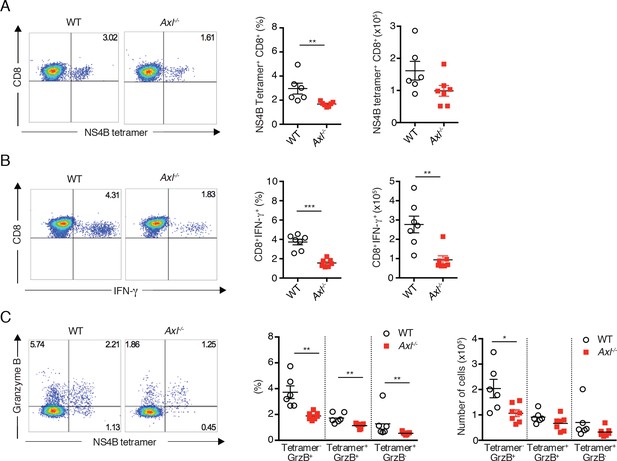
Axl-/- mice mount a deficient CD8+ T cell response to WNV infection.
WT and Axl-/- mice were infected subcutaneously with 102 PFU of WNV, and spleens were harvested 8 days post infection after extensive cardiac perfusion with PBS. (A) Representative flow cytometry plots (left) and percentage and number (right) of NS4B tetramer+ CD8+ T cells. (B) Representative flow cytometry plots (left) and percentage and number (right) of CD8+IFN-γ+ T cells. (C) Representative flow cytometry plots (left) and percentage and number (right) of CD8+ T cells stained for NS4B tetramer and granzyme B. Data are presented as the mean ± SEM of 6–7 mice per genotype. Data are pooled from two independent experiments. p<0.05; **p<0.01; ***p<0.001.
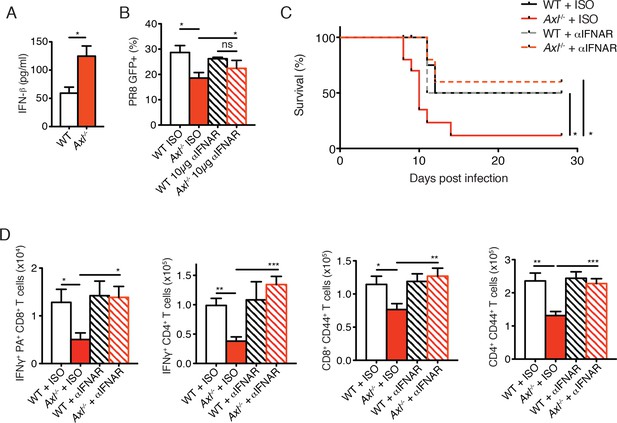
Blockade of IFNAR signaling protects Axl-/- mice to IAV infection and rescues T cell activation.
(A) IFN-β in the supernatant of WT and Axl-/- BMDCs after 12 hr of infection with 0.25 MOI of PR8, as determined by ELISA from 4 independent experiments. (B) Percentage of GFP+ WT and Axl-/- BMDCs infected with 0.05 MOI PR8 for 12 hr treated with 10 μg/ml of IgG1 isotype control or α-IFNAR MAR1-5A3 antibody. Data is compiled from 3 independent experiments. (C) Kaplan-Meier survival curves for WT and Axl-/- mice given α-IFNAR MAR1-5A3 antibody or isotype control by IP injection one day prior to infection with 10 PFU of A/PR8 virus, 8–10 mice per group, 2 independent experiments. (D) WT and Axl-/- mice were treated with antibody and infected as in (C). Number of IFN-γ-producing PA tetramer+ CD8+ T cells (left) and IFN-γ+ CD4+ T cells (middle) in the lung 7 days post infection with 10 PFU of PR8. 4–5 mice in each group, representative of 2 independent experiments. Number of CD8+CD44+ T cells (middle) and CD4+CD44+ T cells (right) in the MLN 9 days post-infection with 10 PFU of PR8. 8–10 mice per group, representing 2 independent experiments. Data are shown as the mean ± SEM. *p<0.05; **p<0.01; ***p<0.001.
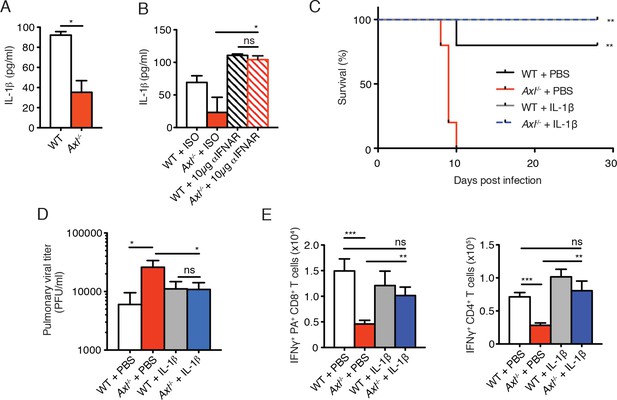
Intranasal administration of IL-1β rescues Axl-/- T cell activation and confers protection to IAV infection.
(A) IL-1β levels in supernatant of WT and Axl-/- BMDCs after 12 hr of infection with 0.25 MOI of PR8, as determined by ELISA from 4 independent experiments. (B) IL-1β in supernatant of WT and Axl-/- BMDCs infected with 0.05 MOI of PR8-GFP for 12 hr treated with 10 μg/ml of isotype control or α-IFNAR MAR1-5A3 antibody, as determined by ELISA from 3 independent experiments. (C-E) WT and Axl-/- mice were intranasally administered PBS or 20 ng of recombinant IL-1β on days 0, 1, 2, and 3 post infection with 10 PFU of PR8. (C) Kaplan-Meier survival curves for mice treated as indicated with 5 mice per group, representative of 4 independent experiments. **Axl-/- mice given PBS succumbed to infection significantly more than the other experimental groups. (D) Viral titers in the bronchoalveolar lavage (BAL) collected 9 days post infection determined by qPCR of PR8 PA RNA. 6–10 mice per group, representing 3 independent experiments. (E) Number of IFN-γ-producing PA tetramer+ CD8+ T cells and IFN-γ+ CD4+ T cells in the lung 7 days post infection with PR8. 4–5 mice in each group, representative of 2 independent experiments. Data are shown as the mean ± SEM. *p<0.05; **p<0.01.
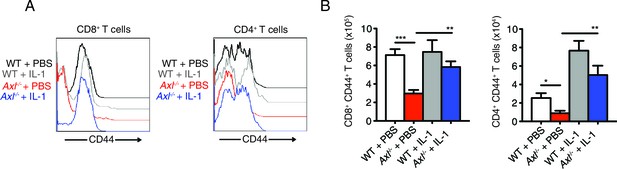
Intranasal IL-1β delivery rescues Axl-/- T cell CD44 expression during IAV infection.
WT and Axl-/- mice were intranasally administered PBS or 20 ng of recombinant IL-1β on days 0, 1, 2, and 3 post infection with 10 PFU of PR8. (A) CD44 expression on CD8+ and CD4+ T cells was assessed by flow cytometry. (B) Number of CD8+CD44+ T cells and CD4+CD44+ T in the lung with 4–5 mice in each group, representative of 3 independent experiments. Data are shown as the mean ± SEM. *p<0.05; **p<0.01; ***p<0.001.
Additional files
-
Supplementary file 1
Primer sequences for the indicated genes.
F = forward, R = reverse.
- https://doi.org/10.7554/eLife.12414.021





