Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons
Figures
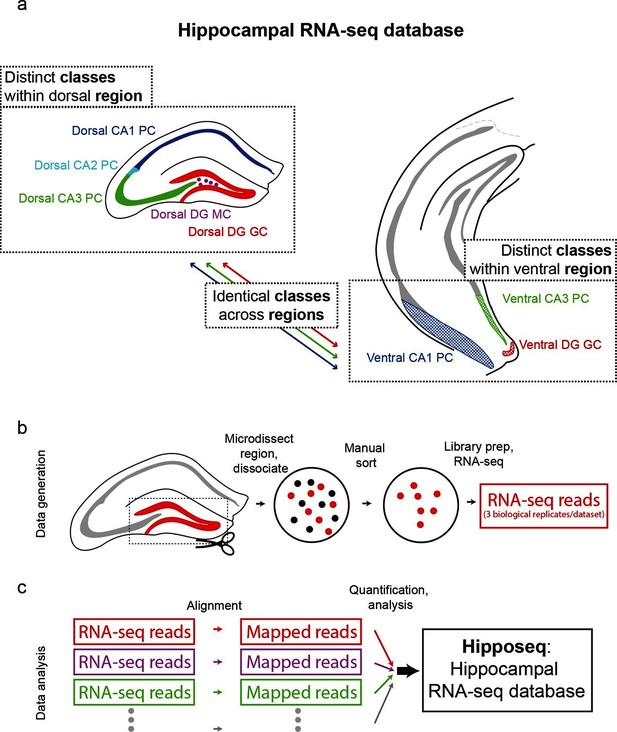
Generation of hippocampal RNA-seq database.
(a) Datasets included in the hippocampal RNA-seq characterization. Note, operationally, cell 'class' refers to gross cell type and 'region' refers to dorsal vs. ventral location. (b) Protocol underlying the generation of raw RNA-seq data. In a transgenic line in which cells of interest were fluorescently labeled (left), the region of interest was microdissected (dashed box). The isolated region was then dissociated, and labeled neurons were manually purified (middle). RNA-seq data was generated from the purified cells. (c) Protocol underlying the processing of RNA-seq data. Raw reads were aligned, and then expression was quantified and statistically analyzed.
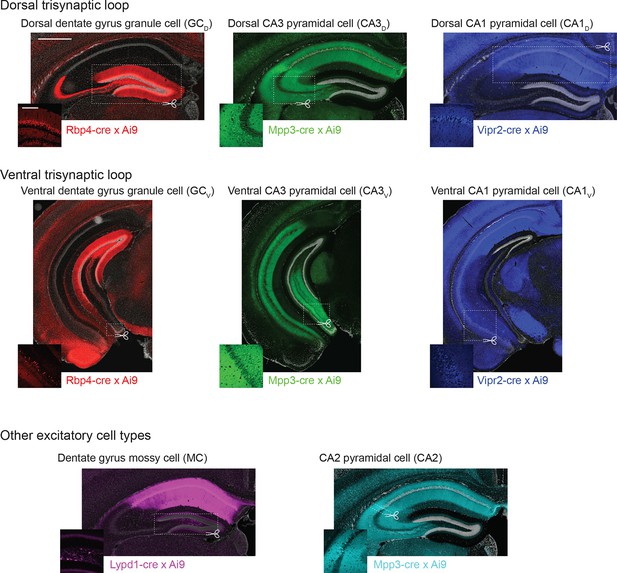

Transgenic lines used to create cell-class- and region-specific transcriptomes.
For each RNA-seq dataset, the corresponding transgenic expression pattern and approximate microdissected region is shown. Scale bar: overview: 500 μm; inset: 200 μm. Images of trisynaptic loop and CA2 expression patterns reprinted from Neuron, 89(2), Cembrowski et al., Spatial Gene-Expression Gradients Underlie Prominent Heterogeneity of CA1 Pyramidal Neurons, 351–368, 2016, with permission from Elsevier.
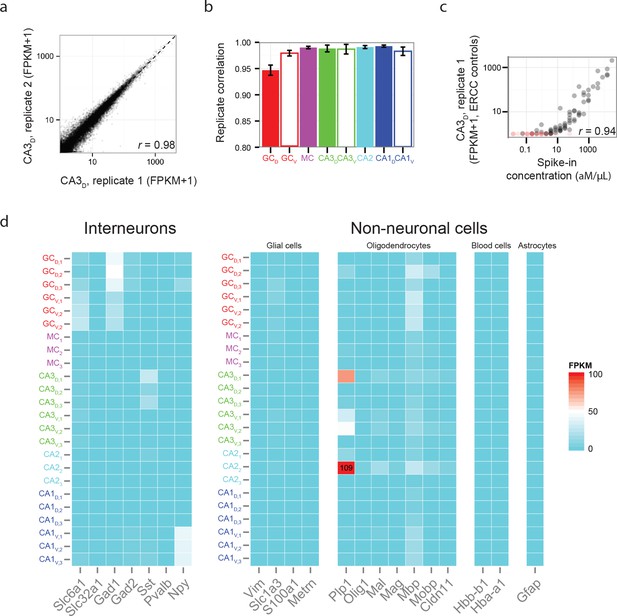
Reproducibility and purity of RNA-seq data.
(a) Representative scatterplot of FPKM values for all genes for two replicates of dorsal CA3, with a Pearson correlation coefficient r = 0.98. (b) Correlation coefficients across replicates for each cell population. (c) Representative FPKM values corresponding to ERCC spike-in controls. Red points indicate undetected spike-in control; i.e., FPKM=0. Here, the Pearson correlation coefficient r = 0.94; for all replicates, r = 0.94 ± 0.01 (n = 24 replicates). (d) FPKM values for genes corresponding to interneurons and non-neuronal cells.
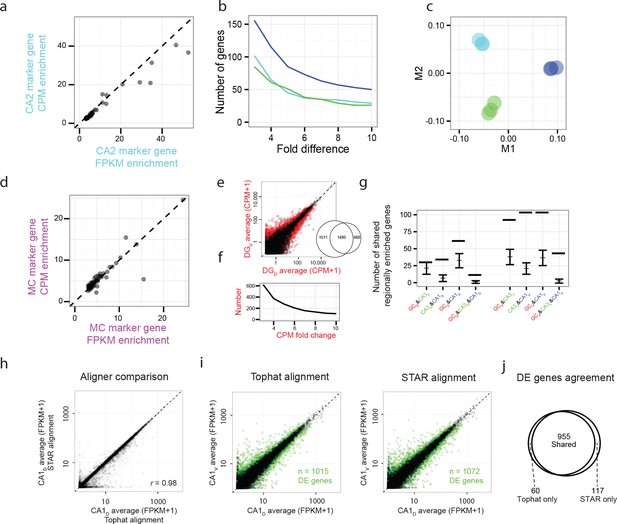
Reproducibility of RNA-seq quantification and differential expression.
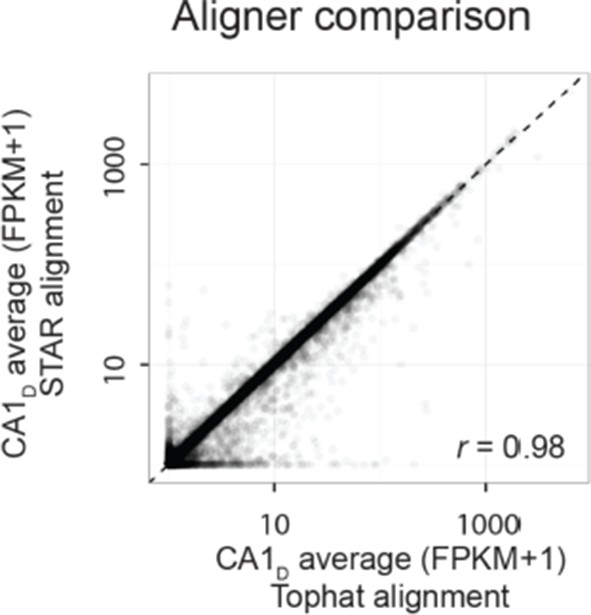
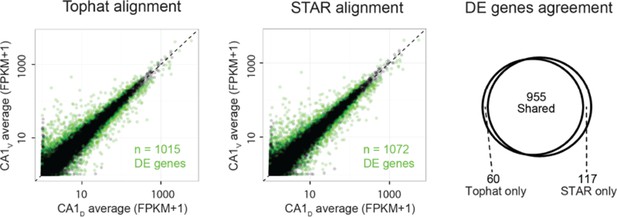
(a) Comparison of FPKM- vs. CPM-based enrichment for CA2 marker genes in Figure 3b. (b,c) As in Figure 3c,d, but for CPM-based analysis. (d) As in a, but for mossy cell marker genes of Figure 4b. (e,f) as in Figure 5b,c, but for CPM-based analysis. Insert: comparison of the number of differentially expressed genes for FPKM- vs. CPM-based approaches. (g) As in Figure 6c, but for CPM-based analysis. (h) Representative example of FPKM values for datasets obtained with TopHat and STAR alignment (dorsal CA1 correlation r = 0.98; all datasets r = 0.98 ± 0.00, Pearson correlation, mean ± SD, n = 8 datasets). (i) Representative example of differential expression results obtained from Tophat and STAR alignment (dorsal vs. ventral CA1: 1015 genes identified using Tophat alignment, 1072 genes identified using STAR alignment). Colored points denote differentially expressed genes, with green color used here to better visualize data points. (j) Overlap in differentially expressed genes from the representative example in i. Here, 955/1015 = 94.1% of genes found using TopHat alignment were also identified with STAR. Across entire dataset, 95.0 ± 1.3% of differentially expressed genes found by TopHat approach were shared with STAR, with STAR identifying 6.8 ± 1.3% more genes than TopHat on average (mean ± SD, n = 28 pairwise comparisons for each).
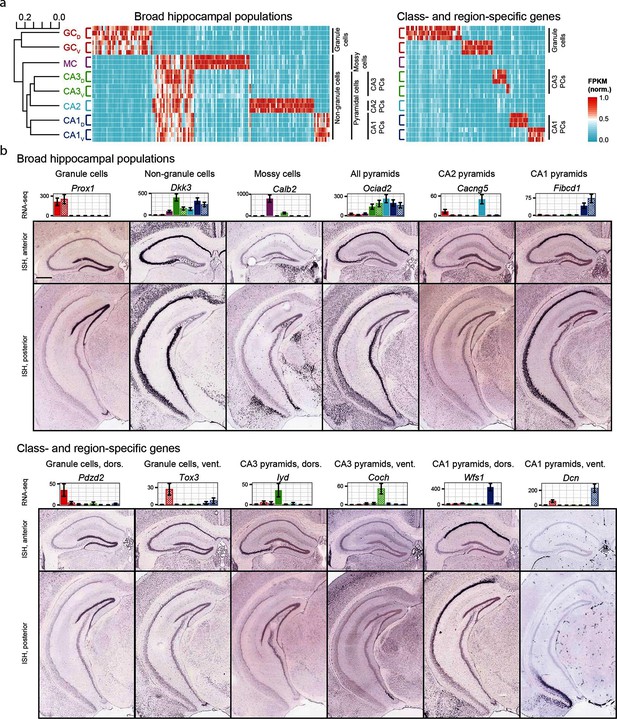
Gene expression in the hippocampus exhibits a variety of cell population- and region-specific expression.
(a) Left: the hierarchical structure of gene expression in the hippocampus calculated by agglomerative clustering. Middle and right: Expression across replicates for marker genes associated with broad hippocampal populations (middle) or specific cell classes and regions (right). Marker genes were selected based upon two-fold enrichment in all replicates in the target population(s) relative to all other replicates (see Materials and methods). FPKM values displayed in the heat map were normalized on a gene-by-gene (i.e., column-by-column) basis by the highest expressing sample for each gene. (b) Confirmation of gene expression profiles by ISH. In corresponding bar plots, RNA-seq FPKM values for each class/region dataset are displayed, with coloring adhering to the conventions of Figure 1a and fill vs. crosshatch indicating dorsal vs. ventral datasets. Scale bar: 500 μm.
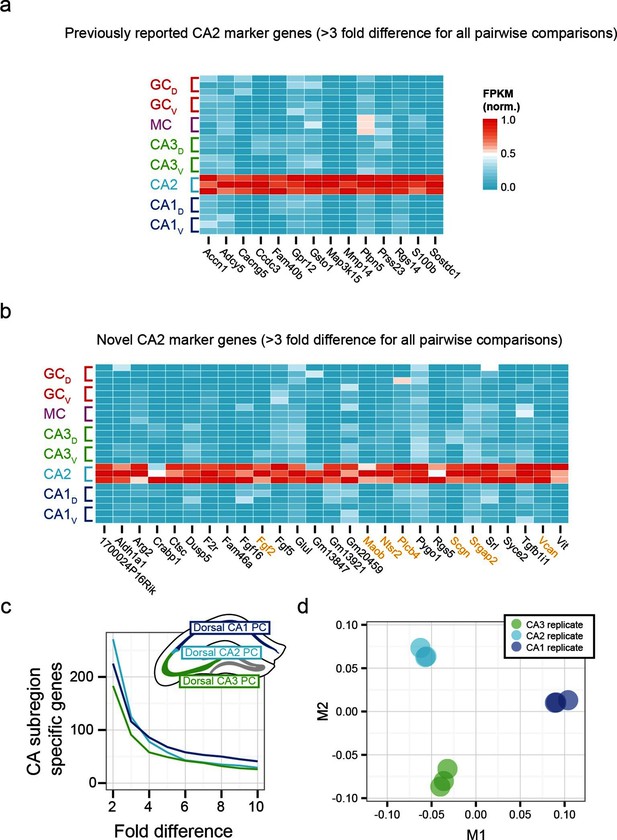
Gene expression properties of CA2 pyramidal cells.
(a) Heat map of replicate FPKM values for previously identified CA2 marker genes. (b) Heat map of replicate FPKM values for novel CA2 marker genes identified by RNA-seq. Orange indicates genes with previously characterized neuronal relevance. (c) The number of CA3-, CA2-, and CA1-specific genes, when restricting comparisons to solely these three cell populations in the dorsal hippocampus. A gene is denoted as X-fold enriched in a given CA region if it the average FPKM value is at least X-fold greater than the other CA regions. (d) Multidimensional scaling demonstrating the distance between CA3, CA2, and CA1 pyramidal cells.
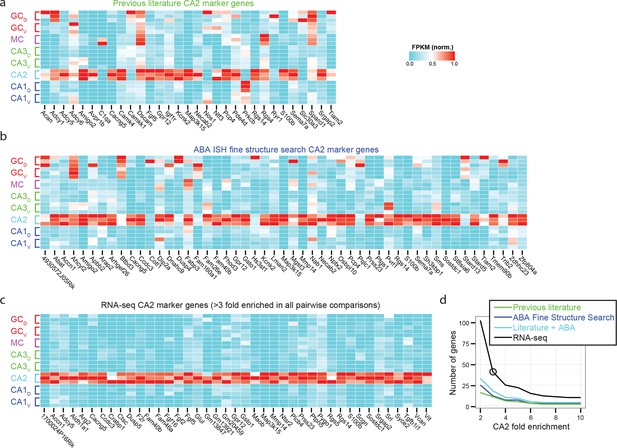
Recapitulation and extension of previous CA2 marker gene results.
(a) Normalized FPKM values for CA2 marker genes identified in previous literature. Note that many previous marker genes have relatively high expression in non-CA2 populations. (b) As in (a), but for all CA2 marker genes identified through ABA ISH CA2 Fine Structure Search. (c) As in (a) and (b), but for RNA-seq-identified marker genes, to compare specificity relative to previous marker genes. (d) Quantitative comparison of the number of marker genes as a function of fold change identified from previous literature (green), ABA ISH fine structure search (blue), the union of previous literature and Fine Structure Search results (teal), and RNA-seq (black). Circular data point illustrates 3-fold enrichment criterion used to obtain RNA-seq marker genes shown in (c).
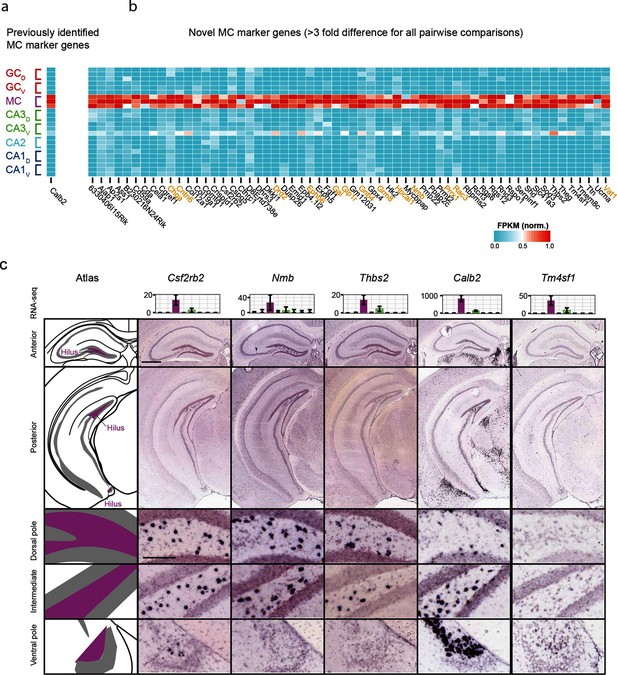
Gene expression properties of hilar mossy cells.
(a) Heat map of replicate FPKM values for the previously identified mossy cell marker gene Calb2. (b) Heat map of replicate FPKM values for novel mossy cell marker genes identified by RNA-seq. Orange indicates genes with previously characterized neuronal relevance. (c) ISH profiles (bottom) for marker genes identified by RNA-seq (top). Scale bar, overview: 500 μm; expanded: 100 μm.
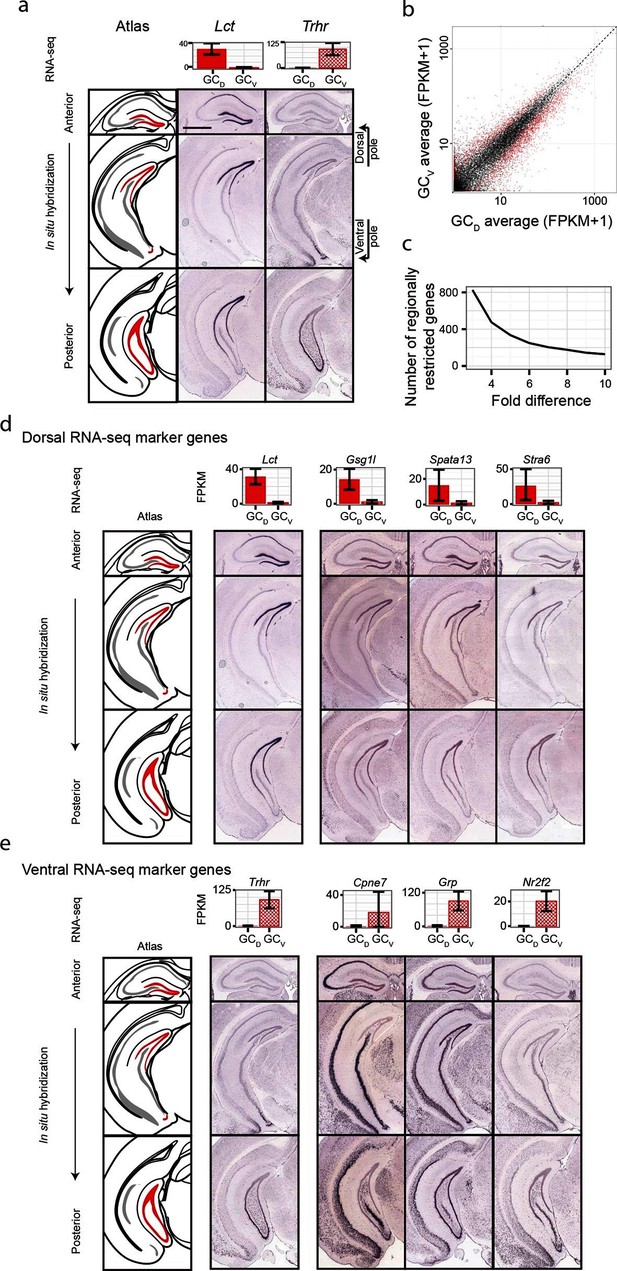
Dorsal-ventral differences in dentate gyrus granule cells.
(a) RNA-seq (top) and ISH (bottom) profiles of Lct and Trhr, two previously identified marker genes respectively enriched in dorsal and ventral granule cells. Scale bar: 500 μm. (b) FPKM scatterplot of average dorsal and ventral GC transcriptomes. Data points represent individual genes, with genes highlighted in red indicating differential expression. (c) Number of genes enriched at the poles of DG as a function of fold change. (d) Example ISH profiles (bottom) of dorsal GC marker genes obtained by RNA-seq (top). (e) As in (d) but for ventral GC marker genes.
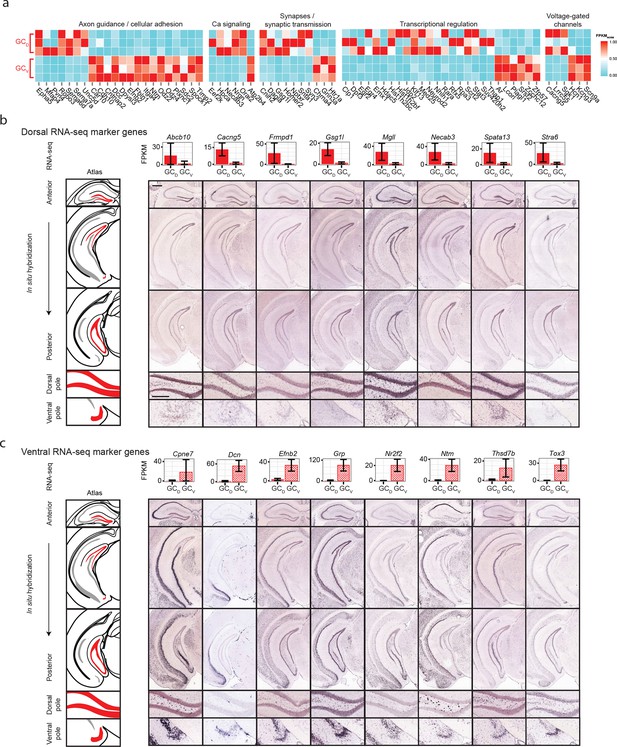

Dorsal-ventral differences in dentate gyrus granule cells.
(a) Examples of regionally enriched marker genes with neuronally relevant functionality. (b) RNA-seq and ISH profiles of representative novel dorsal marker genes. Scale bar, overview: 500 μm; expanded: 100 μm. (c) As in b, but for novel ventral marker genes.
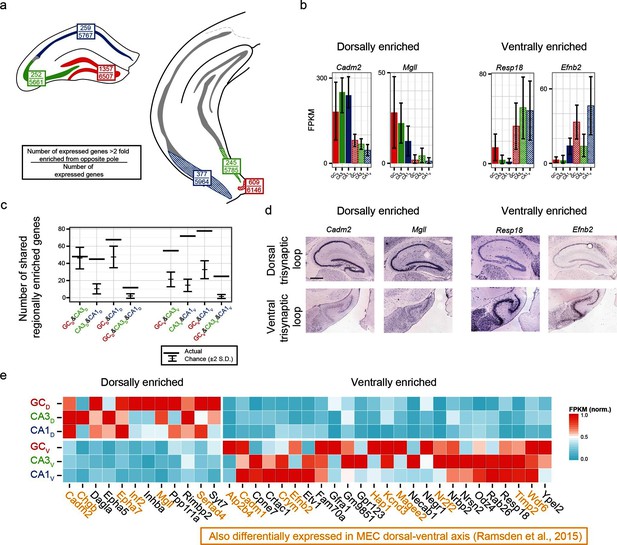
Regionally enriched genes invariant to principal cell class.
(a) For each trisynaptic loop dataset, the number of enriched genes when comparing to the same cell class at the opposite pole (>2-fold difference), as well as the total number of expressed genes for each dataset (FPKMMIN>10), are shown (top and bottom values respectively). (b) Example genes enriched in a region-, but not cell-class-, specific manner. (c) The number of dorsally and ventrally enriched genes shared across cell classes. Both the observed RNA-seq data (horizontal lines) and null distribution (mean ± 2SD) are shown. (d) Sagittal ISH profiles of the example region-enriched genes. Scale bar: 500 μm. (e) Heat map of all genes found enriched in a region-specific manner across the trisynaptic loop (n=37; null distribution predicts 6.0 ± 7.1 (mean ± 2SD), p<1e-6). Orange text: genes also identified as differentially expressed in medial entorhinal cortex (MEC) (Ramsden et al., 2015).
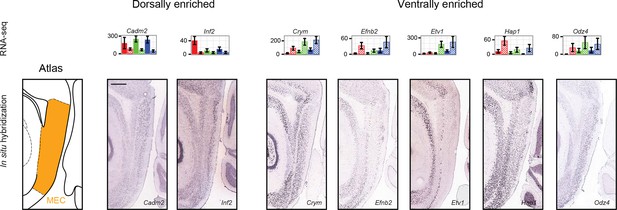
Dorsal and ventral genes enriched across hippocampus and MEC.
Left: atlas showing dorsal-ventral extent of MEC in sagittal section. Middle, right: example ISH profiles of dorsally- and ventrally-enriched genes identified by RNA-seq and found to be predictive of MEC enrichment (Ramsden et al., 2015). Scale bar: 500 μm.
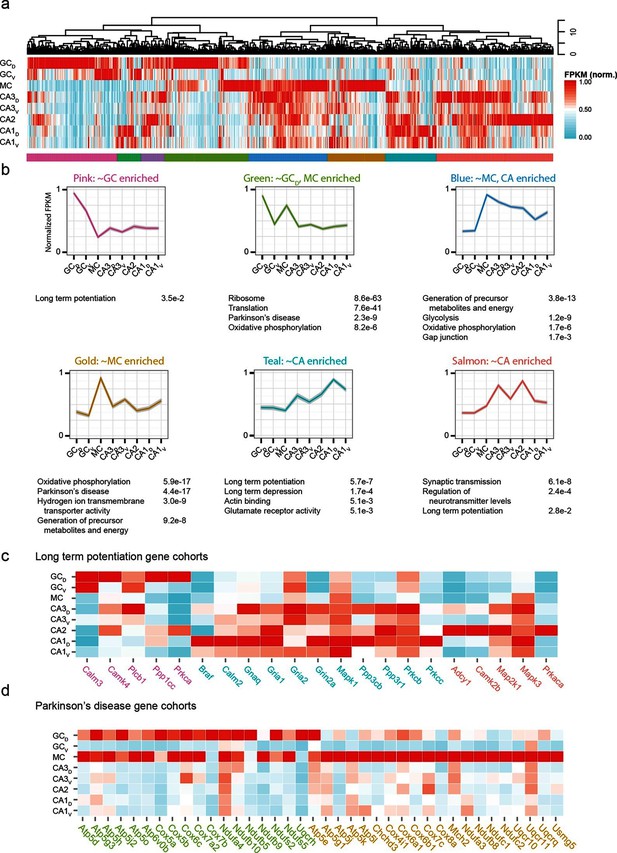
Weighted gene co-expression network analysis (WGCNA) of hippocampal excitatory neuron transcriptomes.
(a) Top and middle: hierarchical clustering and normalized expression of the 1000 most variable genes, respectively. Bottom: colors denoting the modules obtained from WGCNA. (b) Six modules obtained from a, with the average expression shown and significantly enriched terms highlighted. Each module is named according to the gross overall expression profile across datasets. (c) Genes associated with long-term potentiation significantly enriched in modules. (d) As in c, but with genes associated with Parkinson’s disease.
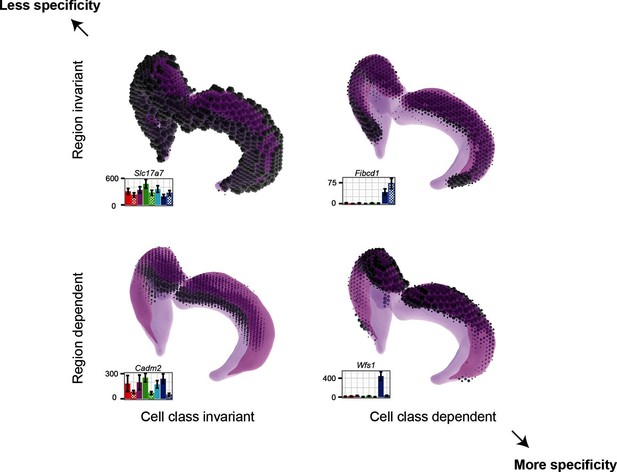
Gene expression patterns of excitatory hippocampal neurons.
Genes can be expressed in relatively similar abundances across the hippocampus (Slc17a7; upper left), vary in a cell-class-specific (Fibcd1; upper right) or region-specific manner (Cadm2; lower left), or vary in both cell-class- and region-specific manners simultaneously (Wfs1; lower right). In each panel, light magenta denotes the spatial extent of the hippocampus, dark magenta illustrates the CA1 region in particular, and the dots indicate the location and intensity of labeling from ISH. RNA-seq profiling results are provided for each gene. Images from the Allen Brain Explorer v2.
Additional files
-
Supplementary file 1
Dendrogram marker genes.
- https://doi.org/10.7554/eLife.14997.017
-
Supplementary file 2
Marker genes for dentate gyrus mossy cells.
- https://doi.org/10.7554/eLife.14997.018
-
Supplementary file 3
Dorsal and ventral dentate gyrus granule cell marker genes.
- https://doi.org/10.7554/eLife.14997.019
-
Supplementary file 4
List of Allen Mouse Brain Atlas images shown in text
- https://doi.org/10.7554/eLife.14997.020





